 Back
BackGenetic Code, Transcription, and Translation: Study Notes for Genetics Students
Study Guide - Smart Notes

Chapter 13: The Genetic Code and Transcription
Central Dogma of Molecular Genetics
The central dogma describes the directional flow of genetic information: DNA → RNA → Protein. This process is fundamental to gene expression, which involves transcription (DNA to RNA) and translation (RNA to protein).
Features of the Genetic Code
The genetic code is the set of rules by which information encoded in mRNA is translated into proteins. It is written in linear form using ribonucleotide bases and is composed of triplet codons.
Codon: Sequence of three ribonucleotides in mRNA that specifies an amino acid.
Unambiguous: Each codon specifies only one amino acid.
Degenerate: Most amino acids are specified by more than one codon.
Start and Stop Codons: Specific codons initiate (AUG) and terminate (UAG, UAA, UGA) translation.
Commaless: Codons are read sequentially without breaks.
Nonoverlapping: Each nucleotide is part of only one codon.
Colinear: The sequence of codons in a gene is colinear with the amino acid sequence in the protein.
Nearly Universal: The code is used by most organisms, with few exceptions.
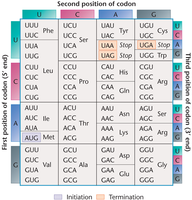
Triplet Binding Assay and Deciphering the Code
Early studies, such as the triplet binding assay by Nirenberg and Leder, established the operational patterns of the code. Ribosomes bind to single codons, and complementary charged tRNAs bind, allowing identification of codon assignments.
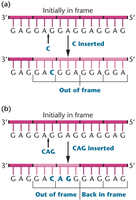
Reading Frame and Frameshift Mutations
The reading frame is a contiguous sequence of nucleotides. Insertions or deletions shift the reading frame, causing frameshift mutations that alter downstream codons.
Degeneracy and Order in the Genetic Code
The genetic code is degenerate, meaning multiple codons can specify the same amino acid. Only tryptophan and methionine are encoded by a single codon. Chemically similar amino acids often share similar codons.

Using the Codon Chart
Students should be able to use the codon chart to determine amino acid sequences from mRNA templates. For example:
mRNA: 5’-AUG GCA AAA AAG UGA-3’ → Met-Ala-Lys-Lys
mRNA: 5’-ACU UCU UCA UGA-3’ → Thr-Ser-Ser

