Back
BackGene Expression and Regulation in Eukaryotes: Mechanisms and Epigenetic Control
Study Guide - Smart Notes

Gene Expression in Eukaryotes
Overview of Gene Regulation
Gene expression in eukaryotes is a highly regulated process that ensures genes are expressed at the right time, place, and amount. Regulation occurs at multiple steps, from transcription initiation to post-translational modification of proteins. This complexity allows for cell type specificity and adaptation to environmental signals.
Transcription Factors: Proteins that bind DNA to regulate transcription.
Activators and Repressors: Increase or decrease gene expression, respectively.
Epigenetics: Heritable changes in gene expression not involving changes to the DNA sequence.
Post-transcriptional Control: Includes RNA splicing and RNA interference (RNAi).
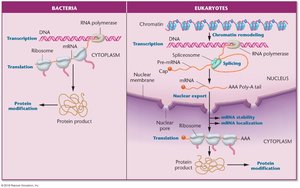
Regulation of Gene Expression: Prokaryotes vs. Eukaryotes
Key Differences
While both prokaryotes and eukaryotes regulate gene expression in response to environmental signals, eukaryotic regulation is more complex, involving multiple layers and cell-type specificity.
Prokaryotes: Regulation mainly at transcription initiation; genes are often organized in operons.
Eukaryotes: Regulation at multiple steps, including chromatin remodeling, transcription, RNA processing, export, translation, and protein modification.

Steps Regulated During Eukaryotic Gene Expression
Major Regulatory Points
Transcription Initiation
Transcript Processing (e.g., splicing, capping, polyadenylation)
Export from Nucleus
Translation of mRNA
Protein Localization
Protein Modification
Transcription Initiation in Eukaryotes
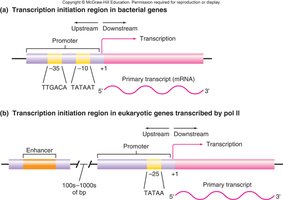
Promoters and Enhancers
Eukaryotic genes often contain enhancers that can be located thousands of base pairs away from the promoter. Enhancers are required for efficient transcription and can function regardless of their position or orientation relative to the gene.
Promoter: Region where RNA polymerase and basal transcription factors bind to initiate transcription.
Enhancer: DNA sequence that augments or represses gene expression, often by binding activators or repressors.
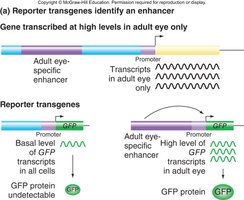
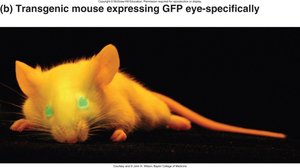
Reporter Constructs and Enhancer Identification
Enhancer regions can be identified using reporter constructs in model organisms or transgenic animals. For example, a gene encoding Green Fluorescent Protein (GFP) can be placed under the control of a suspected enhancer to visualize its activity.
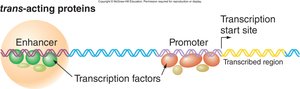
Transcription Factors: Activators and Repressors
Trans-acting Proteins
Transcription factors are sequence-specific DNA-binding proteins that regulate gene expression by binding to promoters and enhancers. They recruit other proteins to influence transcription.
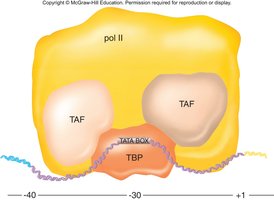
Basal Transcription Factors
Basal transcription factors are required for the assembly of the transcription initiation complex at the promoter. They include the TATA-binding protein (TBP) and TBP-associated factors (TAFs).
Transcriptional Activators
Activators bind to enhancer regions and are responsible for most of the variation in gene expression. They can interact directly or indirectly with basal transcription factors and recruit coactivators such as histone acetyltransferases (HATs) and chromatin remodeling complexes.

