Back
BackGenetic Linkage and Mapping in Eukaryotes: Study Guide
Study Guide - Smart Notes

Genetic Linkage and Mapping in Eukaryotes
Overview of Mapping
Mapping is a fundamental technique in genetics used to determine the position of gene loci on a chromosome. Linked genes are genes that are inherited together because they reside on the same chromosome, which acts as the unit of inheritance.
Linked genes do not undergo independent assortment.
Complete linkage describes genes that are always inherited together.
In a heterozygous cross, the F2 ratio is 1:2:1; in a test cross, it is 1:1.
Example: Linkage groups are sets of genes that tend to be inherited together due to their physical proximity on the chromosome.
Crossing Over and Genetic Recombination
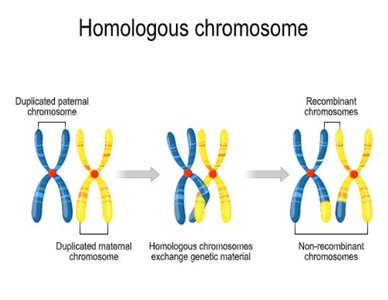
Crossing over is the physical breaking and rejoining of homologous chromosomes during meiosis, resulting in genetic recombination and new combinations of alleles. This process explains why genes on the same chromosome may not always be inherited together.
Occurs during meiosis.
Produces recombinant gametes.
Explains the inheritance patterns of linked genes.
Chromosomal Mapping and Recombination Frequency
Chromosomal mapping is performed by analyzing the frequency of recombination between gene loci. The closer two genes are, the less likely they are to cross over; the farther apart, the more likely.
Recombination frequency is used to determine the relative location of genes.
Map units (m.u.) are used to express genetic distances: 1 m.u. = 1% recombination frequency.
Example: If two genes have a recombination frequency of 10.7%, they are 10.7 map units apart.
Crossing Over Terminology
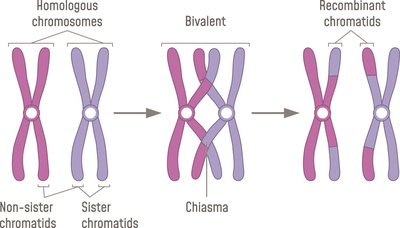
Crossing over occurs between two chromatids, typically non-sister chromatids from homologous chromosomes. During meiosis, homologous pairs form tetrads and undergo crossing over at chiasmata.
Tetrads: Four paired chromatids.
Dyads: Pair of two chromatids.
Bivalent: Pair of homologous chromosomes.
Chiasmata: Structure formed during crossing over.
Linked Genes: Cis and Trans Conformations
Linked genes can be arranged in cis or trans conformations:
Cis conformation: Dominant alleles of two genes are on the same chromosome (e.g., AB/ab).
Trans conformation: Different alleles of two genes are on the same chromosome (e.g., Ab/aB).
Alleles on the same homolog are written together without punctuation; the “/” separates homologs.
Mapping Genes Using Recombination Frequencies
Measuring recombination is the traditional method for mapping gene loci. Recombination frequency is calculated as:
Physical distance is directly correlated with recombination frequency.
Linkage is likely if recombination frequency is less than 50%.
Recombination frequencies are never greater than 50% due to independent assortment.
Modern Mapping Techniques
Gene loci can be mapped using recombination maps, physical maps, and genomic markers:
Recombination maps: Use recombination frequencies.
Physical maps: Use genomic sequencing.
Genomic markers: Include SNPs, RFLPs, and microsatellites.
Trihybrid Crosses for Mapping
A trihybrid cross examines the inheritance of three traits. For linked genes, recombination frequencies in offspring are used to map all three loci.
Calculate recombination frequencies for each pair of genes.
Double crossovers must be counted twice for accurate mapping.
Multiple Crossovers and Interference
Mapping three or more genes can be complicated by multiple crossovers. Interference occurs when crossovers in one region affect the likelihood of crossovers in adjacent regions.
Coefficient of coincidence: Used to measure interference.
Interference is calculated as:
Example: If the observed frequency is 4 and expected is 6, then or 33% interference.
Chi-Square Test for Linkage
The chi-square test is used to evaluate whether observed experimental values differ from predicted values, helping to determine the likelihood of gene linkage.
Null hypothesis: Expected and observed values are not different (genes are not linked).
Degrees of freedom (df) are calculated based on the number of phenotypic classes minus one.
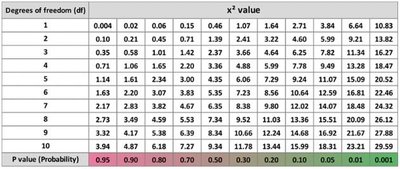
Chi-square value is compared to a table to determine the p-value.
If p > 0.05, accept the null hypothesis; if p < 0.05, reject it.
Practice Problems and Applications
Practice problems are provided throughout the notes to reinforce concepts such as calculating recombination frequencies, determining gene order, and applying the chi-square test.
Calculate recombination frequency from offspring data.
Determine gene order based on genetic distances.
Apply chi-square test to experimental results.
Example: Given offspring numbers for different phenotypes, calculate recombination frequency and use chi-square analysis to test for linkage.
Summary Table: Chi-Square Values
The chi-square value table is essential for interpreting statistical results in genetic linkage studies.