 Back
BackDNA Structure, Chromatin Organization, and Replication: Study Notes for Genetics Students
Study Guide - Smart Notes

DNA Structure and Analysis
Properties of Genetic Material
For a molecule to serve as genetic material, it must be capable of replication, storing information, expressing information, and varying by mutation. DNA fulfills all these criteria, as demonstrated by classic experiments.
Replication: DNA can make exact copies of itself.
Information Storage: DNA encodes genetic instructions.
Expression: DNA directs cellular functions via transcription and translation.
Variation: DNA can mutate, leading to genetic diversity.
Experimental Evidence for DNA as Genetic Material
Griffith's Transformation Experiment: Demonstrated that a "transforming principle" could convert avirulent bacteria into virulent forms.
Avery, MacLeod, and McCarty: Identified DNA as the transforming principle by showing that only DNA, not protein or RNA, could induce transformation.
Hershey-Chase Experiment: Used radioisotopes to show that DNA, not protein, enters bacteria during phage infection and directs viral reproduction.
Evidence in Eukaryotes
Distribution: DNA is localized where genetic functions occur (nucleus).
Mutagenesis: UV light is most mutagenic at 260 nm, matching DNA's absorption peak.
Recombinant DNA: DNA can be spliced and replicated in cells, confirming its genetic role.
RNA Viruses: Some viruses use RNA as genetic material, relying on RNA replicase or reverse transcriptase.
Nucleic Acid Chemistry
Nucleotides and Nitrogenous Bases
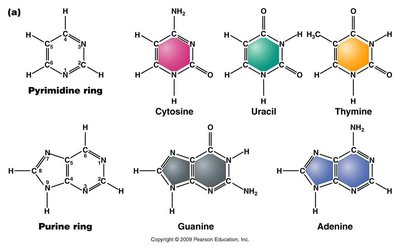
Nucleotides are the building blocks of nucleic acids, consisting of a nitrogenous base, a pentose sugar, and a phosphate group. There are two types of nitrogenous bases: purines and pyrimidines.
Pyrimidines: Single-ring structures (cytosine, thymine, uracil).
Purines: Double-ring structures (adenine, guanine).
Hydrogen Bonding: C-G pairs form three bonds (stronger), A-T/U pairs form two bonds.
Pentose Sugars
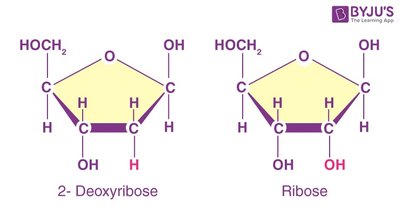
The pentose sugar distinguishes DNA from RNA. DNA contains deoxyribose (lacking an oxygen at the 2' carbon), while RNA contains ribose (with an OH group at the 2' carbon).
Deoxyribose: Less reactive, found in DNA.
Ribose: More reactive, found in RNA.
Nucleosides and Nucleotides
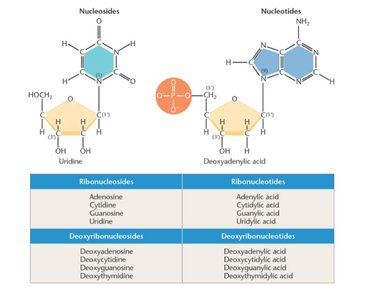
A nucleoside consists of a base and a sugar; a nucleotide adds a phosphate group. Nucleoside triphosphates are precursors for nucleic acid synthesis.
Nucleoside: Base + sugar.
Nucleotide: Base + sugar + phosphate.
Phosphodiester Bond: Links nucleotides in a chain, forming the backbone of DNA/RNA.
Chargaff's Rules and DNA Structure
Chargaff found that the amount of adenine equals thymine, and cytosine equals guanine. Rosalind Franklin's X-ray diffraction revealed the helical structure. Watson and Crick's model described DNA as a right-handed double helix with antiparallel strands, complementary base pairing, and major/minor grooves.
Helix Diameter: 20 Å
Base Pair Spacing: 3.4 Å
Helix Turn: 10.4 base pairs per turn
Stability: Thousands of hydrogen bonds and hydrophobic interactions
Alternative DNA Forms
B-DNA: Standard form, right-handed helix.
A-DNA: More compact, bases tilted.
Z-DNA: Left-handed helix.
RNA Structure
RNA is similar to DNA but contains ribose, uracil instead of thymine, and is usually single-stranded. Types include rRNA (ribosomal), mRNA (messenger), and tRNA (transfer).
rRNA: Structural component of ribosomes.
mRNA: Carries genetic information to ribosomes.
tRNA: Transfers amino acids during translation.
Analytical Techniques
Denaturation: Heat causes DNA to unwind; melting temperature (Tm) depends on GC content.
Molecular Hybridization: Denatured nucleic acids can re-anneal; used in probe detection.
Electrophoresis: Separates nucleic acids by size under an electric field.
DNA Organization in Chromatin
Chromatin Structure in Eukaryotes
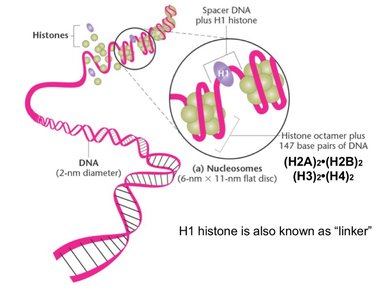
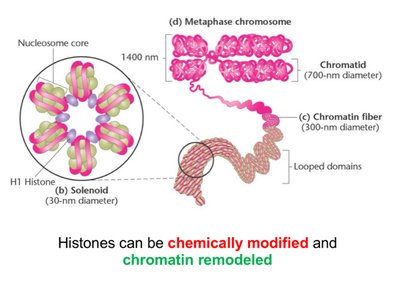
Chromatin is the complex of DNA and proteins found in eukaryotic cells. DNA wraps around histone proteins to form nucleosomes, which further compact into chromatin fibers and chromosomes.
