 Back
BackGene Regulation in Bacteria: Mechanisms and Genetic Analysis
Study Guide - Smart Notes

Gene Regulation in Bacteria
Introduction to Gene Regulation
Gene regulation in bacteria is essential for adapting to environmental changes and efficiently utilizing resources. Bacteria use a variety of genetic mechanisms to control the expression of their genes, allowing them to respond rapidly to environmental signals and metabolic needs.
Constitutive transcription: Genes are continuously transcribed regardless of environmental conditions.
Regulated transcription: Genes are turned on or off in response to environmental changes.
Regulatory proteins: DNA-binding proteins that interact with regulatory DNA sequences to control transcription.
Levels of Gene Regulation
Gene expression in bacteria can be regulated at multiple levels, primarily at the transcriptional and posttranscriptional stages.
Transcriptional Regulation: Involves controlling the initiation and amount of mRNA produced. Mechanisms include inducible and repressible systems, as well as attenuation.
Posttranscriptional Regulation: Involves mRNA stability and translation efficiency, such as mRNA destruction and translation blockage.
Mechanisms of Transcriptional Regulation
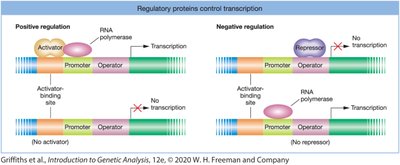
Bacteria use both positive and negative regulatory mechanisms to control gene expression.
Negative regulation: Repressor proteins bind to operators to block transcription.
Positive regulation: Activator proteins enhance RNA polymerase binding to promoters, increasing transcription.
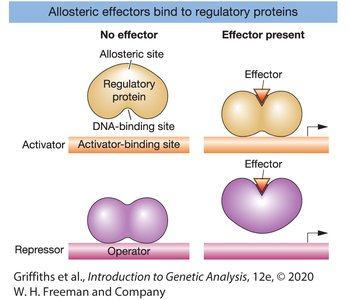
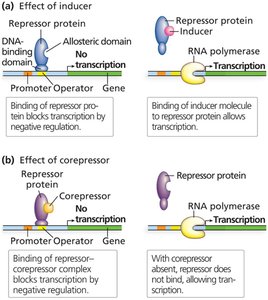
Allosteric Regulation of Regulatory Proteins
Regulatory proteins often have allosteric sites that bind effectors (inducers or corepressors), altering their ability to bind DNA and regulate transcription.
Inducer: A small molecule that binds to a repressor, inactivating it and allowing transcription.
Corepressor: A small molecule that activates a repressor, enabling it to bind DNA and block transcription.

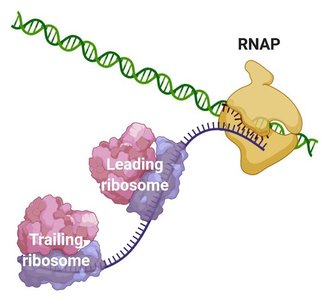
Coupled Transcription and Translation in Bacteria
In bacteria, transcription and translation are tightly coupled, allowing ribosomes to begin translating mRNA while it is still being synthesized by RNA polymerase.
The Lac Operon: A Model for Inducible Gene Regulation
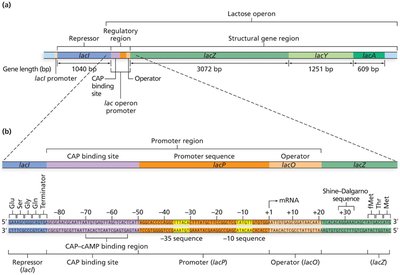
Structure and Function of the Lac Operon
The lac operon is a classic example of an inducible system, allowing E. coli to metabolize lactose only when it is available and glucose is scarce.
Structural genes: lacZ (β-galactosidase), lacY (permease), lacA (transacetylase).
Regulatory elements: Promoter, operator, and the lacI gene (encodes the repressor protein).
The operon is transcribed as a single polycistronic mRNA and translated into three distinct proteins.
Mechanism of Lac Operon Regulation
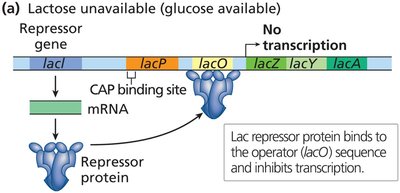
The lac operon is regulated by both negative and positive control mechanisms:
Negative control: The LacI repressor binds to the operator, blocking transcription in the absence of lactose.
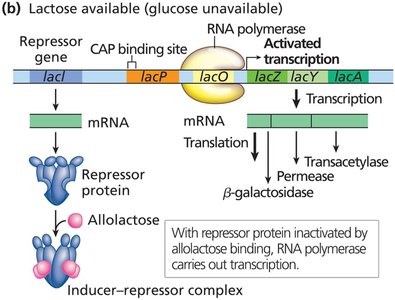
Induction: When lactose is present, allolactose (the inducer) binds to the repressor, inactivating it and allowing transcription.
