 Back
BackEukaryotic DNA Replication and Telomere Biology
Study Guide - Smart Notes

Eukaryotic DNA Replication
Overview of Eukaryotic DNA Replication
Eukaryotic DNA replication is a fundamental process required for cell division and inheritance. While sharing many similarities with bacterial DNA replication, eukaryotic replication is more complex due to the presence of large linear chromosomes, tightly packed chromatin, and intricate cell cycle regulation.
Large linear chromosomes: Eukaryotes possess chromosomes that are much larger and linear, necessitating multiple origins of replication.
Chromatin structure: DNA is tightly packed within nucleosomes, affecting accessibility and regulation.
Cell cycle regulation: Replication is tightly controlled and coordinated with cell cycle phases.
Multiple Origins of Replication
To efficiently replicate their large genomes, eukaryotes utilize multiple origins of replication. Replication proceeds bidirectionally from these origins, forming replication bubbles that eventually merge.
Huberman and Riggs (1968): Provided evidence for multiple origins in eukaryotic chromosomes.
Replication bubbles: Multiple bubbles form and merge to complete chromosome replication.

Eukaryotic Origins of Replication
Origins of replication in simple eukaryotes, such as Saccharomyces cerevisiae, are termed ARS elements (Autonomously Replicating Sequence). These are short, AT-rich DNA sequences. In more complex eukaryotes, origins are often defined by chromatin structure rather than DNA sequence.
ARS elements: About 50 bp, high A/T content, consensus sequence ATTTAT(A or G)TTTA.
Complex eukaryotes: Origins are less defined, often associated with chromatin features.
Assembly of the Prereplication Complex (preRC)
Replication begins with the assembly of the prereplication complex (preRC) at origins. The Origin Recognition Complex (ORC) is a six-subunit initiator, followed by binding of MCM helicase, which completes DNA replication licensing.
ORC: Recognizes and binds origins, recruiting other factors.
MCM helicase: Essential for unwinding DNA and licensing replication.
Eukaryotic DNA Polymerases
Mammalian cells contain over a dozen DNA polymerases, but four are primarily responsible for DNA replication:
DNA pol α: Associates with primase to synthesize RNA-DNA hybrid primers.
DNA pol ε: Elongates the leading strand.
DNA pol δ: Elongates the lagging strand.
DNA pol γ: Replicates mitochondrial DNA.
Polymerase Switch and Strand Elongation
After primer synthesis by DNA pol α/primase, a polymerase switch occurs, replacing α with ε or δ for strand elongation. DNA pol ε is used for the leading strand, while DNA pol δ is used for the lagging strand.
Polymerase switch: Essential for processive DNA synthesis.
Leading strand: Continuous synthesis by DNA pol ε.
Lagging strand: Discontinuous synthesis by DNA pol δ.
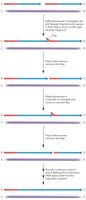
Flap Endonuclease Removes RNA Primers
Unlike prokaryotes, eukaryotic lagging strand synthesis involves flap endonuclease to remove RNA primers. DNA pol δ displaces the primer, creating a flap, which is then cleaved by flap endonuclease or Dna2 nuclease/helicase if the flap is too long.
Flap endonuclease: Removes short flaps of RNA primer.
Dna2 nuclease/helicase: Cleaves long flaps into short flaps for removal.
Telomeres and DNA Replication
Structure and Function of Telomeres
Telomeres are specialized structures at the ends of linear chromosomes, consisting of repetitive DNA sequences and associated proteins. They protect chromosome ends from degradation and fusion, and play a critical role in genome stability.
Telomeric DNA: Repetitive sequences (e.g., TTAGGG in humans).
Telomere-associated proteins: Form a protective cap and regulate telomere maintenance.