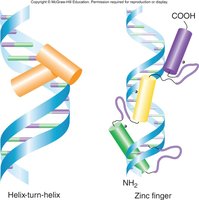
Structure of Activator Proteins
DNA Binding Domain: Recognizes specific DNA sequences.
Activation Domain: Interacts with other proteins to enhance transcription.
Dimerization Domain: Allows activators to form dimers, increasing binding specificity and strength.
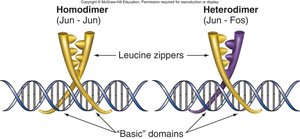
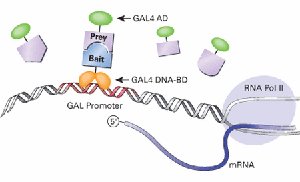
Coactivator Example: Yeast Two-Hybrid Screens
Yeast two-hybrid screens are used to identify protein-protein interactions by bringing together a DNA-binding domain and an activation domain, which activates a reporter gene if the proteins interact.
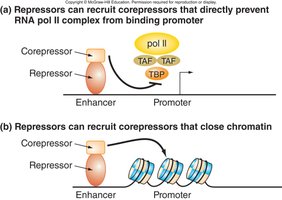
Transcriptional Repressors
Repressors bind to enhancer regions and decrease gene expression. In eukaryotes, they often act indirectly, for example by recruiting corepressors such as histone deacetylases (HDACs) or histone methyltransferases (HMTs), which modify chromatin to a closed state.
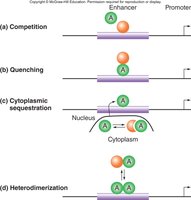
Indirect Repression Mechanisms
Repressors can interfere with activator function by overlapping binding sites, masking activation domains, preventing nuclear entry, or blocking dimerization.
Epigenetics and Gene Regulation
Epigenetic Mechanisms
Epigenetics refers to heritable changes in gene expression that do not involve changes to the DNA sequence. Major mechanisms include DNA methylation, histone modification, and noncoding RNAs.
DNA Methylation: Addition of methyl groups to cytosine residues in CpG dinucleotides, often leading to gene silencing.
Histone Modification: Alters chromatin structure and gene accessibility.
Noncoding RNA: Regulates gene expression at the transcriptional and post-transcriptional levels.
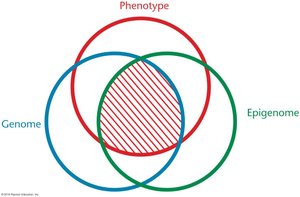
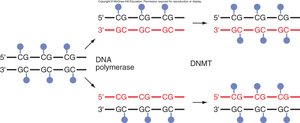
DNA Methylation and CpG Islands
Methylation of CpG islands in promoter regions is associated with transcriptional silencing. DNA methyltransferases (DNMTs) add methyl groups, and methylation patterns are heritable through cell division.
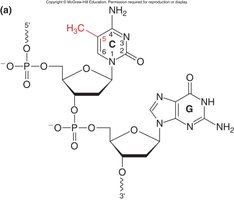
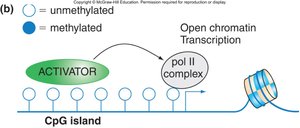
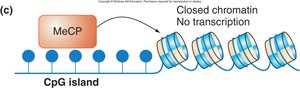
Heritability of CpG Methylation
Once established, DNA methylation patterns are maintained through DNA replication and cell division, contributing to stable gene silencing.
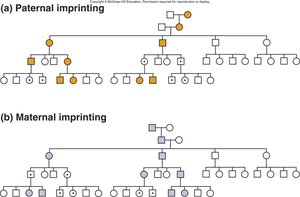
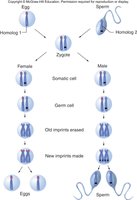
Genomic Imprinting
Genomic imprinting is an epigenetic phenomenon where only one parental allele of a gene is expressed, depending on the parent of origin. This is regulated by DNA methylation at imprinting control regions (ICRs).
Paternal Imprinting: Paternal allele is silenced; expression from maternal allele.
Maternal Imprinting: Maternal allele is silenced; expression from paternal allele.
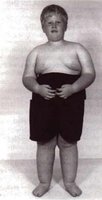
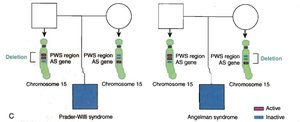
Prader-Willi and Angelman Syndromes
Both syndromes are caused by mutations in the same region of chromosome 15 (q11-13), but the phenotype depends on whether the maternal or paternal copy is affected due to imprinting.
Prader-Willi Syndrome: Paternally inherited mutation; symptoms include obesity, short stature, and intellectual disability.
Angelman Syndrome: Maternally inherited mutation; symptoms include jerky movements, happy demeanor, and intellectual disability.

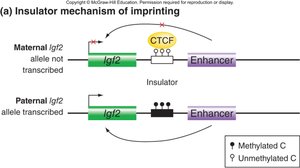
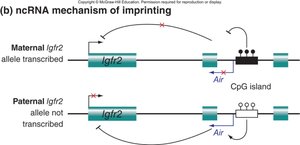
Mechanisms of Imprinting: Insulators and ncRNA
Imprinting can be regulated by insulator elements or noncoding RNAs (ncRNAs). Methylation of insulators can activate or repress gene expression, while ncRNAs can interfere with the expression of sense genes.
Post-Transcriptional Control
RNA Splicing
Alternative splicing allows a single gene to produce multiple protein isoforms by including or excluding specific exons during mRNA processing.
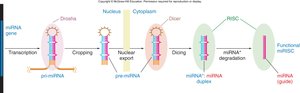
RNA Interference (RNAi)
RNAi is a post-transcriptional gene silencing mechanism involving small noncoding RNAs (miRNAs and siRNAs) that bind to complementary mRNA sequences, leading to mRNA degradation or translational repression.
miRNAs: Endogenously encoded, processed from primary transcripts, and incorporated into RISC complexes.
siRNAs: Derived from double-stranded RNA, processed by Dicer, and guide mRNA cleavage.

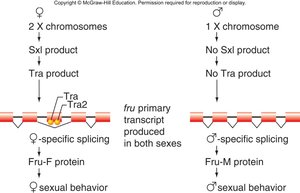
Comprehensive Example: Sex Determination in Drosophila
Genetic Cascade and Alternative Splicing
Sex determination in Drosophila involves a cascade of transcriptional and post-transcriptional controls, including the regulation of the Sex lethal (Sxl), transformer (tra), and doublesex (dsx) genes. Alternative splicing of these genes determines sexual phenotype.
XX (female): Sxl protein produced, leading to female-specific splicing of downstream genes.
XY (male): No Sxl protein, leading to male-specific splicing.

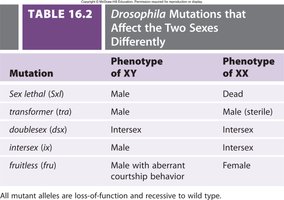
Table: Drosophila Mutations Affecting Sexes Differently
Mutation | Phenotype of XY | Phenotype of XX |
|---|---|---|
Sex lethal (sxl) | Male | Dead |
transformer (tra) | Male | Male (sterile) |
doublesex (dsx) | Intersex | Intersex |
intersex (ix) | Male | Intersex |
fruitless (fru) | Male with aberrant courtship behavior | Female |
All mutant alleles are loss-of-function and recessive to wild type.