The Wobble Hypothesis
The wobble hypothesis states that the third base in a codon is often flexible, allowing non-standard base pairing. The first two bases are more critical for specifying the amino acid.
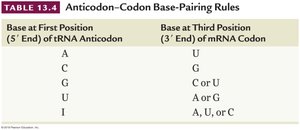
Base at First Position (5' End) of tRNA Anticodon | Base at Third Position (3' End) of mRNA Codon |
|---|---|
A | U |
C | G |
G | C or U |
U | A or G |
I | A, U, or C |
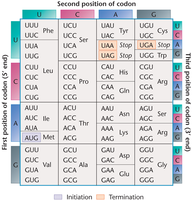
Initiation and Termination Codons
Methionine (AUG) is the initiator codon, and termination codons (UAG, UAA, UGA) signal the end of translation. Termination codons are not recognized by tRNA and do not code for any amino acid.
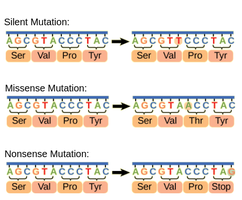
Types of Point Mutations
Single nucleotide changes can have different effects:
Silent Mutation: Changes nucleotide but not the amino acid.
Missense Mutation: Changes nucleotide and amino acid, possibly reducing protein function.
Nonsense Mutation: Changes nucleotide to a stop codon, terminating translation early.
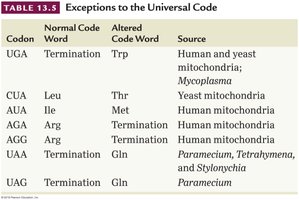
Exceptions to the Universal Genetic Code
Some organisms and organelles (e.g., mitochondria) have exceptions to the universal genetic code. For example, UGA codes for tryptophan in human mitochondria instead of termination.
Codon | Normal Code Word | Altered Code Word | Source |
|---|---|---|---|
UGA | Termination | Trp | Human and yeast mitochondria; Mycoplasma |
CUA | Leu | Thr | Yeast mitochondria |
AUA | Ile | Met | Human mitochondria |
AGA | Arg | Termination | Human mitochondria |
AGG | Arg | Termination | Human mitochondria |
UAA | Termination | Gln | Paramecium, Tetrahymena, and Stylonichia |
UAG | Termination | Gln | Paramecium |
Chapter 14: Translation and Proteins
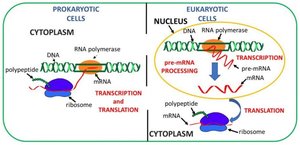
Translation Depends on Ribosomes and tRNA
Translation is the biological polymerization of amino acids into polypeptide chains. It requires amino acids, mRNA, ribosomes, and tRNA. In prokaryotes, transcription and translation occur in the cytoplasm, while in eukaryotes, transcription occurs in the nucleus and translation in the cytoplasm.
Ribosomes: Site of Translation
Ribosomes are essential for gene expression, consisting of ribosomal proteins and rRNAs. They have large and small subunits. Prokaryotic ribosomes are 70S, eukaryotic ribosomes are 80S.
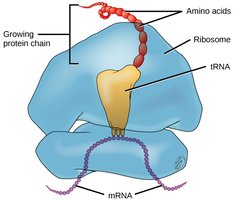
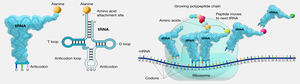
tRNA: Adaptor Molecule
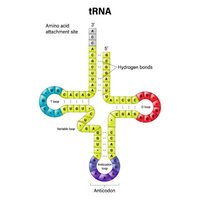
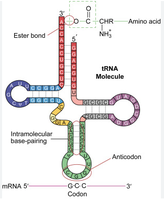
tRNAs are small, stable molecules transcribed from DNA. They have a cloverleaf structure due to hydrogen bonding. tRNAs adapt genetic information from mRNA codons to corresponding amino acids. The anticodon region of tRNA base-pairs with the codon in mRNA, and the amino acid is covalently linked to the 3’ end.
tRNA Charging
Before translation, tRNAs must be charged with their respective amino acids by aminoacyl tRNA synthetases. There are 20 synthetases, each specific for one amino acid.
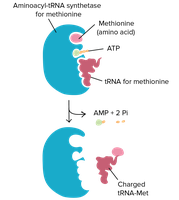
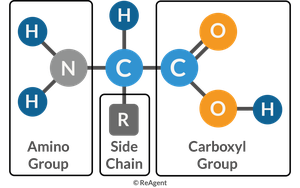
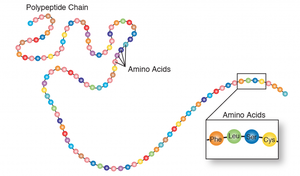
Basic Structure of Amino Acids
Amino acids have a central carbon atom bonded to a carboxyl group, amino group, and an R group (side chain). The R group determines the type of amino acid: nonpolar, polar, positively charged, or negatively charged.
Peptide Bond Formation
Amino acids are linked by peptide bonds, formed by dehydration reactions between the carboxyl group of one amino acid and the amino group of another. Two amino acids form a dipeptide, three a tripeptide, and so on.
Steps of Translation
Translation is divided into three steps:
Initiation: Ribosome assembles on mRNA, and the first tRNA moves into place.
Elongation: Ribosome moves along mRNA, tRNAs read codons, and amino acids are added to the growing chain.
Termination: Stop codon is reached, prompting release of the completed polypeptide and ribosome disassembly.
Translation in Eukaryotes vs Prokaryotes
Eukaryotic translation is more complex, involving larger ribosomes, RNA processing (5’ cap and poly-A tail), and often occurs on endoplasmic reticulum-bound ribosomes. The Kozak sequence increases efficiency of translation initiation in eukaryotes.
One Gene: One Polypeptide Hypothesis
Each gene encodes a single polypeptide. Hemoglobin is a tetrameric protein with two alpha and two beta chains, each containing a heme group for oxygen binding. Mutations in hemoglobin genes can cause disorders such as sickle-cell anemia.
Protein Structure and Diversity
Proteins are polypeptides folded into functional three-dimensional conformations. There are four levels of protein structure:
Primary: Sequence of amino acids.
Secondary: Alpha helices and beta sheets.
Tertiary: Three-dimensional shape.
Quaternary: Composed of more than one polypeptide.
Posttranslational Modifications
Polypeptide chains are often modified after synthesis, which is crucial for the functional capability of the final protein product.
Proteins: Structure and Function
Proteins perform diverse roles, including immunity, hormone regulation, muscle contraction, and gene expression regulation. Their structural diversity underlies biological complexity.
