 Back
BackLecture 6
Study Guide - Smart Notes

Genetic Linkage and Recombination
Introduction to Linkage and Recombination
Genetic linkage and recombination are fundamental concepts in genetics that explain how genes located on the same chromosome can be inherited together, and how crossing-over during meiosis can generate new allele combinations. These principles modify the expectations from simple Mendelian inheritance and are essential for understanding genetic mapping.
Modifications to Mendelian Inheritance
Mendel's experiments with pea plants established the basic laws of inheritance, but most traits and genes behave in more complex ways:
Sex linkage: Genes located on sex chromosomes show unique inheritance patterns.
Incomplete dominance and codominance: Phenotypes may not show complete dominance.
Multiple alleles, lethal alleles, conditional alleles: More than two alleles or special conditions can affect inheritance.
Penetrance and expressivity: Not all individuals with a genotype express the expected phenotype.
Gene interaction: Multiple genes can affect a single trait.
Genetic linkage: Genes on the same chromosome may not assort independently.
Mendel’s Dihybrid Cross and Independent Assortment
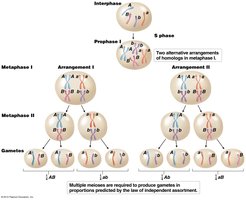
Mendel’s dihybrid crosses demonstrated the law of independent assortment, where genes on different chromosomes segregate independently during gamete formation. This results in a 9:3:3:1 ratio in the F2 generation for two traits.
Law of Independent Assortment: The two copies of a gene for one trait segregate independently from the two copies of a gene for another trait.
Chromosome Theory: Mendel’s factors are located on chromosomes, which segregate randomly during meiosis.
Genetic Linkage: Evidence and Early Experiments
Not all genes assort independently. Genes located close together on the same chromosome are said to be genetically linked. Early experiments by Bateson, Punnet, and Morgan provided evidence for linkage:
Bateson and Punnet (1905): Dihybrid crosses in sweet peas showed more parental phenotypes than expected, indicating linkage.
Morgan (1911): Studies in Drosophila revealed linkage between X-linked traits, with more parental combinations than predicted by independent assortment.
Linkage and Recombination: Mechanisms
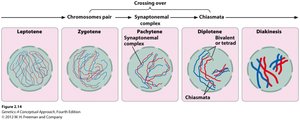
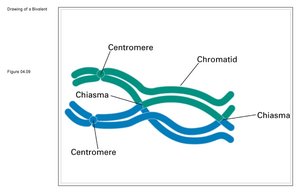
Genes on the same chromosome can be separated by recombination (crossing-over) during meiosis. The physical exchange of chromosome segments occurs at chiasmata during prophase I, generating recombinant gametes.
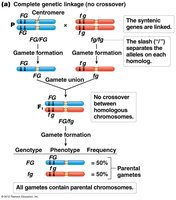
Complete linkage: No recombination occurs; only parental gametes are produced.
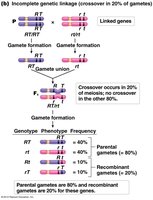
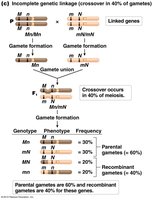
Incomplete linkage: Some recombination occurs, producing both parental and recombinant gametes.
Recombination frequency: The proportion of recombinant gametes indicates the distance between genes.
Linkage Analysis: Gamete Types and Frequencies
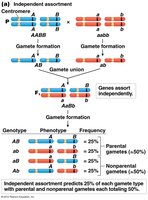
Linkage affects the types and frequencies of gametes produced:
Unlinked genes: 50% parental, 50% recombinant gametes.
Completely linked genes: 100% parental gametes.
Partially linked genes: Intermediate frequencies, depending on recombination rate.
Genetic Mapping: Recombination Frequency and Map Units
The frequency of recombination between two genes is used to estimate their distance on a chromosome. Genetic distances are measured in map units (m.u.) or centiMorgans (cM):
1% recombination = 1 map unit = 1 centiMorgan
Formula:
Double Crossovers and Map Construction
At longer distances, double crossovers can occur, which may reverse the effect of the first crossover and lead to an underestimate of the genetic distance. Genetic maps are most accurate when constructed using small intervals.
Double crossover: Two recombination events between genes can restore parental combinations.
Map construction: Use small intervals to minimize errors from double crossovers.
Historical Development: The First Genetic Map
Alfred Sturtevant, working in Morgan’s lab, created the first genetic map by analyzing recombination frequencies between multiple genes on the X chromosome of Drosophila. He demonstrated that genes are arranged linearly on chromosomes, each with a defined location.
Gene pairs and recombination frequencies:
Gene Pair | # Recombinants/Total | % Recombination |
|---|---|---|
y and w | 214/21,736 | 1.0% |
y and v | 1,464/4,551 | 32.2% |
y and m | 115/324 | 35.5% |
y and r | 260/693 | 37.5% |
w and v | 471/1,584 | 29.7% |
w and m | 2,062/6,116 | 33.7% |
w and r | 406/898 | 45.2% |
v and m | 17/573 | 3.0% |
v and r | 109/405 | 26.9% |
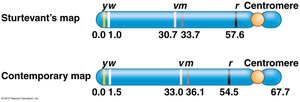

Applications and Modern Genetic Maps
Genetic maps based on recombination frequencies were the foundation of genetics before the molecular nature of genes was understood. Today, genetic mapping is used to locate genes, study inheritance, and understand genome structure in humans, plants, and animals.


Summary Table: Linkage and Recombination Types
Type | Gamete Frequency | Example |
|---|---|---|
Independent Assortment | 50% parental, 50% recombinant | Genes on different chromosomes |
Complete Linkage | 100% parental | Genes very close on same chromosome |
Incomplete Linkage | Intermediate (e.g., 60% parental, 40% recombinant) | Genes moderately apart on same chromosome |
Key Terms and Definitions
Genetic linkage: The tendency of genes located close together on a chromosome to be inherited together.
Recombination: The process by which chromosomes exchange genetic material during meiosis, producing new allele combinations.
Chiasma: The physical site of crossing-over between homologous chromosomes.
Map unit (m.u.)/centiMorgan (cM): A unit of genetic distance corresponding to 1% recombination frequency.
Example: If two genes show a recombination frequency of 25%, they are 25 map units (cM) apart on the chromosome.
Additional info: Double crossovers can lead to an underestimate of genetic distance; modern mapping uses molecular markers and DNA sequencing for greater accuracy.
