 Back
BackGene Regulation in Bacteria: lac and trp Operons
Study Guide - Smart Notes

Gene Regulation in Bacteria
Overview of Transcriptional Regulation
Transcriptional regulation in bacteria is a fundamental mechanism that controls gene expression, allowing cells to respond to environmental changes. This regulation involves specific DNA sequences, regulatory proteins, and small organic molecules that modulate the rate of transcription.
Regulatory Proteins:
Activators increase transcription rate (positive control).
Repressors decrease transcription rate (negative control).
Allosteric Effectors:
Inducers increase transcription by binding to repressors (preventing DNA binding) or activators (enabling DNA binding).
Corepressors decrease transcription by enabling repressors to bind DNA.
Inhibitors decrease transcription by preventing activators from binding DNA.
Regulated Genes:
Inducible genes are usually "off" until turned "on" by an inducer.
Repressible genes are usually "on" until turned "off" by a corepressor or inhibitor.
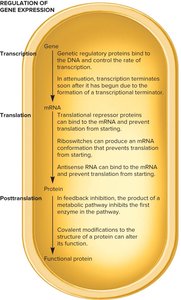
Mechanisms of Transcriptional Regulation
Regulatory proteins interact with DNA and effectors to control transcription. The following mechanisms illustrate how these interactions occur:
Inducible gene (repressor, inducer): Inducer binds to repressor, preventing it from binding DNA, allowing transcription.
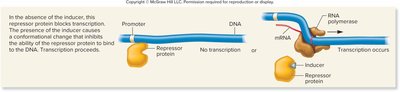
Inducible gene (activator, inducer): Inducer binds to activator, enabling it to bind DNA and activate transcription.

Repressible gene (repressor, corepressor): Corepressor binds to repressor, enabling it to bind DNA and inhibit transcription.
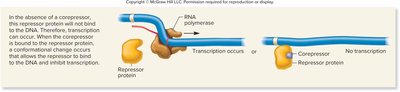
Repressible gene (activator, inhibitor): Inhibitor binds to activator, preventing it from binding DNA and inhibiting transcription.
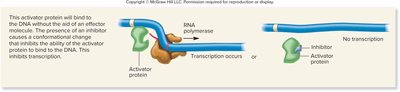
Regulation of the lac Operon
Structure and Function of the lac Operon
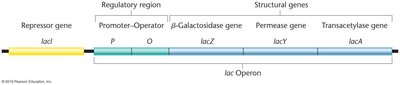
The lac operon is a classic example of gene regulation in bacteria. It encodes enzymes required for lactose metabolism and is tightly regulated to ensure efficient energy use.
Operon: A DNA segment encoding a polycistronic mRNA under a single promoter.
lac Operon Genes:
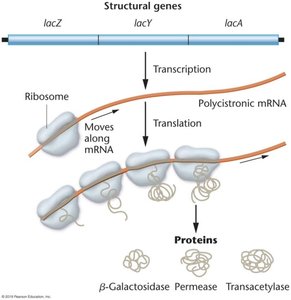
lacZ: β-galactosidase (cleaves lactose).
lacY: Permease (transports lactose).
lacA: Transacetylase (detoxifies by-products).
Regulatory Region: Includes promoter, operator, and repressor gene (lacI).
Regulation of the lac Operon: Induction and Repression
The lac operon is regulated based on the presence of glucose and lactose:
Glucose present, lactose absent: lac operon is OFF.
Glucose absent, lactose present: lac operon is ON.
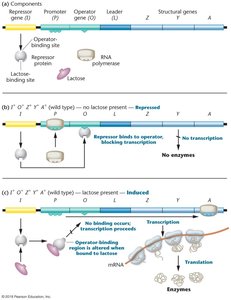
Cycle of lac Operon Repression and Induction
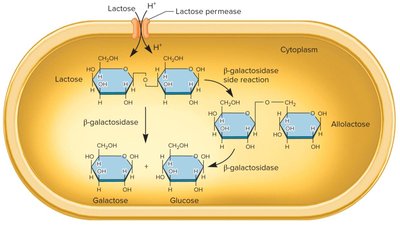
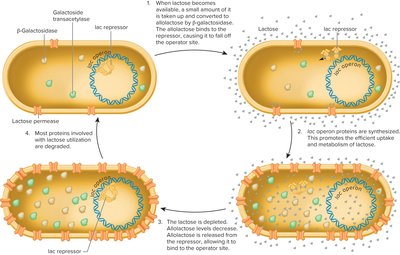
When lactose is available, it is converted to allolactose, which acts as an inducer. Allolactose binds to the lac repressor, causing it to release from the operator, allowing transcription of the operon genes. When lactose is depleted, the repressor binds again, shutting off transcription.
Mutational Analysis of the lac Operon
Experiments with mutant and wild-type E. coli strains revealed:
Operators: cis-acting DNA sequences.
Repressors: trans-acting proteins.
lacI– and lacOC mutations: Cause constitutive transcription (always ON).
lacIS mutation: Super-repressor, always OFF.
