 Back
BackForward and Reverse Genetics: Strategies for Gene Function Analysis
Study Guide - Smart Notes

Forward and Reverse Genetics
Introduction to Forward and Reverse Genetics
Forward and reverse genetics are two complementary approaches used to study gene function and the relationship between genotype and phenotype. Forward genetics starts with a phenotype and seeks to identify the underlying gene, while reverse genetics begins with a gene of interest and investigates the resulting phenotype when that gene is altered.
Forward Genetics: Identifies genes responsible for a particular phenotype.
Reverse Genetics: Determines the function of a gene by analyzing the phenotypic effects of specific gene modifications.
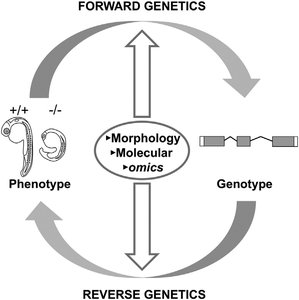
Comparison of Forward and Reverse Genetics
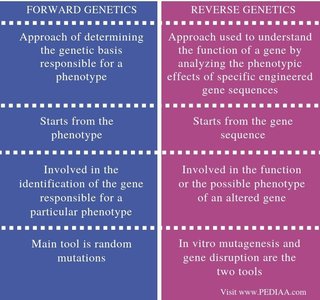
The following table summarizes the main differences between forward and reverse genetics:
Forward Genetics | Reverse Genetics |
|---|---|
Approach of determining the genetic basis responsible for a phenotype | Approach used to understand the function of a gene by analyzing the phenotypic effects of specific engineered gene sequences |
Starts from the phenotype | Starts from the gene sequence |
Involved in the identification of the gene responsible for a particular phenotype | Involved in the function or the possible phenotype of an altered gene |
Main tool is random mutations | In vitro mutagenesis and gene disruption are the two tools |

Forward Genetics
Basic Strategy and Model Organisms
Forward genetics involves mutagenizing a population, screening for abnormal phenotypes, and identifying the gene responsible for the phenotype. Model organisms such as Drosophila melanogaster (fruit fly), Mus musculus (mouse), Saccharomyces cerevisiae (yeast), and Arabidopsis thaliana (plant) are commonly used due to their short generation times, ease of breeding, and genetic tractability.
Step 1: Mutagenize a large population (using chemicals, radiation, or insertional elements).
Step 2: Screen for individuals with abnormal phenotypes.
Step 3: Identify the gene or sequence responsible for the phenotype.
Step 4: Infer the role of the gene product.
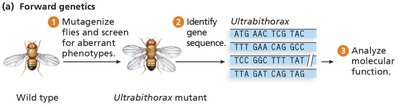
Mutagens Used in Forward Genetics
Mutagens are agents that induce mutations in the genome. Common mutagens include:
Chemical mutagens: e.g., Ethyl methanesulfonate (EMS), which causes point mutations.
Radiation: e.g., X-rays, gamma rays, which cause chromosomal rearrangements.
Insertional elements: e.g., transposons, which disrupt gene function by insertion.
Screening for Dominant and Recessive Mutations
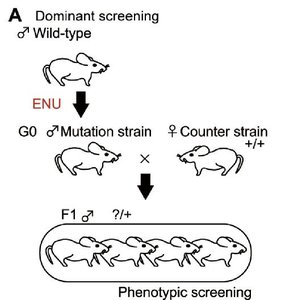
Dominant mutations are typically gain-of-function and can be detected in the first generation after mutagenesis. Recessive mutations, often loss-of-function, require breeding through multiple generations to reveal homozygous mutants.
Dominant screen: Mutagenize sperm, cross with wild-type, screen F1 for phenotype.
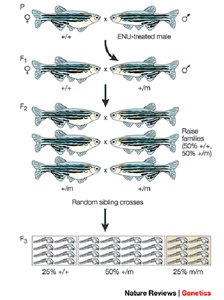
Recessive screen: Mutagenize sperm, cross with wild-type, breed F1 to F2, intercross F2 to produce F3, screen F3 for homozygous mutants.