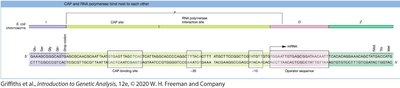
Positive control: The catabolite activator protein (CAP) binds to the promoter in the presence of cAMP (when glucose is low), enhancing RNA polymerase binding and transcription.
Catabolite Repression and the Role of cAMP
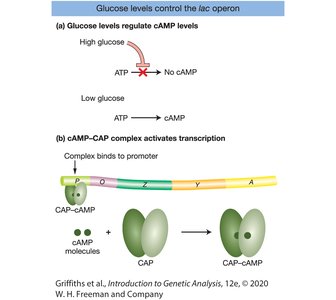
Catabolite repression ensures that the lac operon is only highly expressed when glucose is absent and lactose is present. This is mediated by the cAMP-CAP complex.
High glucose inhibits cAMP production, preventing CAP from binding to the promoter.
Low glucose increases cAMP, allowing CAP to bind and activate transcription.
Summary Table: Lac Operon Regulation
The following table summarizes the regulatory states of the lac operon under different conditions:
Glucose | Lactose | LacI (Repressor) | CAP/cAMP | Operon State |
|---|---|---|---|---|
- | - | Bound | Bound | Off |
- | + | Not bound | Bound | On |
+ | + | Not bound | Not bound | Low (leaky) |
+ | - | Bound | Not bound | Off |
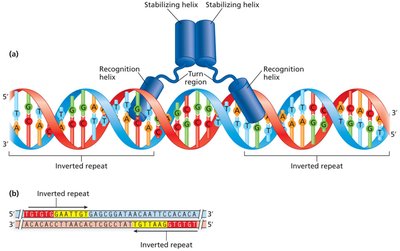
DNA Binding Motifs in Regulatory Proteins
Many bacterial regulatory proteins use the helix-turn-helix motif to recognize and bind specific DNA sequences, such as operators and activator binding sites.
The Tryptophan (trp) Operon: A Model for Repressible Gene Regulation
Structure and Regulation of the trp Operon
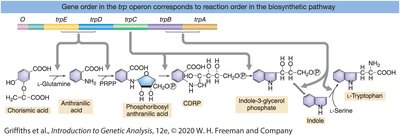
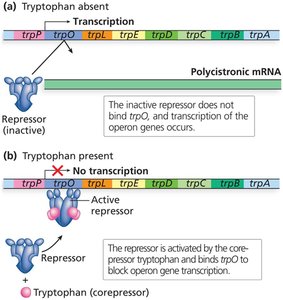
The trp operon encodes enzymes for tryptophan biosynthesis and is regulated by a repressible system. When tryptophan is abundant, it acts as a corepressor, activating the trp repressor to block transcription.
Repressible system: Default state is 'on'; turned 'off' when the end product (tryptophan) is abundant.
trpR gene: Encodes the trp repressor protein.
Corepressor: Tryptophan binds to the repressor, enabling it to bind the operator and inhibit transcription.
Alternative Sigma Factors and Global Regulation
Role of Sigma Factors
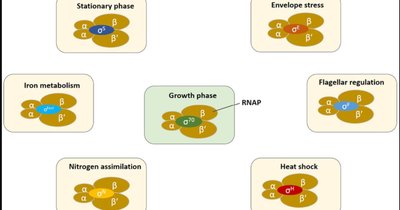
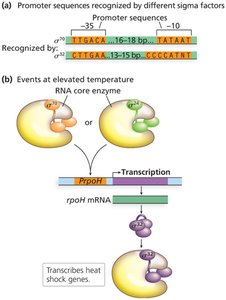
Sigma factors are specialized subunits of bacterial RNA polymerase that direct the enzyme to specific promoter sequences, allowing coordinated expression of large sets of genes in response to environmental changes (e.g., heat shock, stationary phase, nitrogen metabolism).
Alternative sigma factors enable bacteria to rapidly reprogram gene expression in response to stress or developmental signals.
Historical Perspective: The Discovery of Gene Regulation
The foundational work of Jacob and Monod on the lac operon established the concept of regulatory genes and repressor proteins, which has become a central paradigm in molecular genetics.
"Anything found to be true of E. coli must also be true of Elephants." – Jacob and Monod
Key Terms and Concepts
Operon: A cluster of genes under the control of a single promoter and regulatory elements, transcribed as a single mRNA.
Inducible system: Genes are expressed only in the presence of an inducer (e.g., lac operon).
Repressible system: Genes are expressed unless repressed by a corepressor (e.g., trp operon).
Allosteric regulation: Regulatory proteins change conformation upon binding small molecules, altering their DNA-binding ability.
Catabolite repression: A mechanism that ensures the preferential use of certain carbon sources (e.g., glucose) over others (e.g., lactose).
Summary
Gene regulation in bacteria is achieved through a combination of negative and positive control mechanisms, involving regulatory proteins, allosteric effectors, and global regulators such as sigma factors. The lac and trp operons serve as paradigms for understanding inducible and repressible systems, respectively, and illustrate how bacteria efficiently adapt to their environment through precise genetic control.
