 Back
BackEukaryotic Transcription and RNA Processing
Study Guide - Smart Notes

Chapter 12: Eukaryotic Transcription and RNA Editing
Introduction
This chapter explores the mechanisms of eukaryotic transcription and the post-transcriptional modifications that are essential for the maturation of RNA. Key processes include transcription initiation, elongation, termination, 5’ capping, polyadenylation, RNA splicing, and RNA editing. Understanding these processes is fundamental for comprehending gene expression regulation in eukaryotes.
Eukaryotic RNA Polymerases
Types and Functions
Eukaryotic cells possess three main nuclear RNA polymerases, each responsible for transcribing different classes of genes:
RNA Polymerase I: Transcribes all rRNA genes except 5S rRNA.
RNA Polymerase II: Transcribes all protein-coding genes (mRNAs) and some snRNA genes required for splicing.
RNA Polymerase III: Transcribes all tRNA genes, the 5S rRNA gene, and microRNA genes.
Each polymerase recognizes distinct promoter elements and requires specific transcription factors for initiation.
Transcription Initiation in Eukaryotes
Promoter Structure
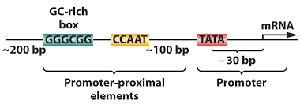
The promoter is a DNA sequence upstream of the gene that serves as the binding site for transcription machinery. Key elements include the TATA box, CAAT box, and GC-rich regions.
TATA Box: Located ~30 bp upstream of the transcription start site; recognized by TATA-binding protein (TBP).
CAAT Box and GC Box: Promoter-proximal elements that enhance transcription efficiency.
Assembly of the Pre-Initiation Complex
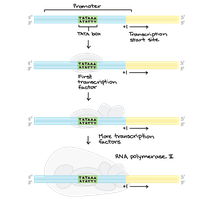
Transcription initiation involves the sequential binding of general transcription factors (GTFs) and RNA polymerase II to the promoter:
TFIID (containing TBP) binds the TATA box.
Other GTFs (TFIIA, TFIIB, TFIIE, TFIIF, TFIIH) assemble, forming the pre-initiation complex.
RNA polymerase II is recruited and positioned at the transcription start site.
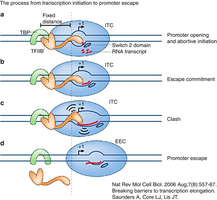
Initiation and Promoter Escape
After the pre-initiation complex forms, RNA polymerase II synthesizes short RNA transcripts. TFIIB can cause abortive initiation, where short transcripts are released until the polymerase escapes the promoter and enters productive elongation.
Transcription Elongation
Process and Regulation
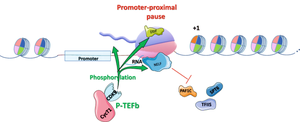
During elongation, RNA polymerase II moves along the DNA, synthesizing RNA in the 5’ to 3’ direction. Elongation is regulated by pausing and the action of elongation factors.
Elongation Pausing: RNA polymerase may pause shortly after initiation, regulated by factors such as NELF and DSIF.
Elongation Factors: P-TEFb phosphorylates components of the paused complex, allowing productive elongation.
Transcription Termination and Polyadenylation
Termination Mechanism
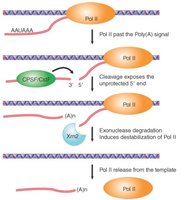
Termination of transcription by RNA polymerase II involves recognition of the polyadenylation signal (AAUAAA) in the nascent RNA. Several protein factors are involved:
CFI and CFII: Cleavage factors that cut the RNA downstream of the polyadenylation signal.
CPSF: Cleavage and Polyadenylation Specificity Factor, binds the polyadenylation signal.
CstF: Cleavage Stimulation Factor, interacts with CPSF to promote cleavage.
PAP: Poly(A) polymerase adds a poly(A) tail to the 3’ end of the RNA.
5’ Capping of mRNA
Mechanism and Purpose
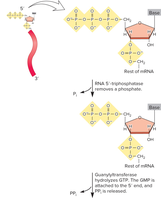
Shortly after transcription initiation, a 7-methylguanosine cap is added to the 5’ end of the nascent mRNA. This process involves:
Removal of the terminal phosphate from the 5’ triphosphate end by RNA 5’-triphosphatase.
Addition of GMP by guanylyltransferase.
Methylation of the guanine base by methyltransferase.
The 5’ cap protects mRNA from degradation, facilitates ribosome binding during translation, and is involved in nuclear export.
RNA Splicing
Removal of Introns
RNA splicing is the process by which non-coding introns are removed from pre-mRNA and exons are joined to form mature mRNA. The spliceosome, a complex of snRNAs and proteins, catalyzes this process.
Splice Sites: Conserved sequences at the exon-intron boundaries guide the spliceosome.
Lariat Formation: The intron is excised as a lariat structure and degraded.

RNA Editing
Post-Transcriptional Base Modification
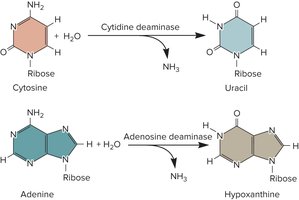
RNA editing refers to changes in the nucleotide sequence of an RNA molecule after it has been synthesized. This can involve:
Addition or deletion of specific bases.
Chemical modification of bases, such as deamination (e.g., cytosine to uracil, adenosine to inosine).
RNA editing increases transcriptome diversity and can affect protein function.
Summary Table: Eukaryotic RNA Polymerases
Polymerase | Transcribes |
|---|---|
RNA pol I | rRNA genes (except 5S rRNA) |
RNA pol II | Protein-coding genes (mRNA), some snRNA genes |
RNA pol III | tRNA genes, 5S rRNA gene, microRNA genes |
Key Terms
Promoter: DNA sequence where transcription machinery assembles.
Transcription Factors: Proteins required for the initiation of transcription.
Spliceosome: Complex responsible for removing introns from pre-mRNA.
Polyadenylation: Addition of a poly(A) tail to the 3’ end of mRNA.
RNA Editing: Post-transcriptional modification of RNA sequence.
