 Back
BackGene Regulation in Prokaryotes and Eukaryotes: Mechanisms and Chromatin Structure
Study Guide - Smart Notes

Gene Regulation in Prokaryotes
Trp Operon: Negative Repressible Operon and Attenuation
The trp operon in Escherichia coli is a classic example of a negative repressible operon, which is regulated both at the level of repression and by attenuation. This operon encodes enzymes required for tryptophan biosynthesis and is tightly regulated to conserve resources.
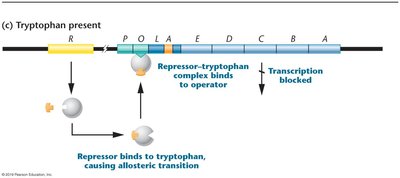
Repression: When tryptophan is abundant, it acts as a corepressor by binding to the trp repressor protein, enabling it to bind the operator and block transcription of the operon.
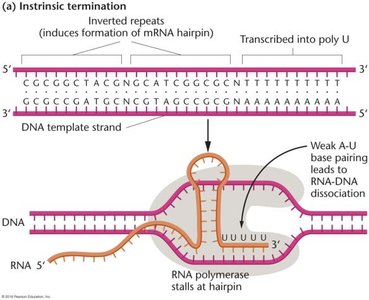
Attenuation: Even with repression, some transcription can occur ('leaky expression'). Attenuation provides an additional regulatory mechanism, using the formation of alternative secondary structures in the leader RNA to terminate transcription prematurely when tryptophan is present.
Leader Region: Contains two tryptophan codons and can form either an antiterminator or terminator hairpin, depending on tryptophan availability.
Mechanism: When tryptophan is low, ribosomes stall at the Trp codons, allowing the formation of the antiterminator hairpin (regions 2-3), and transcription proceeds. When tryptophan is high, ribosomes do not stall, and the terminator hairpin (regions 3-4) forms, causing transcription termination.
Combined Regulation: Repression reduces transcription ~70-fold, attenuation adds another 8-10 fold reduction, resulting in over 600-fold reduction in gene expression when both mechanisms are active.
Generalization: Attenuation is also found in other amino acid biosynthetic operons (e.g., threonine, histidine, leucine, phenylalanine).
Gene Regulation in Eukaryotes
Overview of Eukaryotic Gene Expression
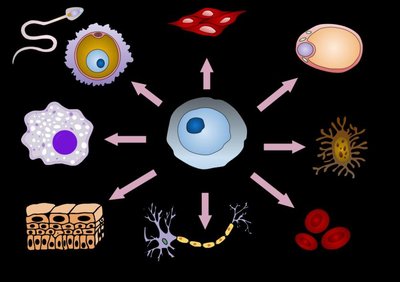
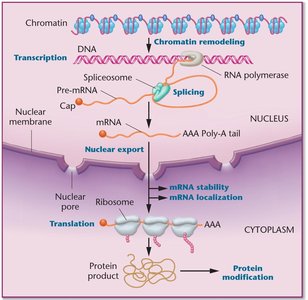
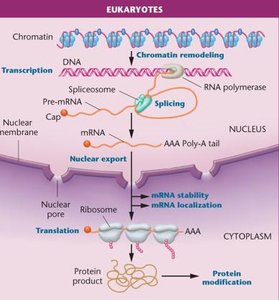
Gene expression in eukaryotes is more complex than in prokaryotes, involving multiple levels of regulation. All cells in a multicellular organism contain the same DNA, but different cell types express different sets of genes in response to internal and external signals.
Levels of Regulation: Transcriptional, post-transcriptional, translational, and post-translational.
Chromatin Structure: DNA is packaged with histones into chromatin, which must be remodeled for gene expression to occur.
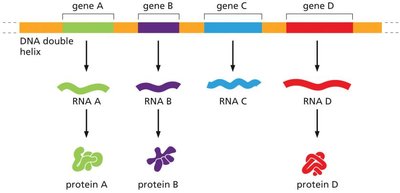
Monocistronic Transcription: Each gene has its own promoter and is transcribed separately.
Compartmentalization: Transcription occurs in the nucleus, translation in the cytoplasm.
mRNA Processing: Eukaryotic mRNAs are spliced, capped, and polyadenylated before export from the nucleus.
