 Back
BackGenetic Code, Translation, and Protein Structure: Study Notes
Study Guide - Smart Notes

The Genetic Code
Degeneracy of the Genetic Code
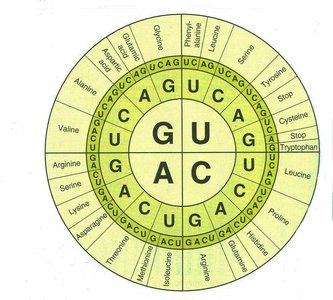
The genetic code is described as degenerate because most amino acids are encoded by more than one codon. This redundancy is primarily due to variations in the third base of the codon, while the first and second bases are more critical for specifying the amino acid.
Partial degeneracy: The third base can be substituted within the same chemical class (purine to purine, pyrimidine to pyrimidine). Example: G <-> A, C <-> U.
Complete degeneracy: Any of the four bases can occupy the third position. Example: GUU, GUC, GUA, and GUG all code for valine.
Biological significance: Degeneracy reduces the impact of mutations, allowing for silent mutations that do not alter the amino acid sequence, thus increasing genetic stability.
tRNA adaptation: Multiple tRNAs may exist for a single amino acid, or a single tRNA anticodon may pair with several codons due to flexible base pairing at the third codon position.
Wobble Hypothesis
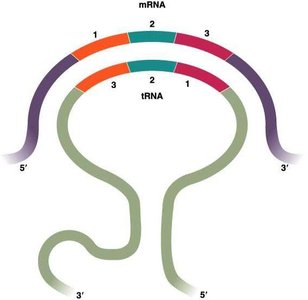
The wobble hypothesis explains how a single tRNA can recognize multiple codons for the same amino acid. Wobble occurs at the first (5') base of the tRNA anticodon and the third (3') base of the mRNA codon, allowing non-standard base pairing.
Hydrogen bonding: The first two bases of the codon and the last two of the anticodon pair tightly, while the third position is more flexible.
Result: This flexibility allows fewer tRNAs to cover all codons, contributing to the efficiency of translation.
Translation: Protein Synthesis
Overview of Translation
Translation is the process by which the genetic information encoded in mRNA is used to assemble a specific sequence of amino acids, forming a polypeptide chain. This process occurs in the ribosome and involves three main stages: initiation, elongation, and termination.
Initiation: Assembly of the ribosome on the mRNA and recruitment of the initiator tRNA.
Elongation: Sequential addition of amino acids to the growing polypeptide chain.
Termination: Release of the completed polypeptide upon encountering a stop codon.
Ribosome Structure and Function
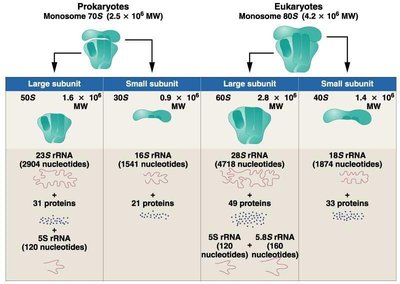
Ribosomes are complex molecular machines composed of rRNA and proteins. They consist of two subunits (large and small) that differ between prokaryotes and eukaryotes.
Prokaryotes | Eukaryotes | |
|---|---|---|
Monosome | 70S (50S + 30S) | 80S (60S + 40S) |
Large Subunit | 23S rRNA, 5S rRNA, 31 proteins | 28S rRNA, 5.8S rRNA, 5S rRNA, 49 proteins |
Small Subunit | 16S rRNA, 21 proteins | 18S rRNA, 33 proteins |
tRNA Structure and Function
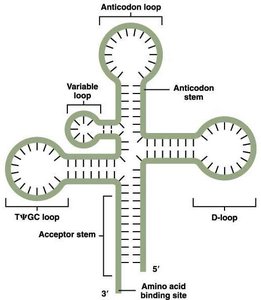
Transfer RNA (tRNA) molecules serve as adaptors that translate the nucleotide sequence of mRNA into the amino acid sequence of proteins. Each tRNA has a specific anticodon that pairs with a codon on the mRNA and an acceptor stem for amino acid attachment.
Anticodon loop: Contains the three-nucleotide sequence complementary to the mRNA codon.
Amino acid binding site: Located at the 3' end of the tRNA.
Other loops: D-loop, TΨC loop, and variable loop contribute to tRNA stability and recognition by aminoacyl-tRNA synthetases.
