 Back
BackCh 14 P2 Gene Mutation, DNA Repair, and Transposition (Essentials of Genetics, Ch. 14) – Study Notes
Study Guide - Smart Notes

Ch 14 P2 Gene Mutation, DNA Repair, and Transposition
Introduction
This chapter explores the molecular mechanisms underlying gene mutation, the cellular systems for DNA repair, and the biology of transposable elements. Understanding these processes is essential for comprehending genetic variation, genome stability, and the evolution of genomes in all organisms.
DNA Mutation and Its Consequences
Types of Mutations
Point Mutations: Changes affecting a single nucleotide pair.
Insertions/Deletions (Indels): Addition or loss of one or more nucleotide pairs.
Frameshift Mutations: Indels that are not multiples of three, altering the reading frame.
Example: The most common mutation causing cystic fibrosis is a three-base-pair deletion at position 508 of the CFTR protein, resulting in the deletion of a single amino acid (phenylalanine) in the protein sequence.
DNA Repair Mechanisms
Proofreading and Mismatch Repair (MMR)
Cells possess multiple systems to correct errors that occur during DNA replication:
Proofreading: DNA polymerase detects and removes incorrectly inserted nucleotides during replication.
Mismatch Repair (MMR): If proofreading fails, MMR detects mismatches, excises the incorrect nucleotide (using endonucleases and exonucleases), and fills the gap with the correct nucleotide.
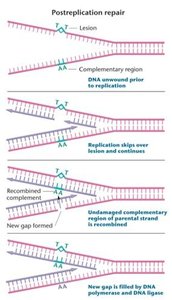
Postreplication Repair and the SOS Repair System
When DNA damage escapes repair and replication proceeds past the lesion, specialized repair systems are activated:
Postreplication Repair: RecA protein mediates recombination between the damaged and undamaged parental DNA strands, allowing the gap to be filled by DNA polymerase and ligase. This is a form of homologous recombination repair.
Excision Repair Pathways
Excision repair mechanisms remove and replace damaged DNA segments:
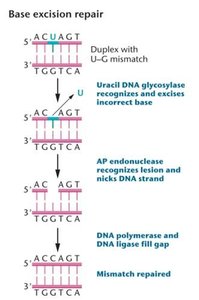
Base Excision Repair (BER)
Corrects DNA containing a damaged base.
DNA glycosylase recognizes and removes the altered base.
AP endonuclease cuts the DNA backbone.
DNA polymerase and ligase fill and seal the gap.
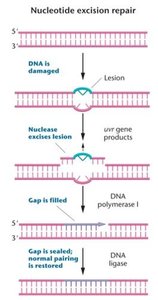
Nucleotide Excision Repair (NER)
Repairs bulky lesions that distort the DNA double helix (e.g., thymine dimers).
uvr gene products recognize and excise the lesion.
In E. coli, 13 nucleotides are removed; in eukaryotes, 28 nucleotides are excised.
DNA polymerase and ligase fill and seal the gap.
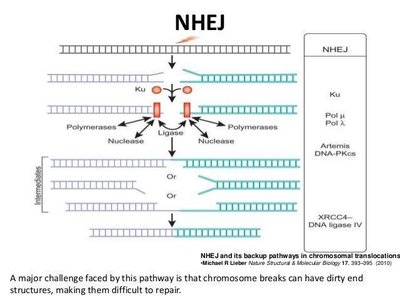
Double-Strand Break (DSB) Repair in Eukaryotes
Double-strand breaks are highly dangerous and can lead to chromosomal rearrangements, cancer, or cell death. Two main pathways repair DSBs:
Homologous Recombination Repair: Recognizes the break, digests the 5' ends, and uses the sister chromatid as a template for accurate repair. Occurs during late S or early G2 phase.
Nonhomologous End Joining (NHEJ): Activated in G1, this pathway directly ligates the broken DNA ends, often with the help of proteins such as Ku, DNA ligase IV, and others. It is more error-prone than homologous recombination.
Transposable Elements (TEs)
Overview of Transposable Elements
Also known as "jumping genes," TEs are DNA sequences that can move within and between chromosomes.
Found in all organisms; their precise function is still under investigation.
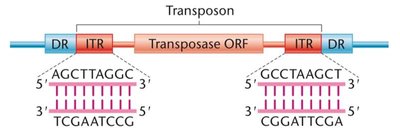
DNA Transposons
Move without an RNA intermediate.
Contain inverted terminal repeats (ITRs) at their ends, recognized by the enzyme transposase.
Transposase is required for the movement of the TE in and out of the genome.
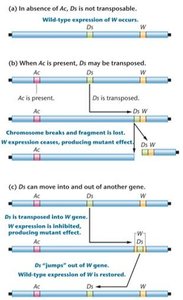
The Ac-Ds System in Maize
Discovered by Barbara McClintock (Nobel Prize, 1983).
Ac (Activator) and Ds (Dissociation) are mobile controlling elements in corn.
Ds is non-autonomous and requires Ac for transposition.
Ds can cause chromosome breakage or disrupt gene expression by inserting into or excising from genes.
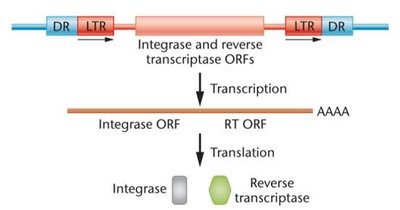
Retrotransposons
Move via an RNA intermediate (copy-and-paste mechanism).
Resemble retroviruses in structure and replication.
Two types: LTR (long terminal repeat) and non-LTR retrotransposons.
Encode enzymes such as integrase and reverse transcriptase.
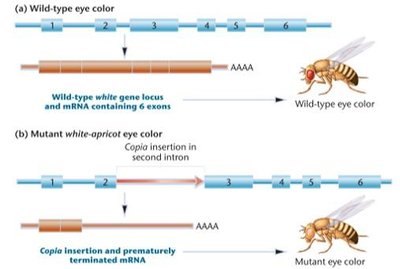
Transposable Elements in Drosophila: The Copia–White-Apricot System
Copia elements are a class of DNA elements in Drosophila, transcribed into large amounts of RNA and dispersed throughout the genome.
The white-apricot mutation is caused by the insertion of copia into the second intron of the white gene, changing eye color from red to orange-yellow due to premature mRNA termination.
Transposable Elements in Humans
LINEs (Long Interspersed Elements) and SINEs (Short Interspersed Elements) make up a significant portion of the human genome.
LINEs are 1–6 kb long, with about 850,000 copies (21% of the genome).
SINEs are 100–500 bp long, with about 1,500,000 copies (13% of the genome).
Other TEs include LTR elements and DNA transposons.
Element Type | Length | Copies in Genome | % of Genome |
|---|---|---|---|
LINEs | 1–6 kb | 850,000 | 21 |
SINEs | 100–500 bp | 1,500,000 | 13 |
LTR elements | <5 kb | 443,000 | 8 |
DNA transposons | 80–300 bp | 294,000 | 3 |
Unclassified | — | 3,000 | 0.1 |
Transposons, Mutation, and Evolution
Effects of Transposable Element Insertions
Insertion into coding regions can disrupt translation.
Insertion into introns can cause premature transcription termination or alter RNA splicing.
Insertion into regulatory regions can affect gene expression.
Identical TEs in the genome can facilitate recombination, leading to chromosomal rearrangements.
Potential Effects of Transposons:
Gene inactivation by insertional mutagenesis
Change in gene regulation by epigenetic modification of nearby chromatin
Altering splice sites of genes
Exon shuffling
All of the above