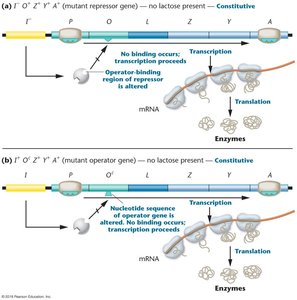
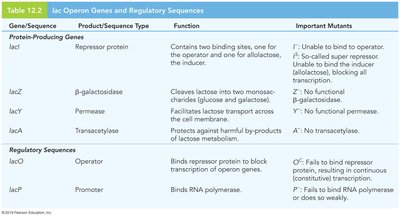
Table: lac Operon Genes and Regulatory Sequences
Gene/Sequence | Product/Sequence Type | Function | Important Mutants |
|---|---|---|---|
lacI | Repressor protein | Binds operator; allolactose-binding site | I-: No binding; IS: Super-repressor |
lacZ | β-galactosidase | Cleaves lactose | Z-: No function |
lacY | Permease | Lactose transport | Y-: No function |
lacA | Transacetylase | Detoxifies by-products | A-: No function |
lacO | Operator | Binds repressor | OC: No binding; constitutive transcription |
lacP | Promoter | Binds RNA polymerase | P-: No transcription |
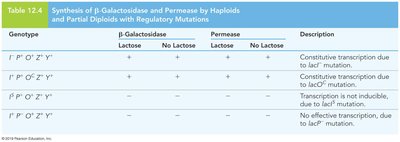
Catabolite Repression of the lac Operon
Catabolite repression ensures that glucose is used preferentially over lactose. When glucose is present, cAMP levels are low, and CAP (catabolite activator protein) cannot bind efficiently, repressing the lac operon. When glucose is absent, cAMP levels rise, cAMP binds to CAP, and the CAP-cAMP complex activates transcription.
CAP-cAMP complex: Binds to CAP site, increases RNA polymerase affinity for lac promoter.
Glucose present: Transcription diminished.
Glucose absent: Transcription occurs.
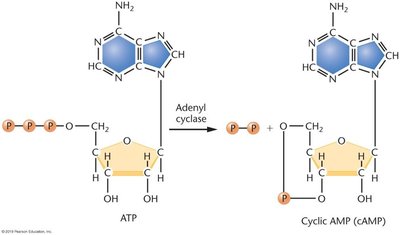
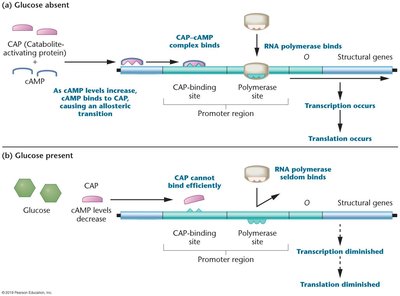
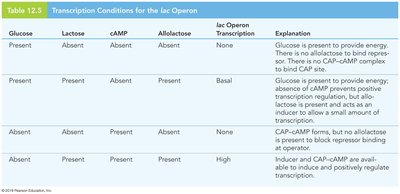
Table: Transcription Conditions for the lac Operon
Glucose | Lactose | cAMP | Allolactose | lac Operon Transcription | Explanation |
|---|---|---|---|---|---|
Present | Absent | Absent | Absent | None | Glucose present, no allolactose, no CAP-cAMP |
Present | Present | Absent | Present | Basal | Glucose present, allolactose induces, but no CAP-cAMP |
Absent | Absent | Present | Absent | None | CAP-cAMP present, but no allolactose |
Absent | Present | Present | Present | High | Inducer and CAP-cAMP present |
Regulation of the trp Operon
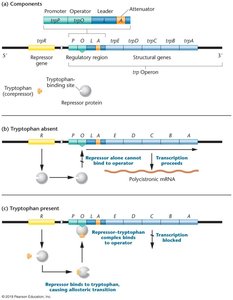
Structure and Function of the trp Operon
The trp operon encodes enzymes for tryptophan biosynthesis. Its regulation ensures that tryptophan is synthesized only when needed.
Tryptophan absent: trp operon is ON (inactive repressor).
Tryptophan present: trp operon is OFF (corepression and attenuation).
Attenuation in the trp Operon
Attenuation is a regulatory mechanism that uses the leader sequence (trpL) to sense tryptophan levels and form stem-loop structures in mRNA, affecting transcription termination.
trpL leader sequence: Encodes a peptide with two Trp codons and forms stem-loop structures.
Low Trp levels: Ribosome stalls, 2-3 stem-loop forms, transcription proceeds.
High Trp levels: Ribosome moves quickly, 3-4 stem-loop forms, transcription terminates.
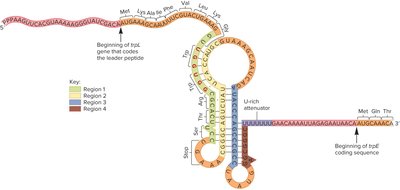
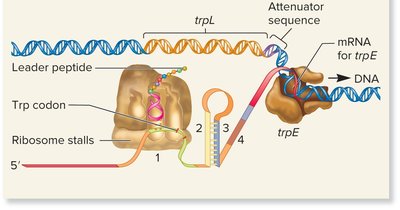
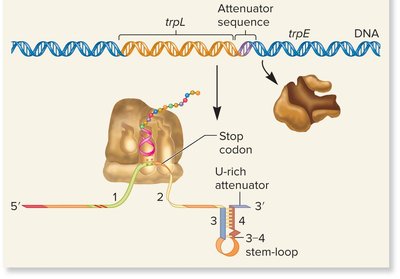
Table: Stem-Loop Structures in trpL Leader RNA
Trp Levels | Stem-Loop Formed | Transcription Outcome |
|---|---|---|
Low | 2-3 | Transcription proceeds |
High | 3-4 | Transcription terminates |
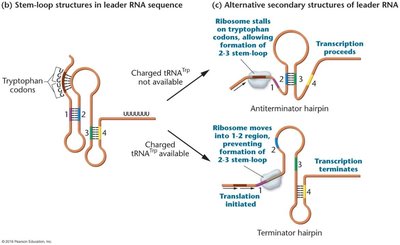
Additional info: Attenuation is unique to certain bacterial operons and allows fine-tuned regulation of biosynthetic pathways based on metabolite availability.
