 Back
BackDNA Replication: Mechanisms, Models, and Chromosome Structure
Study Guide - Smart Notes

DNA Replication and Cell Cycle
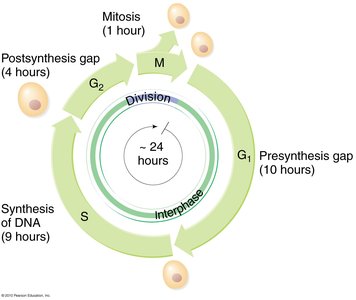
Phases of the Cell Cycle
The cell cycle is divided into distinct phases, each with specific functions related to cell growth and division. DNA replication occurs during the S phase, ensuring that genetic material is duplicated before cell division.
G1 Phase: Presynthesis gap, cell growth and preparation for DNA replication.
S Phase: Synthesis of DNA, chromosomes are duplicated.
G2 Phase: Post-synthesis gap, preparation for mitosis.
M Phase: Mitosis, division of the cell into two daughter cells.
Structure of Chromosomes and Chromatids
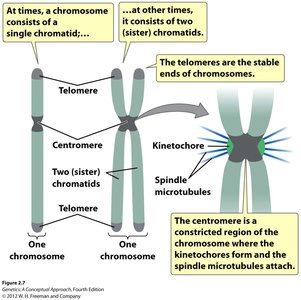
Chromosome Organization
Chromosomes are composed of DNA and associated proteins. They can exist as single chromatids or as pairs of sister chromatids after DNA replication. Key structural features include telomeres and centromeres.
Telomeres: Stable ends of chromosomes, protect genetic material during replication.
Centromere: Constricted region where kinetochores form and spindle microtubules attach during mitosis.
Sister Chromatids: Identical copies formed after DNA replication, joined at the centromere.
Molecular Structure of DNA
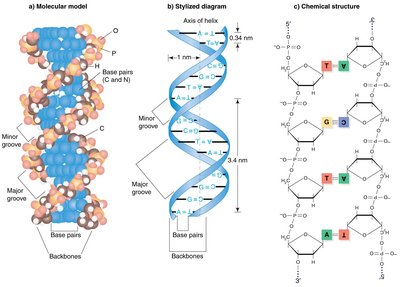
Watson and Crick Model
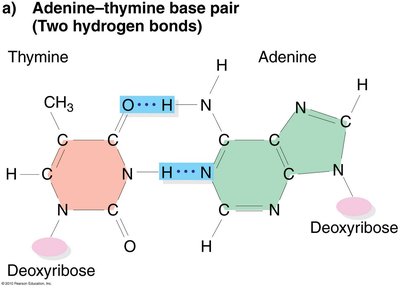
DNA is a right-handed double helix with a sugar-phosphate backbone on the outside and complementary base pairs on the inside. The strands are antiparallel and held together by hydrogen bonds.
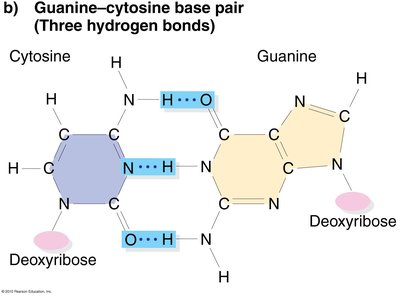
Base Pairing: Adenine (A) pairs with Thymine (T), Guanine (G) pairs with Cytosine (C).
Antiparallel Strands: One strand runs 5' to 3', the other 3' to 5'.
Hydrogen Bonds: Two between A-T, three between G-C.
Models of DNA Replication
Three Proposed Models
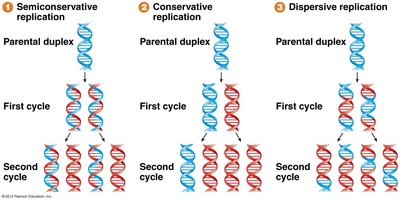
Before the mechanism was established, three models were proposed for DNA replication:
Semiconservative Replication: Each daughter DNA molecule contains one old (parental) strand and one newly synthesized strand.
Conservative Replication: The parental DNA remains intact, and a completely new molecule is synthesized.
Dispersive Replication: Parental DNA is fragmented and dispersed throughout the daughter molecules.

Experimental Evidence: Meselson-Stahl Experiment
Demonstrating Semiconservative Replication
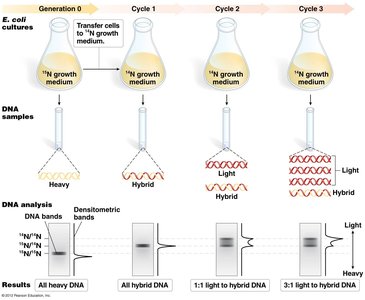
The Meselson-Stahl experiment used isotopes of nitrogen (15N and 14N) to distinguish between old and new DNA strands. E. coli was grown in heavy (15N) and then light (14N) media, and DNA was analyzed by density gradient centrifugation.
Results: After one replication cycle, DNA was of intermediate density, ruling out conservative replication. After two cycles, both intermediate and light DNA were observed, ruling out dispersive replication.
Conclusion: DNA replication is semiconservative.
Mechanism of DNA Replication
Replication Fork and Strand Synthesis
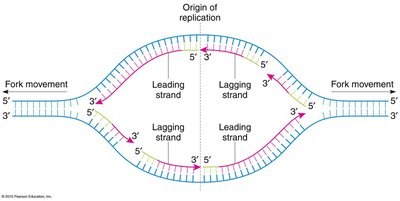
DNA replication begins at origins of replication, forming replication forks where the double helix is unwound. Two types of strand synthesis occur:
Leading Strand: Synthesized continuously in the direction of the replication fork movement.
Lagging Strand: Synthesized discontinuously in short fragments (Okazaki fragments) opposite the fork movement, later joined by DNA ligase.
Enzymes Involved in DNA Replication
Several enzymes coordinate the replication process:
DNA Polymerase: Synthesizes new DNA by adding nucleotides to the 3' end of a primer.
Primase: Synthesizes short RNA primers to initiate DNA synthesis.
Helicase: Unwinds the DNA double helix.
Single-Stranded Binding Proteins (SSB): Stabilize unwound DNA.
DNA Ligase: Joins Okazaki fragments on the lagging strand.
Topoisomerase: Relieves supercoiling ahead of the replication fork.
Chromosome Structure and Telomeres
Telomeres and the End-Replication Problem
Linear chromosomes in eukaryotes face the end-replication problem, where the very ends cannot be fully replicated by DNA polymerase. Telomeres, repetitive DNA sequences at chromosome ends, protect genetic information from loss.
Telomerase: An enzyme with RNA and protein components that extends telomeres by adding repeats to the 3' end.
Function: Maintains chromosome integrity and prevents loss of essential genes during replication.

Summary Table: DNA Replication Models
Model | First Cycle Result | Second Cycle Result | Supported by Meselson-Stahl? |
|---|---|---|---|
Semiconservative | Hybrid DNA | Hybrid and light DNA | Yes |
Conservative | Heavy and light DNA | Heavy and light DNA | No |
Dispersive | Hybrid DNA | Hybrid DNA only | No |
Key Terms and Concepts
DNA Polymerase: Enzyme responsible for DNA synthesis.
Okazaki Fragments: Short DNA fragments synthesized on the lagging strand.
Telomere: Repetitive DNA sequence at chromosome ends.
Semiconservative Replication: Each new DNA molecule contains one old and one new strand.
Replication Fork: Y-shaped region where DNA is actively being unwound and replicated.
Important Equations
Phosphodiester Bond Formation:
Base Pairing:
