 Back
BackGenetic Analysis and Mapping in Bacteria and Bacteriophages: Study Notes
Study Guide - Smart Notes

Genetic Analysis and Mapping in Bacteria and Bacteriophages
Introduction
This chapter explores the specialized genetic methods used to analyze and map genes in bacteria and bacteriophages. Bacteria serve as powerful model organisms due to their simple genomes, rapid growth, and ease of genetic manipulation. The chapter covers the mechanisms of gene transfer—conjugation, transformation, and transduction—and their applications in genetic mapping and genome evolution.
Specialized Methods for Genetic Analysis of Bacteria
Bacterial Utility in Genetics
Genome simplicity: Bacteria have fewer genes and smaller genomes than eukaryotes, simplifying genetic analysis.
Haploid genomes: Mutations are directly observable since only one gene copy exists per cell.
Short generation times: Bacteria can double in minutes, allowing rapid experimental cycles.
Large progeny numbers: Enables detection of rare genetic events.
Ease of propagation: Bacterial cultures are inexpensive and space-efficient.
Numerous heritable differences: Mutants are easily created, identified, and isolated.
Bacterial Culture and Growth Analysis
Bacteria reproduce by binary fission, producing genetically identical colonies. Growth occurs in liquid or solid media containing essential nutrients.
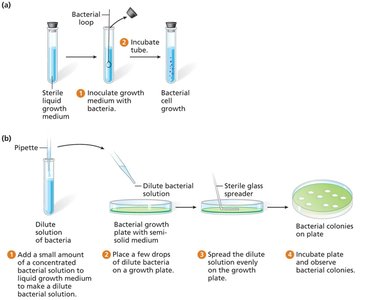
Types of Media
Minimal medium: Contains only essential nutrients (e.g., glucose, nitrogen, water, inorganic salts).
Prototrophs: Bacteria that grow on minimal medium; they lack mutations blocking biosynthetic pathways.
Auxotrophs: Mutant bacteria unable to synthesize a required compound; require complete medium or supplemented minimal medium for growth.
Plating and Replica Plating
Different media are used to determine bacterial genotypes by assessing growth. Replica plating allows identification of auxotrophs by transferring colonies to different media.
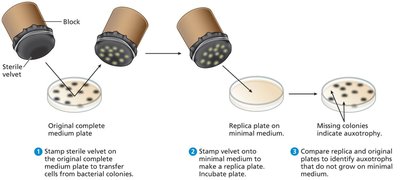
Bacterial Genomes and Plasmids
Characteristics of Bacterial Genomes
Usually a single, covalently closed circular double-stranded DNA molecule.
Small genome size: hundreds of thousands to several million base pairs.
Plasmids in Bacterial Cells
Plasmids: Small, circular, double-stranded DNA molecules carrying nonessential genes.
Types include F (fertility) plasmids (promote gene transfer) and R (resistance) plasmids (carry antibiotic resistance genes).
Plasmids can be high-copy-number (replicate independently) or low-copy-number (depend on chromosome replication).
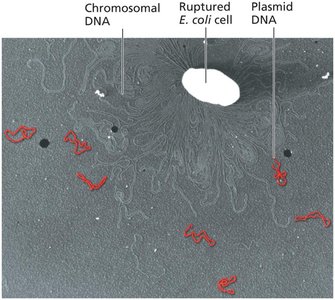
Gene Transfer Mechanisms in Bacteria
Overview of Gene Transfer
Conjugation: Direct transfer of DNA from donor to recipient via cell-to-cell contact.
Transformation: Uptake of free DNA from the environment.
Transduction: Transfer of DNA via bacteriophage (viral) vectors.
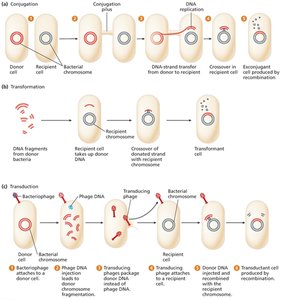
Bacterial Conjugation
F+ (donor) cells: Possess the F factor plasmid; can initiate conjugation.
F− (recipient) cells: Lack the F factor.
Conjugation is mediated by genes on the F plasmid, which direct formation of a conjugation pilus and DNA transfer machinery.
The relaxosome complex nicks the F factor at the origin of transfer (oriT), initiating rolling circle replication and transfer of a single DNA strand to the recipient.
