 Back
BackGenetic Mechanisms: Double-Strand Break Repair, Transposable Elements, and Regulation of Gene Expression
Study Guide - Smart Notes

Double-Strand Break Repair
Overview of Double-Strand Breaks
Double-strand breaks (DSBs) in DNA are among the most severe forms of genetic damage. If unrepaired, they can lead to chromosomal rearrangements, cancer, or cell death. Cells have evolved two primary pathways to repair DSBs: homologous recombination repair and nonhomologous end joining.
Homologous recombination repair: Uses a homologous sequence as a template for accurate repair.
Nonhomologous end joining: Directly ligates the broken DNA ends, often resulting in mutations.
Homologous Recombination Repair
Homologous recombination repair is a high-fidelity mechanism that utilizes a sister chromatid as a template. It is most active during late S or early G2 phase of the cell cycle.
Recognition and processing: The break is recognized, and the 5' ends are digested, leaving 3' overhangs.
Strand invasion: The 3' overhang aligns with a complementary sequence on the sister chromatid.
DNA synthesis: DNA polymerase synthesizes new DNA using the sister chromatid as a template.
Resolution: The newly synthesized DNA is ligated, restoring the integrity of the chromosome.
Example: Homologous recombination is essential for maintaining genome stability and is used during meiosis for crossing over. 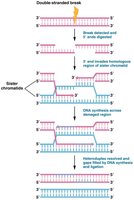
Nonhomologous End Joining (NHEJ)
Nonhomologous end joining is a more error-prone repair mechanism, active primarily in G1 phase before DNA replication.
Protein complex formation: Proteins, including kinases and BRCA1, bind to the free DNA ends.
End processing: The ends are processed and ligated together, sometimes resulting in loss or addition of nucleotides.
Outcome: NHEJ can restore DNA integrity but may introduce mutations at the repair site.
Example: NHEJ is crucial for immune system diversity (V(D)J recombination) but can contribute to genomic instability. 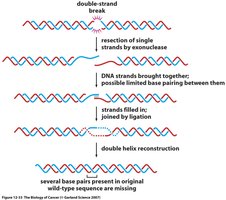
Transposable Elements
Definition and Types
Transposable elements (TEs), also known as "jumping genes," are DNA sequences that can move within and between chromosomes. They are found in all organisms and contribute to genetic diversity and evolution.
Insertion sequences (IS elements): Simple transposons that cause mutations by inserting into genes or regulatory regions.
Bacterial transposons: Larger elements that can carry genes, such as those conferring antibiotic resistance.
Structure of Transposons
Transposons typically contain inverted terminal repeats (ITRs) and direct repeats (DRs) flanking a transposase gene.
ITRs: Short, inverted sequences at each end of the transposon.
DRs: Direct repeats generated during insertion.
Transposase: Enzyme that catalyzes the movement of the transposon.
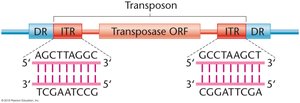
Mechanism of Transposition
The process of transposition involves several steps:
Transposase binding to ITRs
Formation of a transposition complex
Excision of the transposon
Recognition of the target site
Insertion into the new site
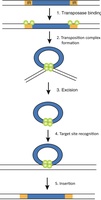
Transposon Insertion and Target Site Duplication
During insertion, transposase makes staggered cuts at the target site, inserts the transposon, and DNA polymerase fills in gaps, creating new direct repeats. 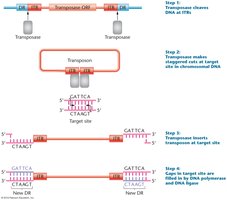
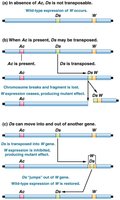
Ac-Ds System in Maize
The Ac-Ds system in maize illustrates autonomous and nonautonomous transposable elements.
Ac (Activator): Autonomous element that produces transposase and can move independently.
Ds (Dissociation): Nonautonomous element that requires Ac for movement.
Phenotypic effects: Insertion of Ds into the C gene disrupts anthocyanin production, resulting in colorless kernels. Removal of Ds by Ac restores color, causing spotted kernels.
