 Back
BackGene Interactions and Functional Effects of Mutations
Study Guide - Smart Notes

Gene Interactions
Haplosufficiency and Haploinsufficiency
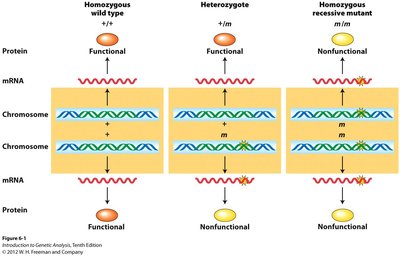
Genes can differ in how many functional copies are required to produce a wild-type phenotype. This concept is central to understanding dominance and recessiveness in genetics.
Haplosufficient gene: A single functional copy is enough to produce the wild-type phenotype. The mutant allele is typically recessive.
Haploinsufficient gene: One functional copy is not enough for the wild-type phenotype. The mutant allele is typically dominant.
Example: If a gene product must reach a threshold (e.g., 45 units) for normal function, a wild-type allele producing 50 units is haplosufficient, while a mutant producing less may be haploinsufficient.
Functional Effects of Mutation
Types of Mutations and Their Effects
Mutations can alter gene function in several ways, leading to different inheritance patterns and phenotypic outcomes.
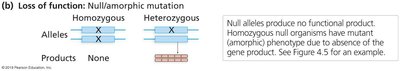
Loss of function: Mutation reduces or eliminates gene activity.
Null (amorphic): No functional product is produced.
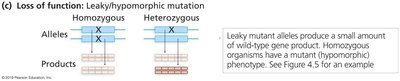
Leaky (hypomorphic): Reduced activity, some product is made.
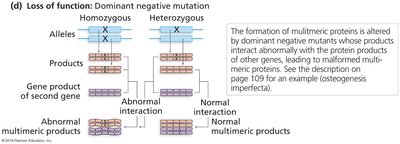
Dominant negative: Mutant protein interferes with normal protein complexes, often causing dominant phenotypes.
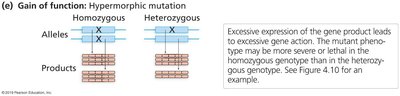
Gain of function: Mutation increases activity or creates a new function.
Hypermorphic: Increased normal activity.
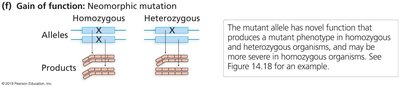
Neomorphic: New activity not seen in the wild-type.

Dominant Negative Mutations and Disease
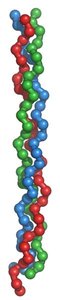
Dominant negative mutations are particularly important in multimeric proteins, where a mutant subunit can disrupt the function of the entire complex. A classic example is osteogenesis imperfecta, a brittle bone disease caused by mutations in collagen genes.
Collagen: A trimeric protein composed of three polypeptides. Mutations in one gene can disrupt the entire structure.
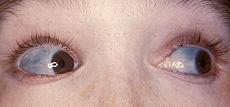
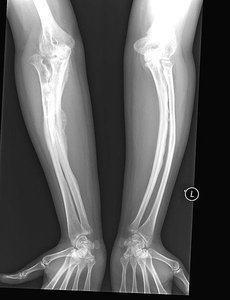
Phenotype: Blue sclera and bone deformities are characteristic symptoms.
Gain-of-Function Mutations
Gain-of-function mutations are almost always dominant and can result in new or excessive activity of the gene product.
Hypermorphic: Excessive normal function.
Neomorphic: New function not present in the wild-type.
Patterns of Dominance
Incomplete Dominance
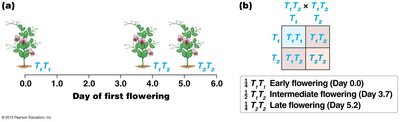
In incomplete dominance, the heterozygote phenotype is intermediate between the two homozygotes, but not necessarily exactly halfway.
Example: Flower color in snapdragons: red (R1R1), white (R2R2), and pink (R1R2).

