 Back
BackEukaryotic Gene Regulation: Promoters, Transcription Factors, and Alternative Splicing
Study Guide - Smart Notes

Regulation of Eukaryotic Gene Expression
Promoters and Their Diversity
Promoters are specific DNA sequences that serve as recognition sites for the transcription machinery and are essential for the initiation of transcription. They are located immediately adjacent to regulatory genes and are critical for determining where and how efficiently transcription begins.
Core Promoter: Determines the accurate initiation of transcription, typically containing the transcription start site (+1).
Proximal-Promoter Elements: Located upstream of the core promoter, these elements modulate the efficiency of basal transcription levels.
There is significant diversity in eukaryotic promoter structure and function:
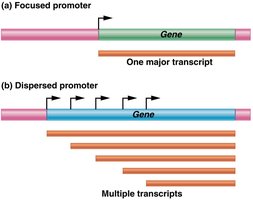
Focused Promoters: Direct transcription initiation at a single start site, producing one major transcript. Common in lower eukaryotes.
Dispersed Promoters: Initiate transcription at multiple weak start sites, resulting in multiple transcripts. More common in higher eukaryotes.
Promoter Structure and Elements
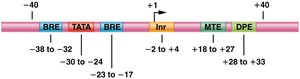
Promoters are composed of several DNA sequence elements, each with specific roles in transcription initiation:
Initiator (Inr): Located around -2 to +4, with the consensus sequence YYANA/TYY (Y = pyrimidine). The first base transcribed is usually an adenine at +1.
TATA Box: Located around -30, crucial for the binding of transcription factors.
TFIIB Recognition Element (BRE): Found near the TATA box, recognized by TFIIB.
Downstream Promoter Element (DPE): Located downstream of the start site, enhances transcription in promoters lacking a TATA box.
Motif Ten Element (MTE): Another downstream element that can enhance transcription.
Proximal-Promoter Elements
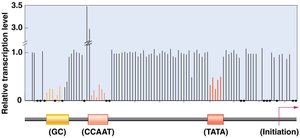
Proximal-promoter elements are DNA sequences located upstream of the core promoter (TATA and BRE motifs) and are important for enhancing basal transcription levels. Common examples include the CAAT box and GC box. Mutations in these regions can significantly affect the amount of mRNA produced.
Enhancers and Silencers
Enhancers and silencers are cis-acting regulatory elements that modulate the transcription of eukaryotic genes:
Enhancers: Can be located upstream, downstream, or within a gene. They increase the maximum level of transcription by facilitating the binding of transcription factors.
Silencers: Repress the level of transcription initiation, often by recruiting repressor proteins.
Transcription Factors
Transcription factors are proteins that regulate gene expression by binding to specific DNA sequences (cis-acting sites). They can act as activators (increasing transcription) or repressors (decreasing transcription). Multiple transcription factors can bind to different enhancers and promoter elements, fine-tuning the level of transcription initiation.
Functional Domains: Transcription factors typically have two main domains:
DNA-binding domain: Binds to specific DNA sequences in regulatory regions.
Trans-activating domain: Interacts with other transcription factors or RNA polymerase to activate or repress transcription.
Human Metallothionein IIA Gene (hMTIIA) as a Model
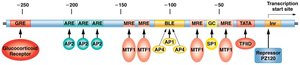
The hMTIIA gene is an example of transcriptional regulation involving promoters, enhancers, and transcription factors. Its protein product binds heavy metals, protecting cells from toxicity and oxidative stress. Expression is low under normal conditions but increases dramatically in response to heavy metals.
Formation of the RNA Polymerase II Initiation Complex
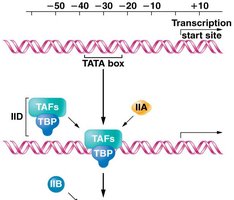
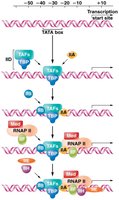
Transcription initiation by RNA Polymerase II (RNAP II) requires the assembly of a pre-initiation complex (PIC) at the promoter. General transcription factors (GTFs) are required for both basal and enhanced transcription. The assembly occurs in a specific order, providing a platform for RNAP II to recognize the transcription start site.
TFIID: Contains the TATA-binding protein (TBP) and TBP-associated factors (TAFs).
Additional factors (TFIIA, TFIIB, TFIIE, TFIIF, TFIIH) are sequentially recruited.
Mechanisms of Transcription Activation and Repression
Transcription activators and repressors can alter the rate of transcription by facilitating or hindering the assembly of the transcription machinery. DNA looping allows distant regulatory elements (such as enhancers) to interact with the promoter region, bringing activators, repressors, and general transcription factors into close proximity.
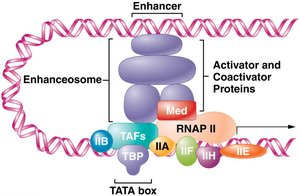
Coactivators and the Enhanceosome
Coactivators are proteins that interact with activators and enable them to contact promoter-bound factors. The complex formed by coactivators and activators is called the enhanceosome, which interacts with the transcription complex to enhance transcription. Repressor proteins at silencer elements can decrease the rate of PIC assembly and RNAP II release.
