 Back
BackEukaryotic Gene Expression and Epigenetic Regulation: Mini-Textbook Study Notes
Study Guide - Smart Notes

Ch 17-18: Eukaryotic Gene Expression
Overview of Eukaryotic Gene Regulation
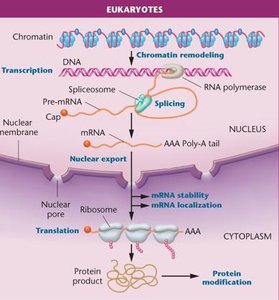
Eukaryotic gene expression is regulated at multiple levels, including transcription, RNA processing, translation, and post-translational modification. The complexity of eukaryotic cells allows for fine-tuned control of gene expression, enabling cells to respond to environmental cues and developmental signals.
Transcriptional Regulation: Chromatin remodeling, histone modification, and DNA methylation affect accessibility of genes for transcription.
Post-transcriptional Regulation: Includes alternative splicing, RNA editing, mRNA stability, localization, and translation initiation.
Post-translational Regulation: Modifications such as phosphorylation, acetylation, methylation, glycosylation, and ubiquitination alter protein function and stability.
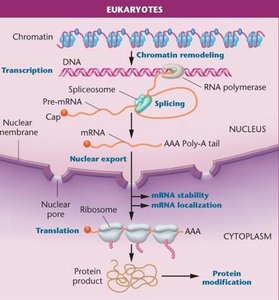
Transcriptional Regulation in Eukaryotes
Gene expression in eukaryotes is influenced by chromatin modifications and the action of transcription factors. Chromatin structure, histone modifications, and DNA methylation play key roles in determining which genes are accessible for transcription.
Histone Modification: Acetylation and methylation of histones can either promote or repress gene expression.
DNA Methylation: Addition of methyl groups to cytosine bases in CpG islands typically silences gene expression.
Transcription Factors: Trans-acting factors bind to cis-acting sites (promoters, enhancers) to regulate transcription initiation.
Post-Transcriptional Regulation in Eukaryotes
Alternative Splicing
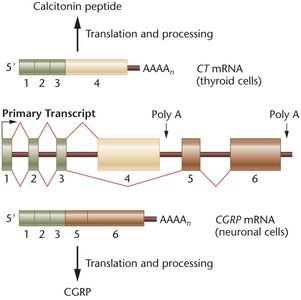
Alternative splicing is a process by which different combinations of exons are joined together to produce multiple mRNA variants from a single gene, greatly expanding the diversity of proteins encoded by the genome.
Definition: The selective inclusion or exclusion of exons during mRNA processing.
Impact: One gene can produce numerous proteins with distinct functions.
Example: The Drosophila Dscam gene can generate up to 38,016 different protein isoforms through alternative splicing.

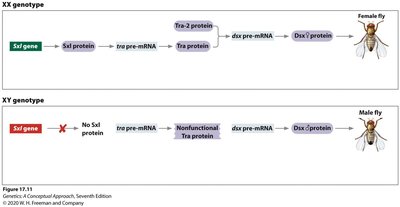
Sex Determination in Drosophila: Alternative Splicing Pathway
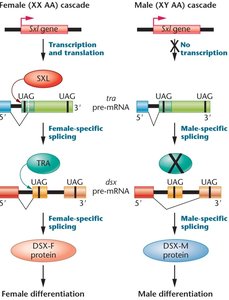
Sex determination in Drosophila is regulated by alternative splicing of key genes: Sex lethal (Sxl), Transformer (tra), and Doublesex (dsx). Sxl acts as a master regulator, controlling the splicing of tra and dsx pre-mRNAs to produce sex-specific proteins.
Sxl: RNA binding protein, ON in females (XX), OFF in males (XY).
tra: Functional in females, nonfunctional in males due to premature stop codon.
dsx: Spliced differently in males and females, leading to sex-specific development.
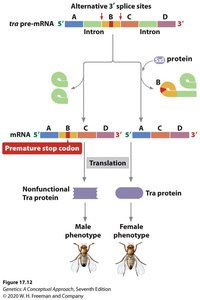
RNA Editing
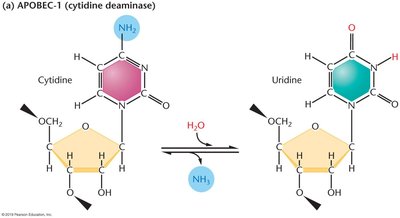
RNA editing is a post-transcriptional process in which the nucleotide sequence of an RNA molecule is altered after synthesis, resulting in a sequence that differs from the DNA template. This can change the encoded protein.
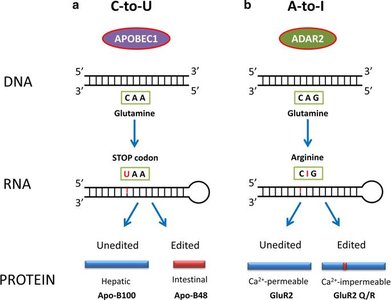
Types: Base changes (e.g., C-to-U, A-to-I), insertions, deletions.
Example: Apolipoprotein B (ApoB) mRNA is edited in the intestine to produce a truncated protein (ApoB-48), while the liver produces full-length ApoB-100.
