 Back
BackGenetic Mechanisms: X-Chromosome Inactivation, Gene Interaction, and Mutation Effects
Study Guide - Smart Notes

X-Chromosome Inactivation and Dosage Compensation
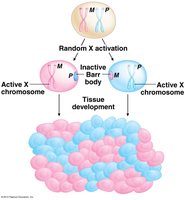
Random X-Chromosome Inactivation in Placental Mammals
In placental mammals, females possess two X chromosomes, while males have one X and one Y chromosome. To balance gene expression between sexes, one X chromosome in each female somatic cell is randomly inactivated early in development. This process, known as the Lyon hypothesis, results in the formation of a condensed structure called the Barr body, which is visible near the nuclear wall.
Dosage Compensation: Ensures equal expression of X-linked genes in males and females.
Permanence: Once inactivation occurs in a cell, it is maintained in all descendant cells.
Mosaicism: Female mammals are mosaics, with some cells expressing the maternal X and others the paternal X.
XIST: The X-inactivation-specific transcript (XIST) is a long non-coding RNA (~17,000 nucleotides) crucial for X inactivation.
Gene Interaction
Types of Gene Interactions
Gene interaction refers to the phenomenon where the expression of a trait is influenced by more than one gene or by the interaction of genes with environmental factors. Multiple alleles, incomplete dominance, and polygenic traits are common forms of gene interaction.
Multiple Alleles: More than two alleles may exist for a gene within a population.
Incomplete Dominance: Dominance of one allele over another may not be complete.
Polygenic Traits: Two or more genes may affect a single trait.
Gene-Environment Interaction: Expression of a trait may depend on both genetic and nongenetic factors.
The Molecular Basis of Dominance
Dominance and recessiveness are determined by the protein products of alleles. The phenotype is a result of the activities of these protein products.
Dominant Allele: Produces enough functional protein for normal phenotype.
Recessive Allele: Produces little or no functional protein.
Haplosufficiency: One copy of the dominant allele is sufficient for normal function.
Haploinsufficiency: One copy of the wild-type allele is not enough for normal function.
Functional Effects of Mutation
Wild-Type vs. Mutant Alleles
The wild-type allele produces a functional gene product, resulting in a normal phenotype. Mutant alleles can alter gene function, leading to various phenotypic consequences.

Loss-of-Function Mutations
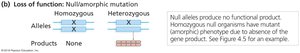
Loss-of-function mutations reduce or eliminate the activity of the gene product. These can be classified as null (amorphic) or leaky (hypomorphic) mutations.
Null Mutations: Produce no functional gene product; often lethal when homozygous.
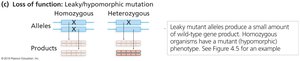
Leaky Mutations: Produce partial gene activity; severity depends on the level of activity.
Dominant Negative Mutations
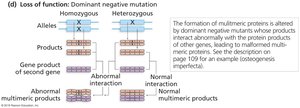
Dominant negative mutations occur in multimeric proteins, where the mutant polypeptide interferes with the function of the normal protein complex, resulting in a loss of function.
Multimeric Proteins: Composed of multiple polypeptides.
Spoiler Effect: Mutant polypeptide prevents normal interaction, causing abnormal protein function.
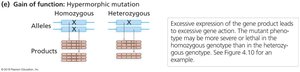
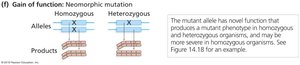
Gain-of-Function Mutations
Gain-of-function mutations increase gene activity or confer new functions. These are classified as hypermorphic or neomorphic mutations and are usually dominant.
Hypermorphic: Produces more gene activity than normal.
Neomorphic: Acquires novel gene activities not found in the wild type.
Incomplete Dominance
Definition and Example
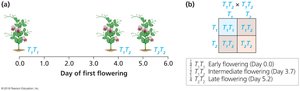
Incomplete dominance occurs when heterozygous individuals display intermediate phenotypes between either homozygous type. This is often observed in traits such as flowering time in pea plants.
Allele Designations: A1, A2 or B1, B2 are used instead of A, a or B, b.
Intermediate Phenotype: Heterozygote is more similar to one homozygous type than the other.
Summary Table: Functional Consequences of Mutation
Mutation Type | Gene Product | Phenotype |
|---|---|---|
Wild-Type | Normal | Wild-type |
Null (Amorphic) | None | Mutant (often lethal) |
Leaky (Hypomorphic) | Partial | Mutant (severity varies) |
Dominant Negative | Abnormal interaction | Mutant (spoiler effect) |
Hypermorphic | Excessive | Mutant (dominant) |
Neomorphic | Novel | Mutant (dominant) |
Key Equations and Concepts
Haplosufficiency and Haploinsufficiency
Haplosufficiency and haploinsufficiency describe whether a single copy of an allele is sufficient for normal function:
Haplosufficient: produces enough enzyme for wild-type phenotype.
Haploinsufficient: does not produce enough enzyme for wild-type phenotype.
Enzyme Activity Thresholds
Phenotype depends on enzyme activity:
Wild-type: threshold units
Mutant: threshold units
Example: If 40 units are required for wild-type phenotype:
: 100 units (wild-type)
: 50 units (wild-type)
: 0 units (mutant)
Additional info: These concepts are foundational for understanding genetic inheritance, gene expression, and mutation effects in eukaryotes.
