Back
BackGene Mutation: Types, Mechanisms, and Effects
Study Guide - Smart Notes

Gene Mutation: Types, Mechanisms, and Effects
Introduction to Genetic Mutations
Genetic mutations are rare alterations in the DNA nucleotide sequence that serve as the foundation for genetic variability and evolution. Mutations can occur anywhere in the genome and may affect coding or noncoding regions, leading to a range of phenotypic outcomes from beneficial to harmful.
Mutation: Any rare change in DNA sequence, including single base pair substitutions, deletions, or major chromosomal alterations.
Genetic variability: Mutations are the raw material for evolution, providing diversity within populations.
Phenotypic effects: Mutations may be beneficial, neutral, or harmful, and may or may not cause observable changes.
Where Do Mutations Occur?
Mutations can occur in any region of the genome, including coding regions, gene regulatory sequences (such as promoters and enhancers), splicing signals, repetitive sequences, and other noncoding regions. They may arise in different cells and chromosomes.
Types of Mutations: Molecular Classification
Mutations are classified based on their molecular nature and impact on the DNA sequence.
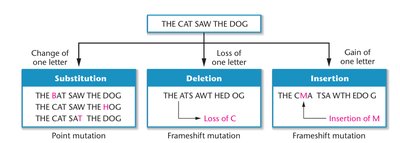
Point mutation (base substitution): Change from one base pair to another.
Transition: Purine-to-purine (A ↔ G) or pyrimidine-to-pyrimidine (C ↔ T) substitution.
Transversion: Purine-to-pyrimidine or vice versa.
Missense mutation: New codon codes for a different amino acid.
Nonsense mutation: Amino acid codon converted to a stop codon, terminating translation prematurely.
Silent mutation: New codon codes for the same amino acid or occurs in a non-coding region.
Frameshift mutation: Caused by insertions or deletions, resulting in a shift in the reading frame and altered downstream codons.
Mutations Classified by Effect on Function
Mutations can be categorized based on their impact on gene product function.
Loss-of-function: Reduces or eliminates gene product function; null mutations eliminate the product entirely.
Recessive: Most loss-of-function mutations; wild-type phenotype when heterozygous with wild-type allele.
Dominant: Mutant phenotype appears even with wild-type allele present.
Dominant negative: Inactive gene product interferes with wild-type product.
Haploinsufficiency: Wild-type allele does not produce enough product to result in wild-type phenotype.
Gain-of-function: Allele produces a gene product with enhanced, negative, or new function; usually dominant.
Suppressor: Second mutation relieves or reverts effects of original mutation; can be intragenic or intergenic.
Mutations Classified by Location
Mutations are also classified by their location within the organism.
Somatic: Occur in any cell except germ cells; not heritable.
Germ-line: Occur in gamete-producing cells; inherited.
Autosomal: On any chromosome other than sex chromosomes.
Sex chromosome linked: On X or Y chromosome.
Spontaneous vs Induced Mutations
Mutations can arise spontaneously due to normal biological processes or be induced by external agents known as mutagens.
Spontaneous mutations: Result from DNA replication errors, tautomeric shifts, depurination, and deamination.
Induced mutations: Caused by mutagens such as chemicals, radiation, and environmental factors.
Rates of Mutation
The spontaneous mutation rate is the likelihood that a nucleotide, gene, or genome will undergo mutation in a single replication or generation. Rates are extremely low and vary between organisms, genes, and genomic regions.
Viruses and bacteria: 1 in 100 million per replication.
Animals: Lower rates due to efficient DNA proofreading and repair.
Mutation hotspots: Regions with higher mutation rates.
Spontaneous Mutations: DNA Replication Errors
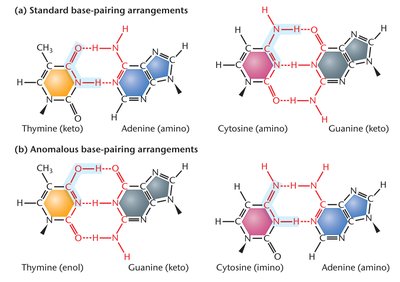
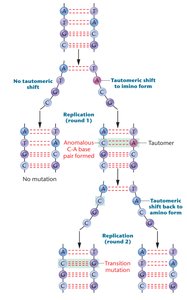
DNA replication is not perfect; DNA polymerase may insert incorrect nucleotides, which can persist if proofreading and repair fail. Tautomeric shifts and replication slippage contribute to these errors.
