 Back
BackEpigenetics and Gene Regulation in Eukaryotes
Study Guide - Smart Notes

Gene Regulation in Eukaryotes: Epigenetics
Introduction to Epigenetics
Epigenetics refers to heritable changes in gene expression that do not involve alterations to the underlying DNA sequence. These changes are mediated by chemical modifications to DNA and histone proteins, as well as by noncoding RNAs. Epigenetic mechanisms play a crucial role in regulating gene activity, cellular differentiation, and development.
Epigenetic traits are heritable and can be passed on during cell division.
Epigenetic modifications include DNA methylation, histone modification, and noncoding RNAs.
Epigenetic changes can be influenced by environmental factors, diet, and lifestyle.
DNA Methylation
DNA methylation is a key epigenetic modification involving the addition of a methyl group to the 5' position of cytosine residues, primarily at CpG dinucleotides. This process is catalyzed by DNA methyltransferases and is often associated with gene silencing.
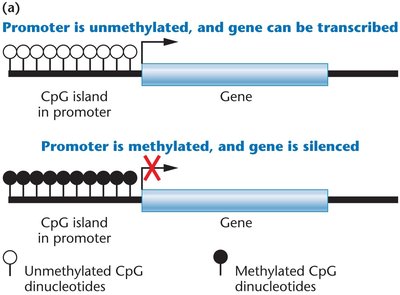
Methylation occurs at CpG islands, which are regions rich in CpG dinucleotides, often found near gene promoters.
Unmethylated CpG islands in promoters are typically associated with active gene transcription, while methylated CpG islands lead to gene silencing.
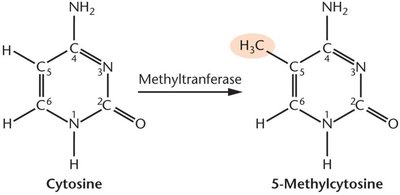
The methyl group is added to cytosine, forming 5-methylcytosine.
Maintenance of DNA Methylation
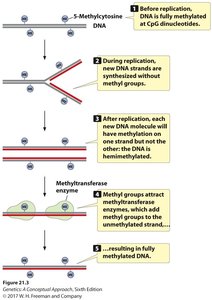
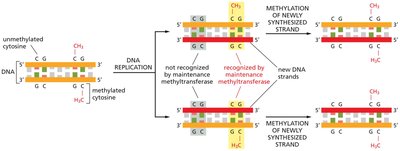
During DNA replication, methylation patterns are preserved by maintenance methyltransferases, which recognize hemimethylated DNA and methylate the newly synthesized strand.
After replication, only the parental strand is methylated, resulting in hemimethylated DNA.
Maintenance methyltransferase restores full methylation to the daughter DNA.
Histone Modification
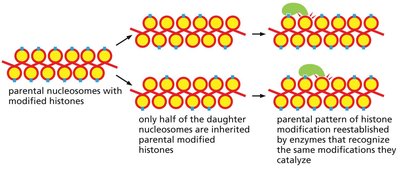
Histone proteins can be chemically modified (e.g., acetylation, methylation, phosphorylation), affecting chromatin structure and gene expression. These modifications can be inherited by daughter chromosomes during cell division.
Each daughter chromosome inherits about half of its parent’s modified histones.
Enzymes recognize and reestablish parental histone modification patterns on new histones.
Noncoding RNAs (ncRNAs) and lncRNAs
Long noncoding RNAs (lncRNAs) are RNA molecules that do not code for proteins but play important roles in regulating gene expression by interacting with chromatin-modifying enzymes.
lncRNAs share features with mRNAs (5′ cap, 3′ poly-A tail, splicing) but lack coding potential.
They can recruit chromatin modifiers to specific genomic loci, influencing gene expression.
Epigenetics and Monoallelic Gene Expression
Genomic Imprinting
Genomic imprinting is an epigenetic phenomenon where certain genes are expressed in a parent-of-origin-specific manner. This involves the silencing of one allele (either maternal or paternal) through DNA methylation and other epigenetic marks.
Most imprinted genes are located on autosomes and are critical for embryonic development.
Imprinting does not alter the DNA sequence but affects gene expression by adding chemical tags during gamete formation.
Imprinting is an exception to standard Mendelian inheritance.

Example: Igf2 Gene in Mice
The Igf2 gene is only expressed from the paternal allele; the maternal allele is silenced by methylation.
Imprinting disorders can result from abnormal methylation patterns, such as Beckwith–Weidemann syndrome (overgrowth) and Prader–Willi/Angelman syndromes (chromosome 15q11-q13).
Random Inactivation of the X Chromosome
In female mammals, one of the two X chromosomes is randomly inactivated in each cell during early embryonic development to balance gene dosage between males and females. The inactive X condenses into a Barr body and remains inactive in all descendant cells.
X inactivation is mediated by the XIST gene, which produces a long noncoding RNA that coats the inactive X chromosome and suppresses transcription.
Females heterozygous for X-linked genes are mosaics for those traits.
Epigenome and Phenotypic Variation
Genome vs. Epigenome
The genome is the complete set of genetic instructions in an organism, while the epigenome refers to the set of chemical modifications to DNA and histones present in a cell at a specific time. The epigenome can vary between cell types and over time, influencing gene expression and phenotype.
Monozygotic twins have identical genomes but can have different epigenomes due to environmental influences, leading to phenotypic differences.
The phenotype of an organism results from interactions between the genome and the epigenome.
Summary Table: Key Epigenetic Mechanisms
Mechanism | Description | Effect on Gene Expression |
|---|---|---|
DNA Methylation | Addition of methyl groups to cytosine in CpG islands | Gene silencing |
Histone Modification | Chemical modification of histone tails (e.g., acetylation, methylation) | Activation or repression, depending on modification |
Noncoding RNAs | lncRNAs and other ncRNAs recruit chromatin modifiers | Gene activation or repression |
Genomic Imprinting | Parent-of-origin-specific gene silencing | Monoallelic expression |
X Inactivation | Random inactivation of one X chromosome in females | Dosage compensation |