Codominance
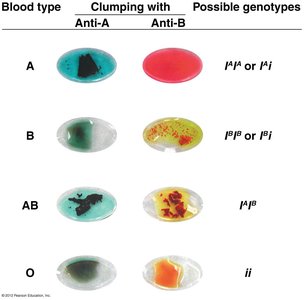
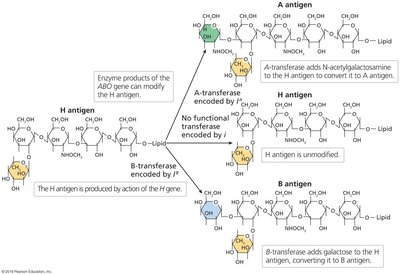
Codominance occurs when both alleles in a heterozygote are fully expressed, resulting in a phenotype that shows both traits distinctly.
Example: Human ABO blood groups, where IA and IB alleles are codominant, and i is recessive.
Allelic Series and Lethal Alleles
Allelic Series
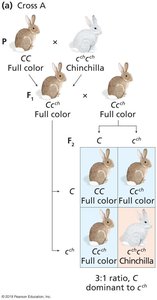
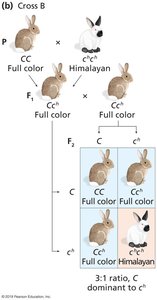
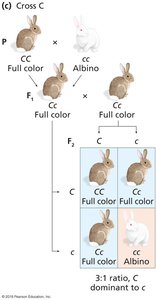
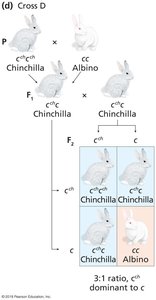
Some genes have multiple alleles that form a dominance hierarchy, resulting in a range of phenotypes.
Example: Coat color in rabbits, with alleles C (full color), cch (chinchilla), ch (Himalayan), and c (albino).

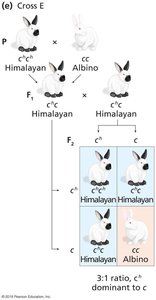
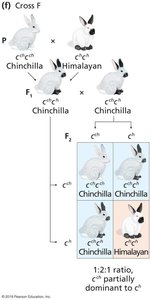
Lethal Alleles
Lethal alleles cause death when present in certain genotypes, often affecting Mendelian ratios.
Dominant lethal: Rare, usually eliminated from populations.
Recessive lethal: More common, only lethal in homozygotes.
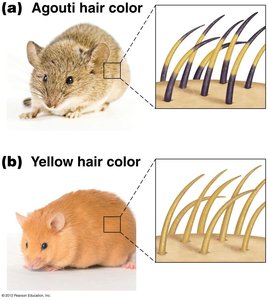
Example: Agouti gene in mice, where the AY allele is dominant for yellow color but recessive lethal.