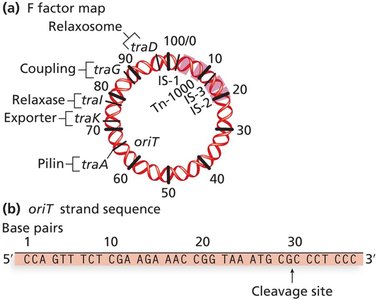
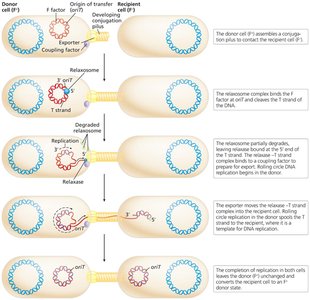
Outcomes of Bacterial Conjugation
Conjugation Type | Exconjugant Becomes Donor? | Bacterial Genes Transferred? |
|---|---|---|
F+ × F− | Yes | No |
Hfr × F− | No | Yes |
F′ × F− | Yes | Yes |
Hfr Strains and Chromosome Mapping
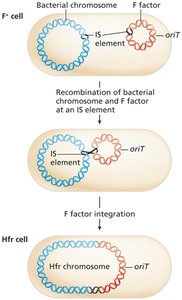
Hfr (High frequency recombination) strains: F factor integrates into the bacterial chromosome at IS elements, allowing transfer of chromosomal genes during conjugation.
Gene transfer occurs in a specific order, starting from oriT.
Complete transfer of the chromosome is rare; only genes near oriT are frequently transferred.
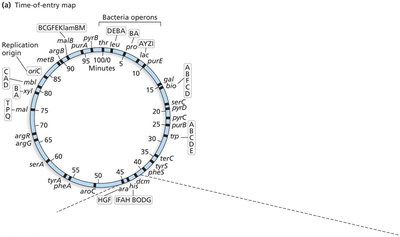
Interrupted Mating and Time-of-Entry Mapping
Interrupted mating: Conjugation is stopped at specific times to map gene order and distance based on the time of entry into the recipient.
Genes closer to oriT enter first and are more frequently incorporated.
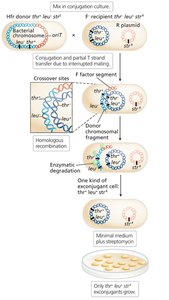
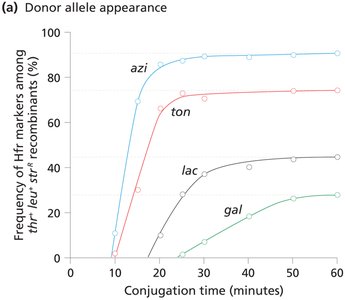
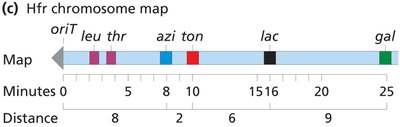
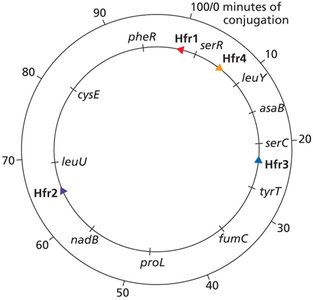
Consolidated Hfr Maps
Multiple Hfr strains are used to map the entire bacterial chromosome, as each strain transfers genes in a unique order and orientation.
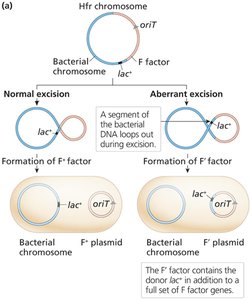
F′ (F prime) Factors and Partial Diploids
F′ factors: Result from imperfect excision of the F factor from the Hfr chromosome, carrying additional bacterial genes.
Conjugation with F′ donors creates partial diploids (merodiploids), useful for studying gene regulation and dominance.
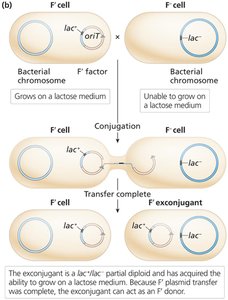
Bacterial Transformation
Mechanism and Steps
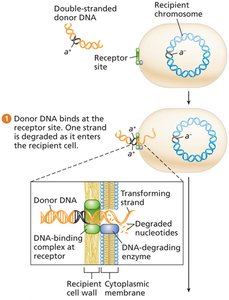
Transformation: Uptake of free DNA fragments from the environment by competent bacterial cells.