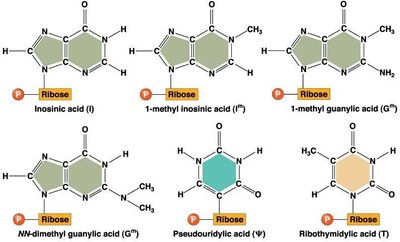
Modified Bases in tRNA
tRNAs contain several unusual or modified bases that are not found in standard RNA. These modifications contribute to the proper folding, stability, and function of tRNA molecules.
Examples: Inosinic acid (I), pseudouridylic acid (Ψ), ribothymidylic acid (T), and various methylated bases.
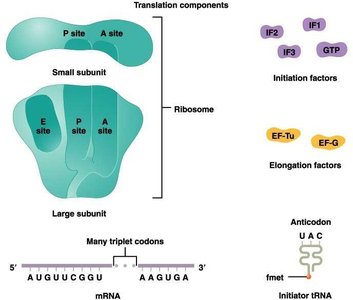
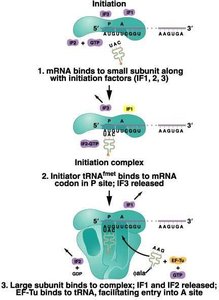
Translation Initiation
Initiation involves the assembly of the ribosome on the mRNA, the recruitment of the initiator tRNA, and the formation of the initiation complex. In prokaryotes, the first amino acid is N-formyl methionine (fMet), while in eukaryotes it is methionine (Met).
Initiation factors (IFs): Proteins that assist in the assembly of the initiation complex.
P site: Peptidyl site, where the initiator tRNA binds.
A site: Aminoacyl site, where the next charged tRNA enters.
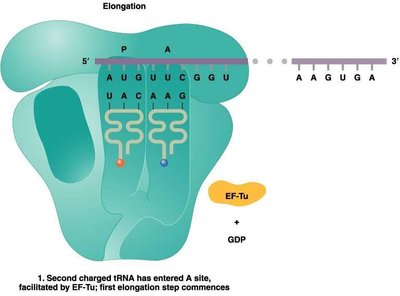
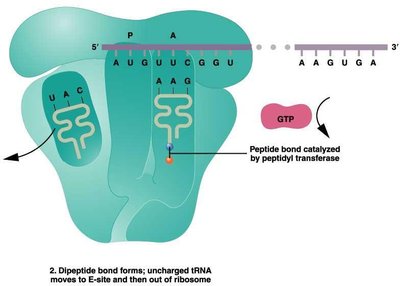
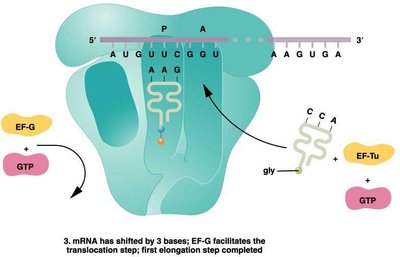
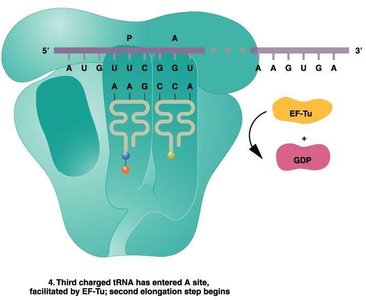
Translation Elongation
During elongation, amino acids are sequentially added to the growing polypeptide chain. This process involves the entry of charged tRNAs into the A site, peptide bond formation, and translocation of the ribosome along the mRNA.
Elongation factors (EFs): Proteins that facilitate tRNA entry and ribosome movement.
Peptidyl transferase: Enzyme activity of the ribosome that forms peptide bonds.
Translocation: Ribosome shifts by one codon, moving the tRNA from the A site to the P site.
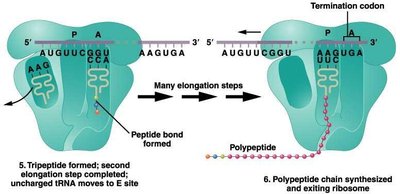
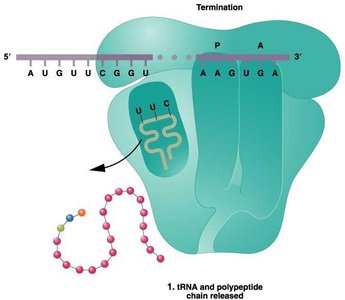
Translation Termination
Termination occurs when a stop codon is encountered in the A site. Release factors bind to the ribosome, prompting the release of the polypeptide chain and dissociation of the ribosomal subunits.