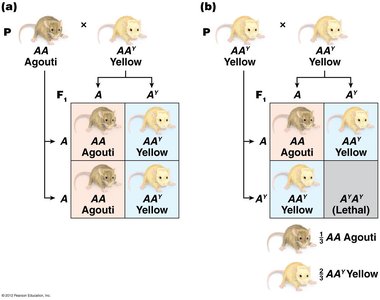
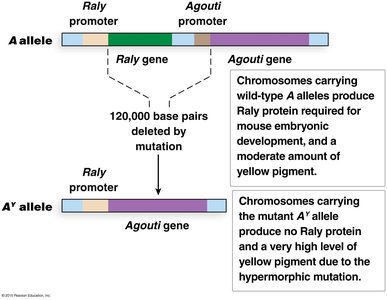
Sex-Influenced and Sex-Limited Traits
Sex-Limited Traits
Sex-limited traits are expressed in only one sex, even though both sexes carry the genes.
Examples: Milk production in mammals, horn development in some animals.
Sex-Influenced Traits
Sex-influenced traits show different phenotypes in males and females for the same genotype, often due to hormonal differences.
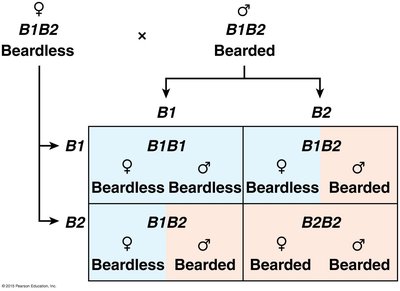
Example: Bearding in goats, where the same genotype can produce bearded males but beardless females.
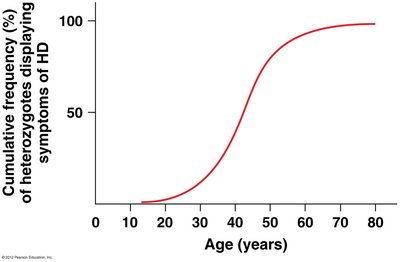
Penetrance and Expressivity
Incomplete Penetrance
Penetrance is the proportion of individuals with a particular genotype who actually express the associated phenotype.
Example: Polydactyly, an autosomal dominant trait, shows incomplete penetrance when not all heterozygotes display extra digits.
Calculation: Penetrance = (number of individuals expressing the phenotype) / (total number with the genotype)
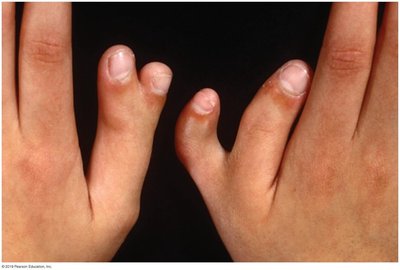
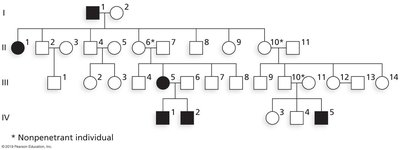
Variable Expressivity
Expressivity refers to the degree or intensity of a phenotype among individuals with the same genotype.
Example: Waardenburg syndrome, where individuals may show different combinations of hearing loss, hair color changes, and eye color.
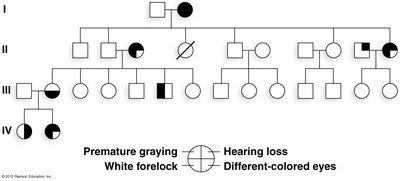
Pleiotropy
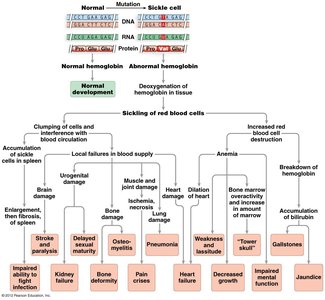
Pleiotropy occurs when a single gene mutation affects multiple traits or systems in an organism.
Example: Sickle cell anemia, where a single mutation affects hemoglobin, red blood cell shape, and multiple organ systems.
Gene Pathways and Genetic Dissection
Anabolic and Catabolic Pathways
Genes often encode enzymes that function in metabolic pathways. The "one gene, one enzyme" hypothesis states that each gene encodes a single enzyme in a pathway.
Anabolic pathways: Build complex molecules from simpler ones.
Catabolic pathways: Break down complex molecules into simpler ones.
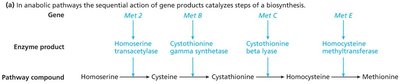
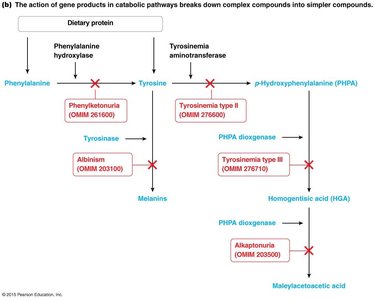
Genetic Dissection of Pathways
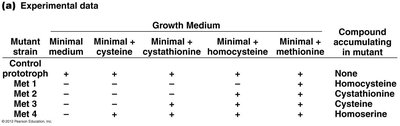
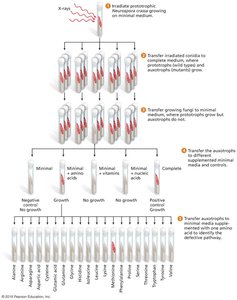
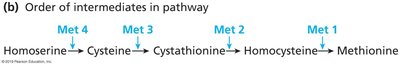
Genetic dissection uses mutants to determine the order and function of genes in a pathway. This approach was pioneered using Neurospora fungi.
Mutants are tested for their ability to grow on media supplemented with pathway intermediates.
The step at which growth is restored indicates the function of the mutated gene.

Epistasis and Gene Interactions
Epistasis
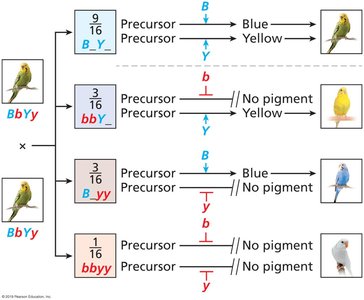
Epistasis is a form of gene interaction where one gene masks or modifies the effect of another gene. This leads to deviations from the classic 9:3:3:1 dihybrid ratio.
Complementary gene interaction (9:7): Both genes are required for the final product.
Duplicate gene interaction (15:1): Either gene can provide the function.
Dominant gene interaction (9:6:1): Both single mutants have the same phenotype.
Recessive epistasis (9:3:4): Homozygosity for a recessive allele masks the effect of another gene.
Dominant epistasis (12:3:1): A dominant allele at one locus masks the effect of another gene.
Dominant suppression (13:3): A dominant allele suppresses the expression of another gene.
Genetic Analysis of Gene Interactions
Geneticists use complementation analysis to determine whether mutations affect the same or different genes. This involves crossing individuals with different mutations and observing the phenotype of the offspring.
Complementation: If the offspring are wild-type, the mutations are in different genes.
No complementation: If the offspring are mutant, the mutations are in the same gene.