Linking Phenotype to Genotype
Once a mutant phenotype is identified, several methods are used to map and identify the causative gene:
Linkage mapping: Uses known genetic markers to locate the gene responsible for the phenotype.
Wild-type complementation: Introduces wild-type DNA to mutant cells to see if the phenotype is rescued.
Genome sequencing: Compares the genomes of mutants and wild-type individuals to identify causative mutations.
Reverse Genetics
Basic Strategy and Approaches
Reverse genetics starts with a known gene and investigates the effects of its alteration on the organism's phenotype. This approach has become increasingly feasible with the availability of whole-genome sequences.
Step 1: Select a gene of interest.
Step 2: Mutate or disrupt the gene (using homologous recombination, insertional mutagenesis, RNAi, or CRISPR).
Step 3: Characterize the resulting phenotype.
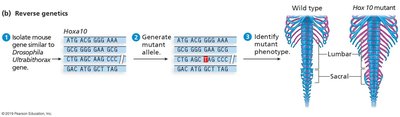
Reverse Genetics Techniques
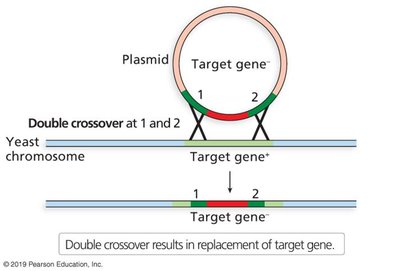
Homologous recombination: Targeted gene disruption or replacement using engineered DNA fragments. Common in yeast and mice.
Insertional mutagenesis: Random insertion of transposons or T-DNA to disrupt gene function.
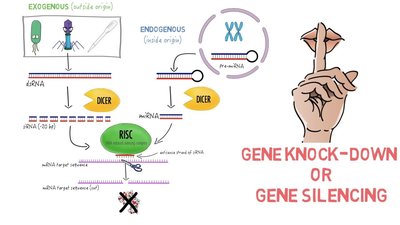
RNA interference (RNAi): Uses small RNA molecules to silence gene expression by degrading mRNA or blocking translation.
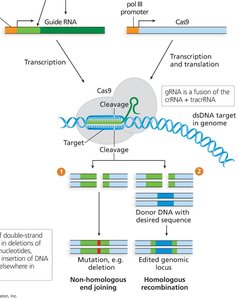
CRISPR-Cas9: A precise genome editing tool that uses guide RNA and Cas9 endonuclease to introduce targeted mutations.
TILLING (Targeting Induced Local Lesions IN Genomes): Screens for point mutations in specific genes in a mutagenized population.
Transgenes and Enhancer Trapping
Transgenes
A transgene is a gene that has been artificially introduced into the genome of another organism. Transgenic organisms are used to study gene function, model diseases, or produce useful products.
Reporter genes: Allow visualization of gene expression (e.g., lacZ, GFP).
Gain-of-function genes: Introduce and express foreign genes to study their effects.
Enhancer Trapping
Enhancer trapping uses transposable elements carrying a reporter gene and a strong promoter. When inserted near a gene of interest, the reporter is expressed whenever the endogenous gene is active, allowing researchers to study gene regulation and expression patterns.
Summary Table: Forward vs. Reverse Genetics
Aspect | Forward Genetics | Reverse Genetics |
|---|---|---|
Starting Point | Phenotype | Gene sequence |
Goal | Identify gene responsible for phenotype | Determine phenotype resulting from gene alteration |
Common Tools | Random mutagenesis, screening | Gene targeting, RNAi, CRISPR |
Applications | Gene discovery, functional genomics | Gene function analysis, disease modeling |
Take Home Message
Both forward and reverse genetics are essential for understanding gene function and the molecular basis of phenotypes. Advances in genome sequencing and gene editing technologies have greatly expanded the toolkit available to geneticists, enabling precise manipulation and analysis of genes in a wide range of organisms.