Comparison of Prokaryotic and Eukaryotic Translation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Location | Free ribosomes | Free and ER-bound ribosomes |
Coupling | Transcription and translation coupled | Separate |
mRNA type | Polygenic | Monogenic |
First amino acid | N-formyl methionine (fMet) | Methionine (Met) |
Ribosome subunits | 50S, 30S | 60S, 40S |
Protein Structure and Function
Levels of Protein Structure
Proteins have four levels of structural organization, each contributing to their function and stability.
Primary structure: Linear sequence of amino acids.
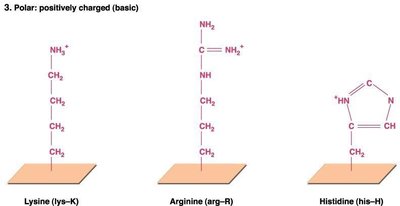
Secondary structure: Local folding into α-helices and β-pleated sheets, stabilized by hydrogen bonds.
Tertiary structure: Three-dimensional conformation of a single polypeptide chain.
Quaternary structure: Association of multiple polypeptide chains.
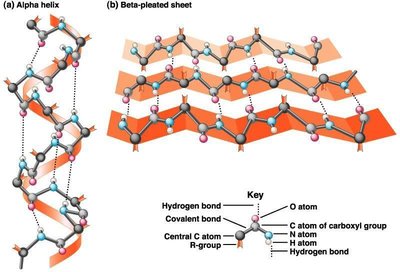
Diversity and Functions of Proteins
Proteins perform a wide variety of functions in the cell, including:
Immunoglobulins: Immune defense in vertebrates.
Transport proteins: Movement of molecules across membranes.
Hormones and receptors: Regulation of physiological processes.
Histones: DNA packaging in eukaryotes.
Transcription factors: Regulation of gene expression.
Enzymes: Catalysis of biochemical reactions, determining metabolic capacity.
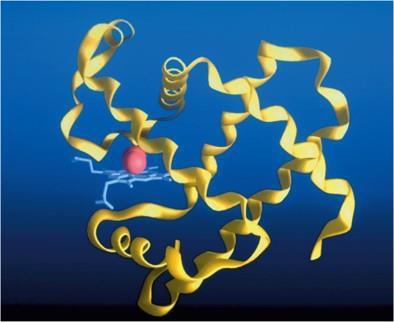
Sickle-Cell Anemia: A Protein Mutation Example
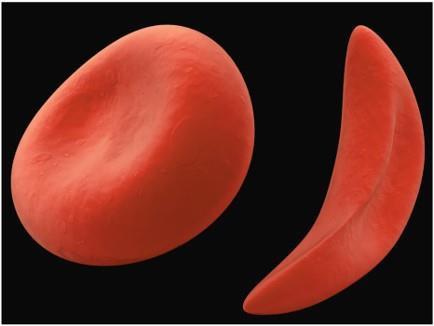
Sickle-cell anemia is a recessive genetic disease caused by a single amino acid substitution (glutamic acid to valine) in the β chain of hemoglobin. This mutation leads to abnormal hemoglobin structure and function, causing red blood cells to become elongated under low oxygen conditions.
Homozygotes: Exhibit disease symptoms.
Heterozygotes: Carriers, usually asymptomatic.
Protein Folding and Disease
Proper protein folding is essential for function. Misfolded proteins can cause diseases such as prion diseases (scrapie, mad cow disease, Creutzfeldt–Jakob disease) and neurodegenerative disorders (Huntington, Alzheimer, Parkinson).
Prions: Infectious misfolded proteins that induce abnormal folding in normal proteins.
Protein aggregates: Linked to neurodegenerative diseases.
Regulation of Gene Expression
Levels and Points of Control
Gene expression is regulated at multiple levels, from chromatin structure to post-translational modification of proteins. This regulation ensures that genes are expressed at the right time, place, and amount.
Chromosome level: Chromatin condensation, gene amplification, and gene activation/inactivation.
Transcriptional control: Activation or repression of gene transcription, mRNA processing, and transport.
Post-transcriptional control: mRNA stability, translation initiation, and mRNA masking.
Post-translational control: Protein modification, storage, secretion, and degradation.
Examples: Inactivation of the X chromosome in female mammals, gene amplification in oocytes, and differential mRNA half-lives in various cell types.
Summary Table: Key Differences in Translation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
First amino acid | N-formyl methionine (fMet) | Methionine (Met) |
Ribosome subunits | 50S, 30S | 60S, 40S |
mRNA type | Polygenic | Monogenic |
Location | Cytoplasm | Cytoplasm and ER |