DNA binds to the cell surface, one strand is degraded, and the other integrates into the chromosome by homologous recombination, forming a heteroduplex.
After replication, one daughter cell is a transformant.
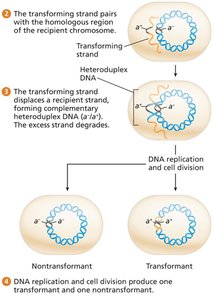
Mapping by Transformation
Transformation is most effective for mapping closely linked genes (cotransformation).
DNA fragments are typically less than 100 kb, so only genes close together are transferred together.
Bacterial Transduction
Mechanism and Bacteriophage Life Cycles
Transduction: Transfer of bacterial DNA by bacteriophages (viruses that infect bacteria).
Bacteriophages have a head (containing DNA), sheath, and tail fibers.
Two life cycles: Lytic (phage replicates, lyses host) and Lysogenic (phage integrates as a prophage, can later enter lytic cycle).
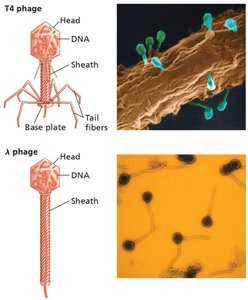
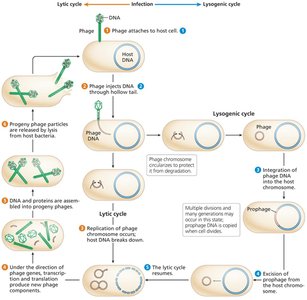
Generalized Transduction
Random fragments of bacterial DNA are mistakenly packaged into phage heads during lytic infection.
These generalized transducing phages can inject donor DNA into new recipients, where homologous recombination may occur.
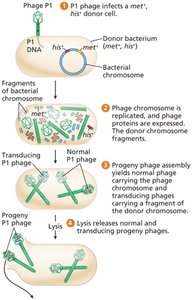
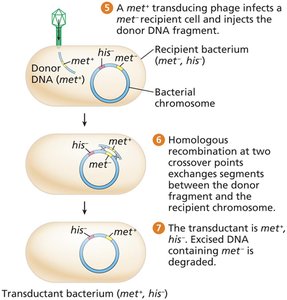
Cotransduction and Mapping
Cotransduction: Simultaneous transfer of two or more genes by a single transducing phage; frequency depends on gene proximity.
Used to determine gene order and distance.
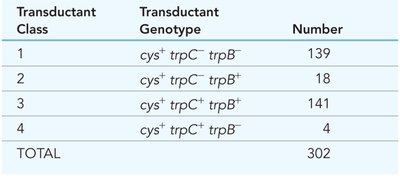
Transductant Class | Transductant Genotype | Number |
|---|---|---|
1 | cys+ trpC− trpB− | 139 |
2 | cys+ trpC− trpB+ | 18 |
3 | cys+ trpC+ trpB− | 141 |
4 | cys+ trpC+ trpB+ | 4 |
TOTAL | 302 |
Specialized Transduction
Occurs when temperate phages excise imprecisely from the host genome, carrying adjacent bacterial genes.
Only genes near the phage integration site (att site) are transferred.
Lateral Gene Transfer (LGT) and Genome Evolution
LGT in Bacteria and Archaea
Lateral gene transfer (LGT): Movement of genetic material between unrelated organisms, contributing to genome diversity and evolution.
LGT can occur between bacteria, archaea, and even eukaryotes (e.g., endosymbiosis, Agrobacterium transfer to plants).
Genomic islands—large, unique DNA regions—are evidence of LGT, often containing genes for pathogenicity or antibiotic resistance.
Medical Importance of LGT
LGT enables rapid adaptation, such as acquisition of antibiotic resistance or virulence factors.
Pathogenicity islands acquired by LGT can turn harmless bacteria into disease-causing strains.
Summary Table: Key Terms and Concepts
Term | Definition |
|---|---|
Prototroph | Bacterium able to grow on minimal medium |
Auxotroph | Mutant unable to synthesize a required compound |
Conjugation | Direct DNA transfer between bacteria via cell contact |
Transformation | Uptake of free DNA from the environment |
Transduction | DNA transfer via bacteriophage |
Hfr | High frequency recombination strain with integrated F factor |
F′ (F prime) | F factor carrying additional bacterial genes |
Lateral Gene Transfer | Transfer of genes between unrelated organisms |
