 Back
BackDNA Replication: Mechanisms, Enzymes, and Experimental Evidence
Study Guide - Smart Notes

DNA Replication: Overview and Significance
Introduction to DNA Replication
DNA replication is a fundamental process that ensures the accurate transmission of genetic information from one generation to the next. It occurs during the S phase of the cell cycle and is essential for cell division and organismal growth.
Replication Timing: DNA synthesis occurs in the S phase, followed by chromosome segregation in M phase.
Genetic Continuity: Accurate replication ensures progeny cells inherit the same genetic information as the parent cell.
Potential for Variation: Occasional errors or mutations during replication provide genetic diversity for evolution.
Structure of DNA and Its Role in Replication
Watson and Crick Model
The double helix model of DNA, proposed by Watson and Crick in 1953, revealed the molecular basis for heredity and replication. The structure consists of two antiparallel strands with a sugar-phosphate backbone and complementary base pairing (A-T, G-C) held together by hydrogen bonds.
Antiparallel Strands: One strand runs 5' to 3', the other 3' to 5'.
Complementary Base Pairing: Adenine pairs with thymine, guanine with cytosine.
Template Function: Each strand can serve as a template for the synthesis of a new complementary strand.
Mechanisms of DNA Replication
Models of Replication
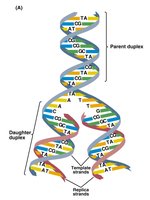
Three models were proposed for DNA replication: semiconservative, conservative, and dispersive. The semiconservative model, supported by experimental evidence, states that each daughter DNA molecule consists of one parental and one newly synthesized strand.
Semiconservative: Each new DNA has one old and one new strand.
Conservative: Parental strands stay together; new strands form a separate molecule.
Dispersive: Parental and new DNA are interspersed in both strands.
Meselson-Stahl Experiment
The 1958 Meselson-Stahl experiment provided strong evidence for the semiconservative model. By growing E. coli in heavy (15N) and light (14N) nitrogen and analyzing DNA density after replication, they demonstrated that each daughter DNA molecule contained one old and one new strand after one replication cycle.
Experimental Design: Bacteria were grown in 15N, then switched to 14N, and DNA was analyzed after each generation.
Results: After one generation, DNA was of intermediate density; after two, both light and intermediate bands appeared, ruling out conservative and dispersive models.

Enzymes and Proteins in DNA Replication
DNA Polymerases
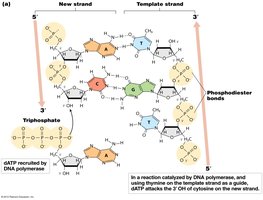
DNA polymerases are the main enzymes responsible for synthesizing new DNA strands. They add nucleotides to the 3' end of a growing DNA chain, using a template strand and a primer with a free 3' OH group.
Directionality: DNA polymerases synthesize DNA only in the 5' to 3' direction.
Proofreading: Many DNA polymerases have 3' to 5' exonuclease activity, allowing them to remove incorrectly paired bases and improve fidelity.
Substrates: Four deoxynucleoside triphosphates (dATP, dGTP, dCTP, dTTP) are required.
Other Key Proteins
Helicase: Unwinds the DNA double helix at the replication fork.
Single-Stranded Binding Proteins (SSB): Stabilize unwound DNA strands.
Primase: Synthesizes short RNA primers to provide a starting point for DNA polymerase.
Ligase: Joins Okazaki fragments on the lagging strand by forming phosphodiester bonds.
Topoisomerase: Relieves supercoiling ahead of the replication fork.
Mechanics of DNA Replication
Replication Fork and Strand Synthesis
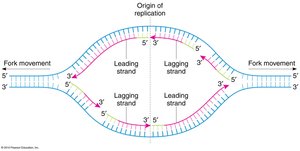
Replication begins at origins of replication, forming a replication bubble with two replication forks. DNA synthesis is continuous on the leading strand and discontinuous on the lagging strand, forming Okazaki fragments.
Leading Strand: Synthesized continuously in the direction of fork movement.
Lagging Strand: Synthesized discontinuously in short fragments, later joined by ligase.
Prokaryotic vs. Eukaryotic DNA Replication
Prokaryotic Replication (e.g., E. coli)
Prokaryotes typically have a single, circular chromosome with one origin of replication. Replication proceeds bidirectionally until the entire chromosome is copied.
Origin of Replication (oriC): Specific sequence where replication begins.
Bidirectional Synthesis: Two replication forks move in opposite directions.
Eukaryotic Replication
Eukaryotic chromosomes are linear and much larger, with multiple origins of replication per chromosome. Special mechanisms are required to replicate chromosome ends (telomeres).
Multiple Origins: Allow rapid replication of large genomes.
Telomeres: Repetitive DNA sequences at chromosome ends, maintained by telomerase to prevent loss of genetic information.
Telomeres and the End-Replication Problem
Structure and Function of Telomeres
Telomeres are repetitive, non-coding DNA sequences at the ends of linear chromosomes. They protect chromosome ends from degradation and prevent the loss of important genetic information during replication.
End-Replication Problem: DNA polymerases cannot fully replicate the 3' ends of linear chromosomes, leading to gradual shortening.
Telomerase: An enzyme with reverse transcriptase activity that extends telomeres using an RNA template.
Summary Table: Key Enzymes and Functions in DNA Replication
Enzyme/Protein | Function |
|---|---|
DNA Polymerase | Synthesizes new DNA strands; proofreads and corrects errors |
Helicase | Unwinds the DNA double helix |
Primase | Synthesizes RNA primers |
Ligase | Joins Okazaki fragments |
SSB Proteins | Stabilize single-stranded DNA |
Topoisomerase | Relieves supercoiling |
Telomerase | Extends telomeres in eukaryotes |
Key Concepts and Equations
Base Pairing Rules: A pairs with T, G pairs with C.
Direction of Synthesis: DNA is always synthesized in the 5' to 3' direction.
Phosphodiester Bond Formation: The 3' OH of the growing strand attacks the 5' phosphate of the incoming dNTP, releasing pyrophosphate.
Example: In the Meselson-Stahl experiment, after two generations in 14N medium, DNA molecules were either hybrid (one 15N and one 14N strand) or light (both 14N strands), confirming semiconservative replication.