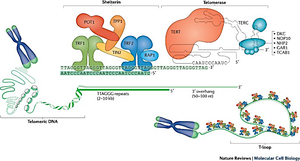
Telomeric DNA Sequences
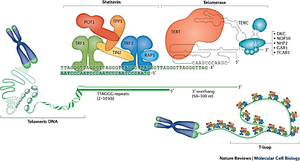
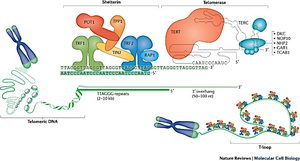
Telomeric DNA consists of tandem repeats and a single-stranded 3' overhang. The shelterin complex binds telomeric DNA, preventing unwanted DNA damage responses and maintaining telomere integrity.
Shelterin complex: Includes proteins such as TRF1, TRF2, POT1, TIN2, RAP1, and TPP1.
3' overhang: Essential for telomere structure and function.
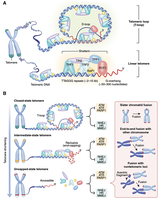
Telomere Structure and Chromosome End Protection
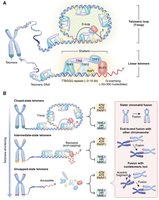
Telomeres form a T-loop structure, which hides the chromosome end and prevents it from being recognized as DNA damage. Different states of telomere structure influence chromosome stability and cellular outcomes.
T-loop: Protects chromosome ends from degradation and fusion.
Linear telomeres: Exposed ends can trigger DNA damage responses.
Telomere dysfunction: Leads to chromosome fusion and genomic instability.
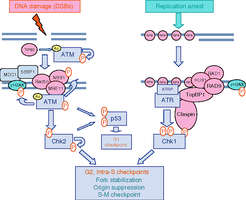
DNA Damage Response at Telomeres
DNA Damage Response Pathways
Telomeres are monitored by DNA damage response pathways, which are activated upon telomere dysfunction or replication stress. Key proteins such as ATM and ATR kinases initiate checkpoint signaling to maintain genome integrity.
ATM/ATR kinases: Detect DNA damage and activate cell cycle checkpoints.
p53: Induces cell cycle arrest or apoptosis in response to damage.
Chk1/Chk2: Mediate checkpoint signaling and repair processes.
Telomerase and Telomere Maintenance
Telomerase Structure and Function
Telomerase is a ribonucleoprotein enzyme that extends telomeres by adding repetitive DNA sequences to chromosome ends. It is active in germ cells, stem cells, and most cancer cells, but inactive in most somatic cells.
Telomerase components: Includes TERT (reverse transcriptase) and TERC (RNA template).
Function: Counteracts telomere shortening during DNA replication.
Telomere Length, Cellular Senescence, and Cancer
Telomere Shortening and Cellular Senescence
Telomeres shorten with each cell division due to the end-replication problem. When telomeres become critically short, cells enter senescence and lose their ability to divide, a phenomenon known as the Hayflick effect.
Telomere length: About 8,000 bp at birth, can shorten to 1,500 bp in elderly individuals.
Senescence: Triggered by short telomeres, preventing further cell division.
Hayflick effect: Cells have a limited number of divisions before senescence.
Telomerase Activation and Cancer
In cancer cells, telomerase is often reactivated, allowing cells to maintain telomere length and proliferate indefinitely. This bypasses senescence and contributes to tumorigenesis.
Telomerase activation: Enables unlimited cell division in cancer and germ cells.
Telomere maintenance: Essential for cancer cell immortality.
Summary Table: Key Features of Eukaryotic DNA Replication and Telomere Biology
Feature | Eukaryotes | Bacteria |
|---|---|---|
Chromosome Structure | Linear, large, chromatin-packed | Circular, small, nucleoid |
Origins of Replication | Multiple, ARS elements, chromatin-defined | Single, defined DNA sequence (oriC) |
DNA Polymerases | α, δ, ε, γ (many others) | Pol III (main), Pol I (primer removal) |
Primer Removal | Flap endonuclease, Dna2 | DNA Pol I |
Telomeres | Present, repetitive DNA, shelterin complex | Absent |
Telomerase | Active in germ/cancer cells | Not required |
Additional info: Academic context was added to clarify the roles of telomerase, shelterin, and DNA damage response pathways, as well as to provide a summary table for comparison.