Tautomeric shifts: Alternate chemical forms of bases increase mispairing during replication.
Replication slippage: DNA polymerase slips in repeat sequences, causing insertions or deletions.
Spontaneous Mutations: Depurination and Deamination
Depurination and deamination are chemical changes that alter DNA bases, leading to mutations.
Depurination: Loss of a purine base (adenine or guanine), creating an apurinic site; random nucleotide is introduced during replication.
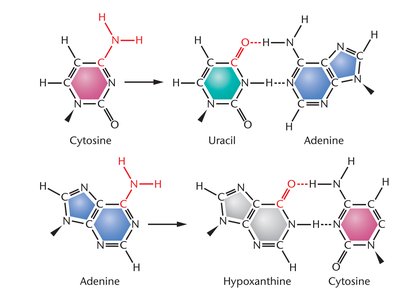
Deamination: Amino group in cytosine or adenine is converted to a keto group, resulting in altered base pairing (cytosine to uracil, adenine to hypoxanthine).
Induced Mutations: Mutagens
Mutagens are natural or artificial agents that induce mutations by interacting with DNA.
Examples: Fungal toxins, cosmic rays, ultraviolet light, industrial pollutants, medical X-rays, chemicals in tobacco smoke and cooked meats, certain dyes.
Induced Mutations: Base Analogs
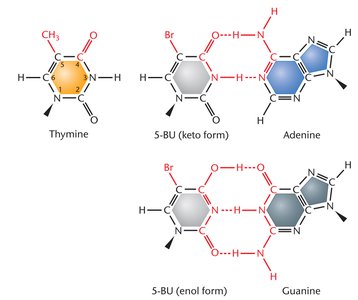
Base analogs are chemicals that resemble DNA bases and can substitute for purines or pyrimidines during nucleic acid biosynthesis, increasing mutation rates.
Example: 5-Bromouracil (5-BU) behaves as a thymine analog, increasing tautomeric shifts and sensitivity to UV light.
Application: Used in some cancer treatments.
Induced Mutations: Alkylating Agents
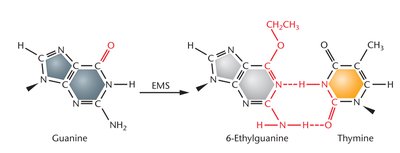
Alkylating agents donate alkyl groups to amino or keto groups in nucleotides, altering base-pairing affinity and causing transition mutations.
Example: Ethyl methanesulfonate (EMS) modifies guanine to 6-ethylguanine, which pairs with thymine.
Induced Mutations: UV Radiation
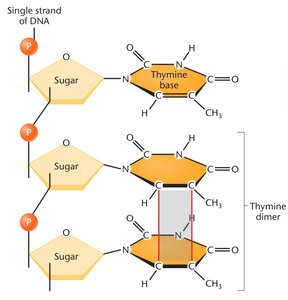
Ultraviolet (UV) radiation disrupts organic molecules by creating pyrimidine dimers, which distort DNA conformation and affect replication.
Pyrimidine dimers: Two identical pyrimidines (usually thymine) covalently bonded, interfering with DNA replication.
Absorption: Purines and pyrimidines absorb UV at 260 nm.
Induced Mutations: Ionizing Radiation
Ionizing radiation penetrates cells and transforms stable molecules into free radicals, which directly or indirectly affect genetic material.
Free radicals: Chemical species with unpaired electrons that alter purines and pyrimidines, break phosphodiester bonds, and produce deletions, translocations, and fragmentation.
Example: Lactase Persistence and Mutation
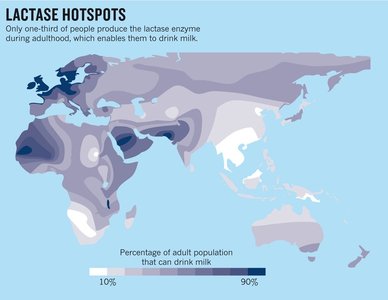
Lactase persistence is an example of a point mutation allowing continuous production of lactase enzyme in some individuals, enabling them to digest milk into adulthood. The distribution of lactase persistence varies globally.
Genetic basis: Mutation in regulatory region of the lactase gene.
Population variation: Only one-third of adults worldwide can digest milk.