Retrotransposons
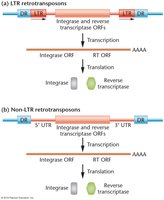
Retrotransposons are a class of TEs that move via an RNA intermediate.
LTR retrotransposons: Contain long terminal repeats and encode integrase and reverse transcriptase.
Non-LTR retrotransposons: Lack LTRs but also encode integrase and reverse transcriptase.
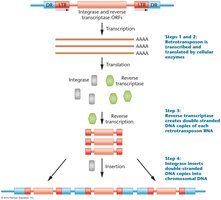
Mechanism of Retrotransposition
Transcription of retrotransposon DNA to RNA
Translation of RNA to produce reverse transcriptase and integrase
Reverse transcriptase synthesizes DNA from RNA
Integrase inserts the new DNA copy into the genome
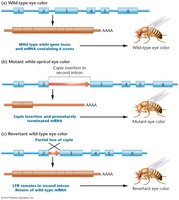
Copia Elements in Drosophila
Copia elements are retrotransposons found in Drosophila, characterized by direct terminal repeats (DTRs) and inverted terminal repeats (ITRs).
Dispersed throughout the genome and can cause mutations by insertion.

Regulation of Gene Expression in Eukaryotes
Levels of Regulation
Gene expression in eukaryotes is regulated at multiple levels, including chromatin remodeling, transcription, RNA processing, translation, and post-translational modification.
Chromatin remodeling: Alters DNA accessibility for transcription.
Transcriptional regulation: Controls initiation and rate of transcription.
Post-transcriptional regulation: Includes mRNA splicing, stability, and translation.
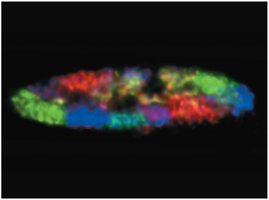
Chromatin Structure and Modification
Chromosome territory: Each chromosome occupies a distinct domain in the nucleus.
Transcription factories: Nuclear sites with concentrated RNA polymerase and regulatory molecules.
Histone modification: Covalent addition of acetyl, methyl, or phosphate groups to histone tails affects transcription.
Acetylation: Decreases histone-DNA affinity, promoting transcription.
Methylation: Often represses transcription.
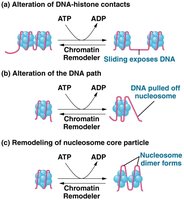
Chromatin Remodeling Complexes
SWI/SNF complex: Loosens histone-DNA interactions, making DNA accessible for transcription.
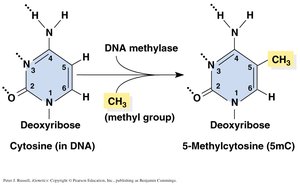
DNA Methylation and Genetic Imprinting
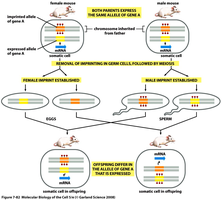
DNA methylation: Addition of methyl groups to cytosine, often silencing genes.
Genetic imprinting: Only one allele (maternal or paternal) is expressed, depending on parent of origin.
Transcriptional Regulation
Gene Structure and Regulatory Elements
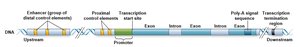
Eukaryotic genes contain coding regions, promoters, enhancers, silencers, and polyadenylation sites.
Promoters: On/off switch for transcription initiation.
Enhancers: Increase transcription in specific tissues or conditions.
Silencers: Repress transcription.
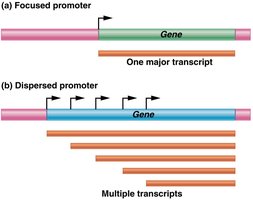
Promoter Diversity and Structure
Focused promoters: Initiate transcription at a single site.
Dispersed promoters: Initiate transcription at multiple sites.
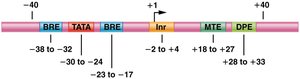
Core promoter elements: Include TATA box, initiator (Inr), BRE, DPE, and MTE.
