 Back
BackDNA Replication: Semiconservative Mechanism and Experimental Evidence
Study Guide - Smart Notes

DNA Replication and Experimental Evidence
Chargaff’s Rules and DNA Base Composition
Erwin Chargaff’s findings were foundational for understanding DNA structure. He discovered that in double-stranded DNA, the amount of adenine (A) equals thymine (T), and the amount of cytosine (C) equals guanine (G). However, the total amount of (C + G) does not necessarily equal (A + T), and the ratios of purines to pyrimidines are consistent within species but vary between species.
Chargaff’s Rules: A = T and C = G in double-stranded DNA.
Implication: The base-pairing rules support the double-helix model and the mechanism of replication.
Incorrect Statement: (C + G) = (A + T) is not consistent with Chargaff’s findings.
Structure of Nucleotides in DNA and RNA
Nucleotides are the building blocks of nucleic acids. Each nucleotide consists of a phosphate group, a five-carbon sugar (ribose in RNA, deoxyribose in DNA), and a nitrogenous base (purine or pyrimidine).
Purines: Double-ring structures (adenine and guanine).
Pyrimidines: Single-ring structures (cytosine, thymine, and uracil).
RNA vs. DNA: RNA contains ribose (with a 2'-OH group), while DNA contains deoxyribose (lacking the 2'-OH group).
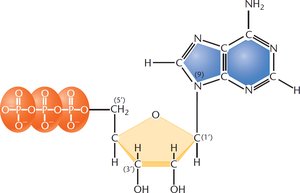
Example: The molecule shown above is a ribonucleotide (RNA) with a purine base, as indicated by the presence of ribose and the double-ring structure.
Chapter 11: DNA Replication and Recombination
Semiconservative Replication of DNA
DNA replication is semiconservative, meaning each new DNA molecule consists of one parental (old) strand and one newly synthesized strand. The complementarity of DNA strands allows each to serve as a template for the other, ensuring accurate genetic information transfer.
Template Mechanism: Each strand guides the synthesis of its complement.
Semiconservative Model: After replication, each daughter DNA contains one old and one new strand.
Experimental Evidence: The Meselson–Stahl Experiment
The Meselson–Stahl experiment (1958) provided strong evidence for semiconservative replication in prokaryotes using Escherichia coli and isotopic labeling.
Step 1: Grow E. coli in medium containing heavy nitrogen (15N) to label DNA.
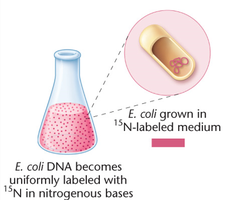
Step 2: Transfer bacteria to normal (14N) medium and allow replication for several generations.

Step 3: Isolate DNA and separate by density using equilibrium density gradient centrifugation with cesium chloride (CsCl).
Observation: After one generation, DNA is of intermediate density (hybrid 15N/14N), supporting the semiconservative model. With further generations, more DNA is light (14N), but hybrid DNA persists.
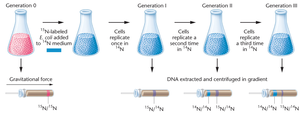
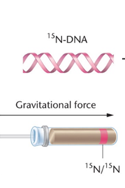
Conclusion: The experiment demonstrated that DNA replication is semiconservative, not conservative or dispersive.
Semiconservative Replication in Eukaryotes
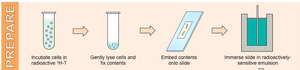
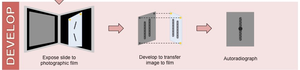
Semiconservative replication also occurs in eukaryotes, as shown by the Taylor–Woods–Hughes experiment (1957) in Vicia faba (broad bean). They used radioactive 3H-thymidine labeling and autoradiography to track newly synthesized DNA.
Autoradiography: A technique to visualize the location of radioactively labeled molecules in cells.
Result: Only one chromatid is labeled after one round of replication, confirming semiconservative replication.
Origin and Directionality of DNA Replication
DNA replication begins at specific sites called origins of replication (ORI). In bacteria, there is a single origin (OriC) on the circular chromosome. Replication is bidirectional, creating two replication forks that move away from the origin.
Replication Fork: The Y-shaped region where the DNA is unwound and new strands are synthesized.
Replicon: The length of DNA replicated from a single origin.
Bacterial Chromosomes: Typically have one origin and replicate bidirectionally.
Summary Table: Key Features of DNA Replication
Feature | Description |
|---|---|
Replication Model | Semiconservative |
Experimental Evidence | Meselson–Stahl (prokaryotes), Taylor–Woods–Hughes (eukaryotes) |
Origin of Replication | Single (bacteria), multiple (eukaryotes) |
Directionality | Bidirectional |
Key Techniques | Isotopic labeling, density gradient centrifugation, autoradiography |
Additional info: The above notes integrate foundational concepts, experimental evidence, and technical terminology to provide a comprehensive overview of semiconservative DNA replication, as required for college-level genetics study.
