 Back
BackGene Regulation in Prokaryotes: Operons, DNA-Binding Proteins, and Mechanisms of Control
Study Guide - Smart Notes

Gene Expression in Prokaryotes
Overview of Gene Expression
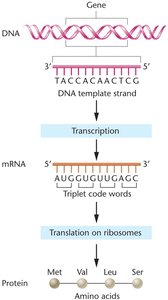
Gene expression is the process by which genetic information is converted from DNA into functional products, such as proteins or RNAs. The regulation of gene expression is essential for all organisms, allowing them to respond to environmental changes and maintain cellular function. In prokaryotes, gene regulation is especially critical for adapting to varying nutrient availability and other environmental factors.
Replication: The genetic material must be faithfully copied for inheritance.
Information Storage: DNA contains complex information necessary for cellular function.
Expression: Genes are expressed to produce phenotypes via transcription and translation.
Variation: Mutations allow genetic diversity and evolution.
Importance of Gene Regulation
Gene regulation ensures that only necessary genes are expressed at any given time.
In bacteria, this allows rapid adaptation to environmental changes by turning genes on or off as needed.
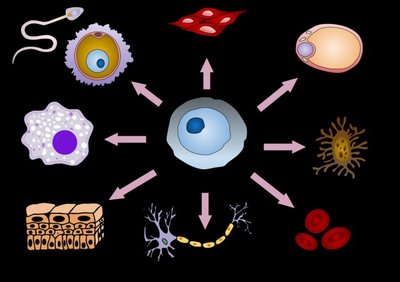
In multicellular eukaryotes, gene regulation is crucial for cell differentiation and specialization, even though all cells contain the same genome.
Levels and Mechanisms of Gene Regulation
Stages of Regulation
Gene expression can be regulated at multiple stages, but transcriptional regulation is often the most significant point of control in both prokaryotes and eukaryotes.
Transcriptional Regulation: Determines when, where, and how much a gene is transcribed.
Regulatory Elements: DNA sequences that control gene expression but are not themselves transcribed.
Regulatory Genes: Encode proteins (regulatory factors) that interact with DNA to influence transcription and translation.
Structural Genes: Encode proteins or RNAs that are not directly involved in regulation.
Types of Gene Expression
Constitutive vs. Regulated Expression
Constitutive Genes: Expressed continuously under normal conditions (e.g., housekeeping genes like actin, tubulin, rRNA).
Regulated Genes: Expression is controlled in response to environmental or cellular signals.
Regulated expression can be further classified as:
Inducible: Genes are usually off but can be turned on (e.g., lac operon).
Repressible: Genes are usually on but can be turned off (e.g., trp operon).
Operons: Coordinated Gene Regulation in Bacteria
Structure and Function of Operons
An operon is a cluster of genes under the control of a single promoter and regulatory elements, allowing coordinated expression. Operons are common in prokaryotes and enable efficient regulation of genes with related functions.
Promoter: Site where RNA polymerase binds to initiate transcription.
Operator: DNA sequence where regulatory proteins (repressors or activators) bind.
Structural Genes: Genes encoding proteins with related functions.
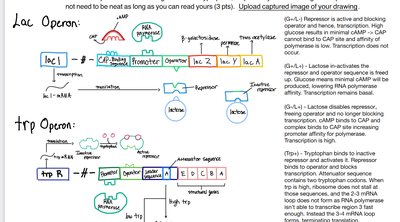
Lac Operon (Inducible System)
The lac operon in Escherichia coli controls the metabolism of lactose. It is an inducible system, meaning it is usually off but can be turned on in the presence of lactose.
In the absence of lactose, a repressor binds to the operator, blocking transcription.
When lactose is present, it binds to the repressor, inactivating it and allowing transcription of the operon.
Catabolite repression: When glucose is present, cAMP levels are low, and the catabolite activator protein (CAP) cannot bind, reducing transcription of the lac operon even if lactose is present.


Trp Operon (Repressible System)
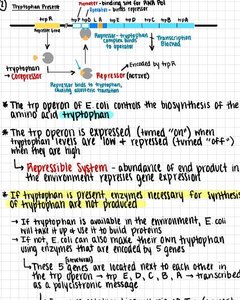
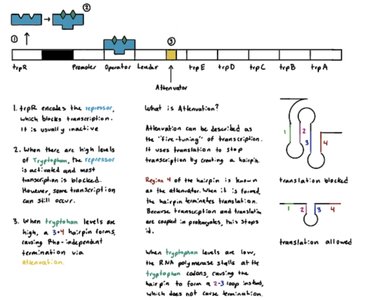
The trp operon in E. coli controls the biosynthesis of tryptophan. It is a repressible system, meaning it is usually on but can be turned off when tryptophan is abundant.
When tryptophan is scarce, the repressor is inactive, and the operon is transcribed.
When tryptophan is abundant, it binds to the repressor, activating it. The active repressor binds to the operator, blocking transcription.
Attenuation: A secondary regulatory mechanism where transcription is prematurely terminated if tryptophan levels are high, further reducing expression of the operon.

Mechanisms of Gene Regulation
Positive and Negative Control
Positive Control: Regulatory protein (activator) binds to DNA and stimulates transcription.
Negative Control: Regulatory protein (repressor) binds to DNA and inhibits transcription.
Operons can be inducible or repressible under either positive or negative control, depending on the regulatory proteins involved.
DNA-Binding Proteins and Motifs
Regulatory proteins often contain specific domains that allow them to bind DNA and influence gene expression.
DNA-binding domain (DBD): Recognizes and binds specific DNA sequences, often through non-covalent interactions such as hydrogen bonds.
Activation domain (AD): Interacts with other proteins to facilitate transcription activation.
Protein motifs: Common structural elements (e.g., helix-turn-helix, zinc finger, leucine zipper) found in DNA-binding proteins.
Most DNA-protein interactions occur in the major groove of the DNA double helix.
Summary Table: Types of Operon Regulation
Type | Regulatory Protein | Gene Expression State | Example |
|---|---|---|---|
Inducible (Negative Control) | Repressor (inactive when inducer present) | Usually off, turned on by inducer | lac operon |
Repressible (Negative Control) | Repressor (active when corepressor present) | Usually on, turned off by corepressor | trp operon |
Inducible (Positive Control) | Activator (active when inducer present) | Usually off, turned on by activator | lac operon (CAP-cAMP) |
Key Terms and Concepts
Operon: Cluster of genes under coordinated control in prokaryotes.
Inducible System: Genes are off unless an inducer is present (e.g., lac operon).
Repressible System: Genes are on unless a corepressor is present (e.g., trp operon).
Attenuation: Regulatory mechanism that causes premature termination of transcription in response to metabolite levels.
DNA-binding Protein: Protein that interacts with specific DNA sequences to regulate gene expression.
Motif: Conserved structural element in proteins, often involved in DNA binding.