Regulation by mRNA Stability and Degradation
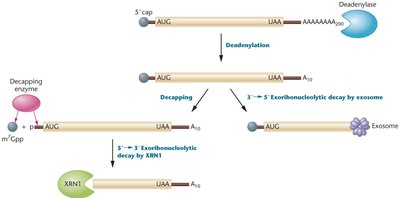
The stability and degradation of mRNA molecules in the cytoplasm are crucial for controlling protein synthesis. Eukaryotic mRNAs are generally more stable than prokaryotic mRNAs, and their half-life is regulated by sequences in the untranslated regions (UTRs).
Degradation Pathways: Deadenylation-dependent decay, nonsense-mediated mRNA decay (NMD).
Key Enzymes: Deadenylase shortens poly(A) tail, decapping enzymes remove 5' cap, exoribonucleases degrade mRNA.
Regulatory Elements: Poly-A binding proteins stabilize mRNA; sequences in 3' UTR influence degradation.
Translational Regulation
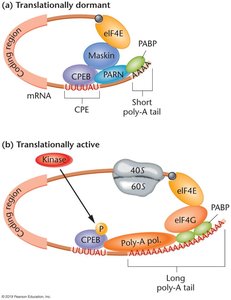
Translation initiation can be regulated to keep mRNAs dormant or activate them in response to cellular signals. Regulatory elements in the 3' UTR, such as the cytoplasmic polyadenylation element (CPE), are recognized by binding proteins that control translation.
CPEB: Cytoplasmic polyadenylation element binding protein recruits factors to keep mRNA translationally dormant.
Activation: Phosphorylation of CPEB leads to polyadenylation and translation activation.
RNA Interference (RNAi) and Noncoding RNAs
RNA interference is a mechanism by which noncoding RNAs (ncRNAs) regulate gene expression by targeting mRNAs for degradation or translational inhibition. Small interfering RNAs (siRNAs) and microRNAs (miRNAs) are processed from double-stranded precursors and incorporated into the RNA-induced silencing complex (RISC).
siRNA: Perfect base pairing with target mRNA leads to degradation.
miRNA: Imperfect pairing often results in translational repression.
Applications: RNAi is used in biotechnology and medicine to silence specific genes without altering DNA.
*Additional info: RNAi introduces no mutations into the organism’s DNA, making it a powerful tool for gene silencing and therapeutic applications.*
Post-Translational Regulation
Post-Translational Modifications (PTMs)
Proteins can be modified after translation to regulate their activity, stability, localization, and interactions. Common PTMs include phosphorylation, acetylation, methylation, glycosylation, and ubiquitination.
Phosphorylation: Regulates enzyme activity and signaling pathways.
Acetylation: Affects gene expression, especially histone acetylation.
Methylation: Influences DNA binding and protein interactions.
Glycosylation: Essential for protein folding and cell recognition.
Ubiquitination: Marks proteins for degradation by the proteasome.
Epigenetics
Definition and Mechanisms
Epigenetics refers to heritable changes in gene expression that do not involve alterations in the DNA sequence. These changes affect how cells read genes and can be influenced by environmental factors.
DNA Methylation: Addition of methyl groups to cytosine bases, typically silencing gene expression.
Histone Modification: Acetylation, methylation, and phosphorylation alter chromatin structure and gene accessibility.
Noncoding RNA: Regulates gene expression post-transcriptionally.
DNA Methylation and CpG Islands
Methylation occurs on cytosine bases adjacent to guanine (CpG dinucleotides), clustered in CpG islands near promoter regions. Methylation of CpG islands is usually negatively correlated with gene activity.
Unmethylated CpG Islands: Associated with active transcription of essential and cell-specific genes.
Methylated CpG Islands: Associated with transcriptional silencing.
Epigenetic Traits and Environmental Influence
Epigenetic traits are stable, heritable phenotypes resulting from changes in gene expression without DNA sequence alterations. Environmental factors such as nutrition, chemicals, and stress can induce epigenetic changes.
Example: The Dutch Hunger Winter showed that prenatal famine exposure led to increased risk of metabolic and neuropsychiatric disorders in offspring.
Honeybee Development: Royal jelly suppresses DNA methyltransferase, leading to queen development.
Heritability of Epigenetic Traits
Epigenetic changes induced by environmental factors can be inherited across generations. For example, stress-induced changes in DNA methylation and histone acetylation affect behavior in rats and are passed to offspring.
High Maternal Nurturing: Leads to increased serotonin, histone acetylation, decreased DNA methylation, and higher glucocorticoid receptor expression, resulting in better stress adaptation.
Low Maternal Nurturing: Results in decreased serotonin, decreased histone acetylation, increased DNA methylation, and lower glucocorticoid receptor expression, leading to poor stress adaptation.
Summary Table: Major Mechanisms of Eukaryotic Gene Regulation
Level | Mechanism | Key Features |
|---|---|---|
Transcriptional | Chromatin remodeling, histone modification, DNA methylation | Controls gene accessibility |
Post-transcriptional | Alternative splicing, RNA editing, mRNA stability, RNA interference | Expands protein diversity, regulates mRNA fate |
Translational | Translation initiation regulation | Controls timing and amount of protein synthesis |
Post-translational | Protein modification (PTMs), ubiquitin-mediated degradation | Regulates protein function and turnover |
Epigenetic | DNA methylation, histone modification, ncRNA | Heritable changes in gene expression |