Common DNA-Binding Motifs in Transcription Factors
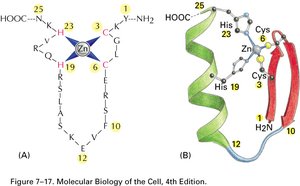
Transcription factors often contain conserved DNA-binding motifs that facilitate their interaction with DNA:
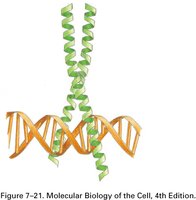
Zinc Finger: Characterized by coordination of a zinc ion, stabilizing the fold of the protein domain.
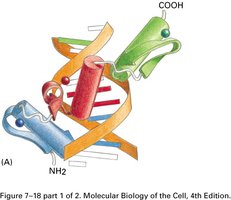
Helix-Loop-Helix: Consists of two α-helices connected by a loop, facilitating dimerization and DNA binding.
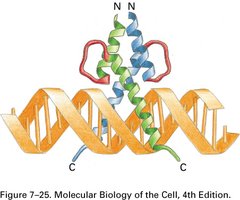
Leucine Zipper: Contains leucine residues at every seventh position, allowing dimerization and DNA binding.
Alternative Splicing and Post-Transcriptional Regulation
Alternative Splicing
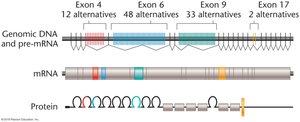
Alternative splicing is a post-transcriptional mechanism that allows a single gene to produce multiple mRNA variants, greatly increasing proteomic diversity. This process involves the selective inclusion or exclusion of exons during mRNA processing.
Cassette Exons: Exons that may be included or skipped in the final mRNA.
Alternative Splice Sites: Use of different 5' or 3' splice sites within an exon.
Intron Retention: Introns are retained in the mature mRNA, potentially producing novel isoforms.
Mutually Exclusive Exons: Only one exon from a cluster is included in the mature mRNA.
Alternative Promoters: Different transcription start sites produce mRNAs with different 5' exons.
Alternative Polyadenylation: Use of different polyadenylation signals results in mRNAs with different 3' ends.
Regulation of Alternative Splicing
Alternative splicing is regulated by cis-acting sequences (splicing enhancers and silencers) and trans-acting proteins:
SR Proteins: Bind to splicing enhancers and promote splicing by recruiting spliceosome components.
Heterogeneous Nuclear Ribonucleoproteins (hnRNPs): Bind to splicing silencers and inhibit splicing.
Biological Significance of Alternative Splicing
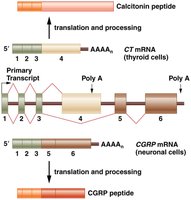
Alternative splicing allows for the generation of protein isoforms with different, sometimes tissue-specific, functions. For example, the calcitonin gene produces two distinct peptides—calcitonin in the thyroid and CGRP in neurons—through alternative splicing.
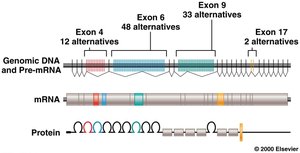
The Dscam gene in Drosophila is another example, capable of producing over 38,000 unique isoforms through alternative splicing, which is critical for neural development.
Control of mRNA Stability
The steady-state level of mRNA in a cell is determined by the balance between its synthesis and degradation. mRNA stability is regulated by:
Shortening of the poly-A tail by exonucleases
Removal of the 5' cap by decapping enzymes
Endonucleolytic cleavage within the mRNA
Translational and Post-Translational Regulation: The p53 Protein
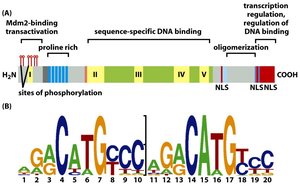
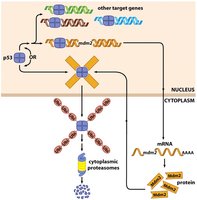
The p53 protein is a transcription factor whose levels are tightly regulated. Under normal conditions, p53 is rapidly degraded via ubiquitin-mediated proteasomal degradation. In response to DNA damage or stress, p53 is stabilized and activates the transcription of genes involved in cell cycle arrest, DNA repair, or apoptosis.
Ubiquitin: A small protein that tags other proteins for degradation by the proteasome.
Mdm2: An E3 ubiquitin ligase that targets p53 for degradation.
Summary Table: Types of Alternative Splicing
Type of Alternative Splicing | pre-mRNA | mRNA Spliceforms |
|---|---|---|
Cassette exons | Exon may be included or skipped | Two possible mRNAs: with or without the exon |
Alternative 5' (or 3') splice site | Different splice sites within an exon | mRNAs with different exon boundaries |
Intron retention | Intron may be retained | mRNA with or without the intron |
Mutually exclusive exons | Cluster of exons, only one included | mRNAs with different exons from the cluster |
Alternative promoters | Multiple transcription start sites | mRNAs with different 5' exons |
Alternative polyadenylation | Multiple polyadenylation signals | mRNAs with different 3' ends |