Comparison: Prokaryotic vs. Eukaryotic Gene Regulation
The following table summarizes key differences between gene regulation in bacteria and eukaryotes:
Characteristic | Bacterial Gene Control | Eukaryotic Gene Control |
|---|---|---|
Levels of regulation | Primarily transcription | Many levels |
Cascades of gene regulation | Present | Present |
DNA-binding proteins | Important | Important |
Role of chromatin structure | Absent | Important |
Presence of operons | Common | Uncommon |
Negative and positive control | Present | Present |
Initiation of transcription | Relatively simple | Relatively complex |
Enhancers | Less common | More common |
Transcription and translation | Occur simultaneously | Occur separately |
Regulation by small RNAs | Rare | Common |
Chromatin Structure and Gene Regulation
Chromatin structure plays a central role in eukaryotic gene regulation. DNA is wrapped around histone proteins to form nucleosomes, which can be chemically modified to influence gene expression.
Nucleosome: The basic unit of chromatin, consisting of DNA wrapped around an octamer of histone proteins (H2A, H2B, H3, H4).
Histone H1: Stabilizes higher-order chromatin structure but is not part of the core octamer.
Chromatin Remodeling: Chromatin-remodeling complexes reposition nucleosomes to expose regulatory DNA regions for transcription.
Histone Modifications: Acetylation (by HATs), methylation (by methyltransferases), and phosphorylation (by kinases) of histone tails alter chromatin accessibility and gene expression.
Acetylation: Neutralizes positive charge on lysines, loosening chromatin and increasing gene expression.
Deacetylation: Carried out by HDACs, condenses chromatin and represses gene expression.
Epigenetics: Heritable changes in gene expression not involving changes in DNA sequence, often mediated by chromatin modifications.
Example: Flowering in Arabidopsis
Histone acetylation regulates flowering time in Arabidopsis thaliana:
FLC gene: Encodes a transcriptional repressor of flowering. High FLC expression delays flowering.
FLD gene: Encodes a histone deacetylase that represses FLC, promoting earlier flowering.
Mutagenesis Strategies: Mutations that increase FLC delay flowering; mutations that activate FLD promote early flowering.
Transcriptional Regulation in Eukaryotes
Transcriptional regulation is the primary level of gene control in eukaryotes, involving the interplay of DNA regulatory elements and transcription factors.
Regulatory Elements: Promoters, enhancers, silencers, and insulators are DNA sequences that control gene transcription.
General Transcription Factors (GTFs): Required for all RNA polymerase II-mediated transcription; assemble at the promoter (TATA box) to form the preinitiation complex (PIC).
Specific Transcription Factors: Bind to enhancers or silencers to increase or decrease transcription of particular genes.
RNA Polymerase II: The enzyme responsible for transcribing protein-coding genes; its C-terminal domain (CTD) is essential for mRNA processing.
TFIIH: A GTF with kinase and helicase activity; phosphorylates RNA pol II CTD to initiate transcription and helps unwind DNA.
Posttranscriptional Regulation in Eukaryotes
Gene expression is also regulated after transcription through several mechanisms:
Alternative Splicing: Generates multiple mRNA isoforms from a single gene.
mRNA Stability and Degradation: Determines the half-life of mRNAs and thus the level of protein produced.
Noncoding RNAs: Small RNAs (e.g., miRNAs, siRNAs) regulate gene expression posttranscriptionally.
mRNA Localization and Translation Initiation: Control where and when proteins are synthesized.
Posttranslational Modifications: Protein activity can be regulated by chemical modifications after translation.

Summary Table: Levels of Eukaryotic Gene Regulation
Level | Mechanism |
|---|---|
Transcriptional | Chromatin remodeling, transcription factors, promoter/enhancer activity |
RNA Processing | Splicing, capping, polyadenylation |
RNA Transport/Localization | Export from nucleus, localization in cytoplasm |
Translational | Initiation, mRNA stability, noncoding RNAs |
Posttranslational | Protein modification, folding, degradation |
Key Terms and Concepts
Operon: A cluster of genes under the control of a single promoter (common in prokaryotes).
Attenuation: A regulatory mechanism that controls transcription termination in response to metabolite levels.
Chromatin: The complex of DNA and proteins (mainly histones) that forms chromosomes in eukaryotic cells.
Histone Acetylation: Addition of acetyl groups to histone tails, associated with gene activation.
Transcription Factor: A protein that binds to specific DNA sequences to regulate gene expression.
Enhancer: A DNA sequence that increases transcription of a gene, often located far from the gene itself.
Epigenetics: Heritable changes in gene expression that do not involve changes to the DNA sequence.