Histones: Basic proteins (H2A, H2B, H3, H4) form octamers; H1 is a linker.
Nucleosome: DNA wrapped around histone octamer (~200 base pairs).
Solenoid: Higher-order structure formed by stacked nucleosomes.
Chromatin Remodeling: Chemical modifications (acetylation, methylation, phosphorylation) alter compaction and gene expression.
DNA Replication
Semiconservative Replication
DNA replication is semiconservative: each new molecule contains one old and one new strand. The Meselson-Stahl experiment confirmed this in prokaryotes; Taylor, Woods, and Hughes confirmed it in eukaryotes.
Replication Fork: Site where DNA is unwound for replication.
Replicon: Region of DNA replicated from a single origin.
Bidirectional Replication: Occurs in both directions from the origin.
DNA Synthesis in Bacteria
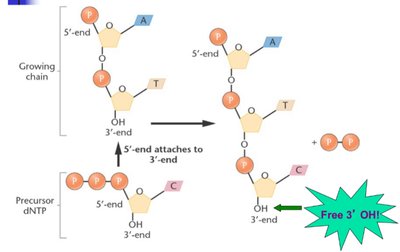
DNA polymerases catalyze DNA synthesis, requiring a template, dNTPs, Mg2+, and a primer. Chain elongation occurs in the 5' to 3' direction, with energy provided by the release of pyrophosphate.
DNA Polymerase I: Removes primer, gap-filling, proofreading.
DNA Polymerase II: DNA repair.
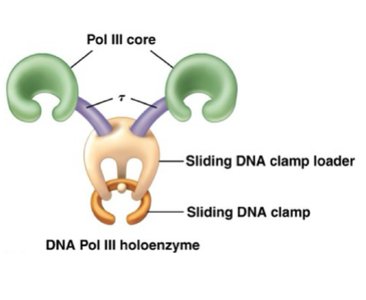
DNA Polymerase III: Main enzyme for DNA synthesis, proofreading.
Exonuclease Activity: 3'-5' for proofreading; only Pol I has 5'-3' exonuclease.
Primer: Required for initiation; synthesized by primase.
Unwinding and Initiation
DnaA: Initiator protein binds to origin, opens helix.
Helicase: Unzips DNA, requires ATP.
Single-Stranded Binding Proteins: Stabilize unwound DNA.
DNA Gyrase: Topoisomerase relieves supercoiling.
Primase: Synthesizes RNA primer for DNA polymerase.
Leading and Lagging Strand Synthesis
Leading Strand: Synthesized continuously.
Lagging Strand: Synthesized discontinuously as Okazaki fragments, joined by DNA ligase.
Eukaryotic DNA Replication
Multiple Origins: Replication bubbles in linear chromosomes.
Origin Recognition Complex (ORC): Tags origins for initiation.
Polymerases: α, δ, and ε required for nuclear DNA replication.
Telomeres: Specialized sequences at chromosome ends; telomerase extends telomeres using an RNA template.
The Genetic Code and Transcription
The Genetic Code
The genetic code is a linear sequence of triplet codons, each specifying an amino acid. It is degenerate (multiple codons per amino acid), unambiguous, non-overlapping, and nearly universal.
Start Codon: AUG (methionine)
Stop Codons: UAA, UAG, UGA
Frameshift Mutations: Addition or deletion of nucleotides shifts the reading frame.
Wobble Hypothesis: Third base in codon is less critical, allowing relaxed pairing.
Transcription in Bacteria
RNA polymerase synthesizes RNA from DNA without a primer. Promoters (TATA box, TTGACA) are recognized by sigma factors. Transcription proceeds through initiation, elongation, and termination (intrinsic or rho-dependent).
Consensus Sequences: Cis-acting elements regulate transcription.
Trans-acting Factors: Proteins that bind to cis-elements.
Termination: Hairpin structures or rho factor cause dissociation.
Transcription in Eukaryotes
Occurs in the nucleus, requires chromatin remodeling, multiple RNA polymerases, and general transcription factors. Promoters, enhancers, and silencers modulate transcription.
RNA Processing: Includes 5' capping, 3' poly-A tail addition, and splicing to remove introns.
Spliceosome: Removes introns, joins exons.
mRNA Export: Nuclear export receptors guide mRNA through nuclear pores.
Translation and Proteins
Ribosomes and tRNA
Ribosomes are composed of rRNA and proteins, with large and small subunits. tRNA carries amino acids to the ribosome, with a specific anticodon and amino acid binding site.
Charging tRNA: Aminoacyl tRNA synthetase links amino acid to tRNA using ATP.
Translation in Prokaryotes and Eukaryotes
Initiation: Small subunit binds mRNA, initiator tRNA, and initiation factors.
Elongation: Peptide bonds formed, ribosome shifts, tRNA moves through E, P, A sites.
Termination: Stop codon recognized by release factors, polypeptide released.
Post-Translational Modifications: Proteins are modified, folded, and tagged for degradation if misfolded.
Protein Domains: Modular regions determine function and location; exon shuffling creates new genes.
Additional info: These notes cover DNA structure, chromatin organization, replication, transcription, and translation, with relevant images included to reinforce key concepts.
