 Back
BackGene Linkage and Chromosome Mapping in Eukaryotes
Study Guide - Smart Notes

Gene Linkage and Chromosome Mapping
Gene Linkage: Basic Concepts
Gene linkage refers to the physical association of genes located on the same chromosome. Linked genes tend to be inherited together because their loci are close to each other, reducing the likelihood of recombination separating them during meiosis.
Locus: The specific location of a gene on a chromosome.
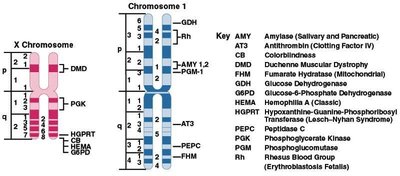
Linkage Group: Each linkage group corresponds to a pair of homologous chromosomes. Genes within a linkage group are inherited together unless separated by crossing over.

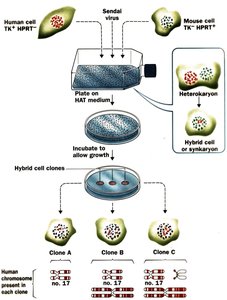
Clone: Definition and Relevance
A clone is a group of genetically identical individuals derived from a single parent through nonsexual propagation. Cloning is important in genetics for studying gene function and inheritance.
Application: Clones are used in genetic mapping and functional studies.
Types of Linkage
Linkage can be classified based on the frequency of recombination between genes:
Independent Assortment (IA): Genes assort independently; parentals and recombinants are produced in equal proportions (P = 50%, R = 50%).
Complete Linkage (CL): No recombination occurs; only parental types are produced (P = 100%, R = 0%).
Normal Linkage (NL): Some recombination occurs; parental types are more frequent than recombinants (0% < R < 50%, 50% < P < 100%).
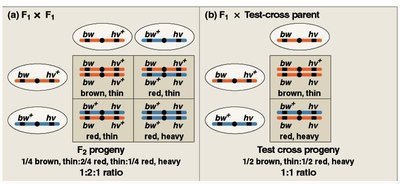
Testcross and Linkage Analysis
Testcrosses are used to determine linkage and calculate recombination frequencies. The proportion of recombinant offspring indicates the distance between genes.
Recombination Frequency: Calculated as .
Map Units (m.u.): 1 m.u. = 1% recombination.
Gene Mapping and Crossing Over
Gene mapping uses recombination frequencies to estimate distances between genes. Crossing over during meiosis produces recombinant gametes, which are used to construct genetic maps.
Single Crossover: Produces recombinant gametes for the interval between two loci.
Double Crossover: Occurs between three loci, helping determine gene order.
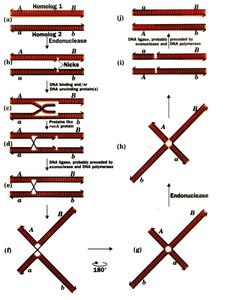
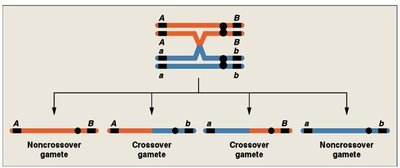
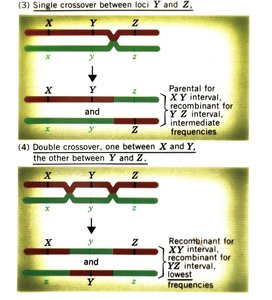
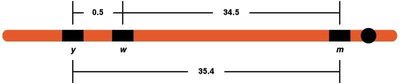
Calculating Map Distances
Map distances are calculated using recombination frequencies. For three-point crosses, the gene order can be determined by comparing parental and double crossover types.
Example Calculation:
Gene Order: The gene that differs in double crossover types compared to parental types is placed in the middle.
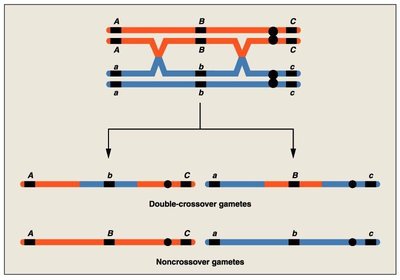
Interference and Coincidence
Interference describes how one crossover event can affect the probability of another nearby crossover. Coincidence is the ratio of observed to expected double crossovers.
Coincidence (C):
Interference (I):
Chromosome Mapping: Practical Examples
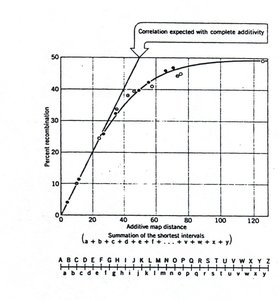
Chromosome maps show the relative positions of genes based on recombination frequencies. These maps are essential for understanding genetic linkage and inheritance patterns.
Summary Table: Linkage Types and Recombination
Linkage Type | Parentals (%) | Recombinants (%) |
|---|---|---|
Independent Assortment (IA) | 50 | 50 |
Complete Linkage (CL) | 100 | 0 |
Normal Linkage (NL) | 50-100 | 0-50 |
Applications and Examples
Drosophila Mapping: Classic experiments with fruit flies demonstrate linkage, recombination, and gene mapping.
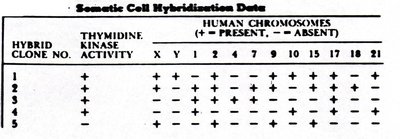
Somatic Cell Hybridization: Used to assign genes to specific chromosomes.


Key Terms
Linkage Group
Locus
Clone
Recombination Frequency
Map Unit (m.u.)
Interference
Coincidence
Additional info:
Gene linkage and chromosome mapping are fundamental for understanding inheritance patterns and genetic diseases.
Somatic cell hybridization is a technique used to map genes to chromosomes by fusing cells from different species and analyzing gene expression.