Proximal-Promoter Elements
Located upstream of core promoter elements (e.g., CAAT and GC boxes).
Enhance basal transcription levels.
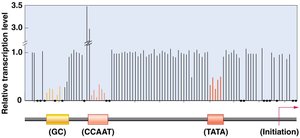
Enhancers and Silencers
Enhancers: Can be located far from the gene, increase transcription.
Silencers: Repress transcription initiation.
Transcription Factors
Transcription factors: Proteins that bind to cis-acting elements to regulate gene expression.
Activators: Increase transcription.
Repressors: Decrease transcription.
Functional domains: DNA-binding domain and trans-activating domain.
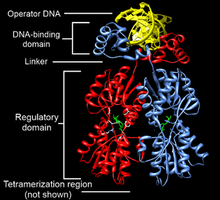
Common DNA Binding Motifs
Zinc finger: Coordinates zinc ions for DNA binding.
Helix-loop-helix: Facilitates dimerization and DNA binding.
Leucine zipper: Enables protein-protein interactions and DNA binding.
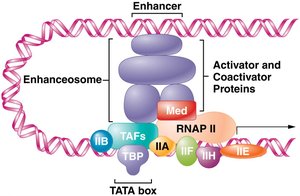
RNA Polymerase II Initiation Complex
General transcription factors: Required for transcription initiation.
Pre-initiation complex (PIC): Assembly of proteins provides a platform for RNA polymerase II.
Enhanceosome and Coactivators
Coactivators: Enable activators to contact promoter-bound factors.
Enhanceosome: Complex that interacts with transcription machinery to enhance transcription.
GAL Gene System in Yeast
Inducible Gene Expression
The GAL gene system in yeast is a model for inducible gene expression.
Structural genes: GAL1, GAL2, GAL7, GAL10 transport and metabolize galactose.
Regulatory genes: GAL3, GAL4, GAL80 regulate transcription.
Positive control: Activator protein must be present for transcription.
Post-Transcriptional Regulation
Alternative Splicing
Alternative splicing generates multiple mRNA variants from a single gene, increasing protein diversity.
Introns: Non-coding sequences removed during splicing.
Exons: Coding sequences joined to form mature mRNA.
Example: Calcitonin gene produces different peptides in thyroid and neurons.
Dscam Gene in Drosophila
The Dscam gene undergoes extensive alternative splicing, producing over 38,000 isoforms that guide neural connections.
Control of mRNA Stability
Steady-state level: Amount of mRNA available for translation.
Degradation pathways: Poly-A tail shortening, decapping, and endonuclease cleavage.
Translational and Posttranslational Regulation
Proteasome: Cylindrical structure that recycles amino acids.
Ubiquitin: Protein that tags other proteins for degradation.
p53 protein: Transcription factor regulated by ubiquitin-mediated degradation; increases in response to DNA damage.
Summary Table: Double-Strand Break Repair Pathways
Pathway | Mechanism | Accuracy | Cell Cycle Phase |
|---|---|---|---|
Homologous Recombination | Uses sister chromatid as template | High | Late S/G2 |
Nonhomologous End Joining | Direct ligation of ends | Low (mutagenic) | G1 |
Summary Table: Types of Transposable Elements
Type | Structure | Mechanism | Example |
|---|---|---|---|
IS Element | ITRs, transposase | Cut-and-paste | Bacteria |
Bacterial Transposon | ITRs, DRs, multiple genes | Cut-and-paste | Antibiotic resistance |
Retrotransposon | LTRs, integrase, RT | Copy-and-paste (RNA intermediate) | Copia in Drosophila |
Summary Table: Levels of Gene Regulation
Level | Mechanism | Example |
|---|---|---|
Chromatin Remodeling | Histone modification, nucleosome repositioning | SWI/SNF complex |
Transcriptional | Promoters, enhancers, silencers | GAL gene system |
Post-transcriptional | Alternative splicing, mRNA stability | Calcitonin gene |
Translational/Posttranslational | Proteasome, ubiquitin tagging | p53 regulation |
