 Back
BackGenetics: Key Concepts in the Genetic Code, Transcription, Translation, and Gene Regulation
Study Guide - Smart Notes

Q1. The genetic code is _________, meaning that an amino acid may be encoded by more than one codon.
Background
Topic: Genetic Code Properties
This question tests your understanding of the redundancy in the genetic code and how multiple codons can specify the same amino acid.
Key Terms:
Genetic code: The set of rules by which information encoded in genetic material (DNA or RNA sequences) is translated into proteins by living cells.
Codon: A sequence of three nucleotides that together form a unit of genetic code in a DNA or RNA molecule.
Degeneracy: The property that allows more than one codon to specify the same amino acid.
Step-by-Step Guidance
Recall that there are 64 possible codons but only 20 amino acids.
Think about what it means if more than one codon can code for the same amino acid.
Identify the term that describes this redundancy in the genetic code.
Try solving on your own before revealing the answer!
Q2. The wobble hypothesis predicts that codons coding for the same amino acid _____.
Background
Topic: Wobble Hypothesis in Translation
This question examines your understanding of how tRNA anticodons can pair with more than one codon due to flexibility at the third codon position.
Key Terms:
Wobble hypothesis: The theory that the third base in a codon can form non-standard base pairs, allowing some tRNAs to recognize more than one codon.
Anticodon: A sequence of three bases in a tRNA molecule that pairs with a complementary codon in mRNA.
Step-by-Step Guidance
Recall that the first two bases of a codon are usually the most important for specifying an amino acid.
Consider what the wobble hypothesis says about the third position of the codon.
Think about how this flexibility affects the number of tRNAs needed and the degeneracy of the code.
Try solving on your own before revealing the answer!
Q3. The sigma subunit of bacterial RNA polymerase _____.
Background
Topic: Transcription Initiation in Prokaryotes
This question tests your knowledge of the role of the sigma subunit in bacterial transcription initiation.
Key Terms:
Sigma subunit: A protein component of bacterial RNA polymerase that recognizes and binds to promoter sequences, enabling the initiation of transcription.
Promoter: A DNA sequence that signals the start site for transcription.
Step-by-Step Guidance
Recall the structure of bacterial RNA polymerase and the function of its subunits.
Think about what must happen for transcription to begin in bacteria.
Identify which subunit is responsible for recognizing the promoter region.
Try solving on your own before revealing the answer!
Q4. What is the role of general transcription factors (GTFs)?
Background
Topic: Eukaryotic Transcription Initiation
This question focuses on the proteins required for the initiation of transcription by RNA polymerase II in eukaryotes.
Key Terms:
General transcription factors (GTFs): Proteins that assemble at the core promoter to help recruit RNA polymerase II and initiate transcription.
Core promoter: The DNA sequence where the transcription machinery assembles.
Step-by-Step Guidance
Recall the steps required for transcription initiation in eukaryotes.
Consider the role of GTFs in recruiting RNA polymerase II to the promoter.
Think about what would happen to transcription without GTFs.
Try solving on your own before revealing the answer!
Q5. In humans, the phenylalanine hydroxylase gene is 90,000 bases long, yet the mRNA is only 2,400 bases. What explains this difference?
Background
Topic: Gene Structure and RNA Processing
This question tests your understanding of the structure of eukaryotic genes and the processing of pre-mRNA to mature mRNA.
Key Terms:
Exon: A coding region of a gene that remains in the mature mRNA.
Intron: A non-coding region of a gene that is removed during RNA processing.
Splicing: The process of removing introns and joining exons in pre-mRNA.
Step-by-Step Guidance
Recall the structure of eukaryotic genes, including exons and introns.
Think about what happens to the primary transcript (pre-mRNA) after transcription.
Consider why the mature mRNA is much shorter than the gene itself.
Try solving on your own before revealing the answer!
Q6. Considering transcription initiation, the core promoter in eukaryotes is homologous to what in prokaryotes?
Background
Topic: Promoter Elements in Transcription
This question asks you to compare the DNA sequences that serve as the site of transcription initiation in eukaryotes and prokaryotes.
Key Terms:
Core promoter (eukaryotes): The region containing the TATA box where transcription factors and RNA polymerase assemble.
Pribnow box (prokaryotes): The -10 region in bacterial promoters, similar in function to the TATA box.
Step-by-Step Guidance
Recall the sequence elements that define the start of transcription in both eukaryotes and prokaryotes.
Identify the functional similarities between the TATA box and the Pribnow box.
Match the eukaryotic core promoter to its prokaryotic counterpart.
Try solving on your own before revealing the answer!
Q7. The poly(A) tail of mRNA _____.
Background
Topic: mRNA Processing in Eukaryotes
This question tests your understanding of the function and significance of the poly(A) tail in eukaryotic mRNA.
Key Terms:
Poly(A) tail: A stretch of adenine nucleotides added to the 3' end of most eukaryotic mRNAs after transcription.
mRNA stability: The poly(A) tail helps protect mRNA from degradation.
Step-by-Step Guidance
Recall the steps of mRNA processing in eukaryotes, including capping, splicing, and polyadenylation.
Think about the functions of the poly(A) tail in mRNA stability and translation.
Consider whether the poly(A) tail is involved in transcription termination, mRNA export, or other processes.
Try solving on your own before revealing the answer!
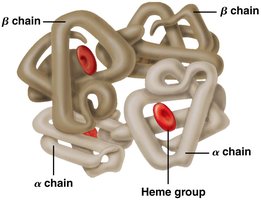
Q8. In this drawing of hemoglobin, what level(s) of structure are visible?
Background
Topic: Protein Structure
This question asks you to identify the levels of protein structure (primary, secondary, tertiary, quaternary) visible in a diagram of hemoglobin.
Key Terms:
Primary structure: The sequence of amino acids in a polypeptide chain.
Secondary structure: Local folding patterns such as alpha helices and beta sheets.
Tertiary structure: The overall 3D shape of a single polypeptide chain.
Quaternary structure: The arrangement of multiple polypeptide subunits in a protein complex.
Step-by-Step Guidance
Examine the diagram for evidence of alpha helices or beta sheets (secondary structure).
Look for the overall folding of each polypeptide chain (tertiary structure).
Identify if multiple polypeptide chains are interacting (quaternary structure).
Try solving on your own before revealing the answer!
Q9. Eukaryotic regulation of gene expression occurs at the level of _____.
Background
Topic: Regulation of Gene Expression
This question tests your understanding of the multiple levels at which gene expression can be regulated in eukaryotic cells.
Key Terms:
Transcriptional regulation: Control of gene expression at the level of mRNA synthesis.
Post-transcriptional regulation: Control at the level of mRNA processing, stability, and translation.
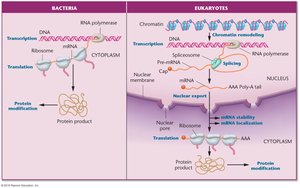
Step-by-Step Guidance
Recall the steps from DNA to protein in eukaryotic cells.
Think about where regulation can occur: transcription, RNA processing, mRNA stability, translation, and post-translational modification.
Identify which of these levels are used in eukaryotes.
Try solving on your own before revealing the answer!
Q10. What is the difference between the Initiator element (Inr) and the TATA box?
Background
Topic: Eukaryotic Promoter Elements
This question examines your understanding of the specific DNA sequences involved in the initiation of transcription in eukaryotes.
Key Terms:
TATA box: A DNA sequence found in the core promoter region of genes, important for the binding of transcription factors and RNA polymerase II.
Initiator element (Inr): A core promoter element where transcription actually begins, located downstream of the TATA box.
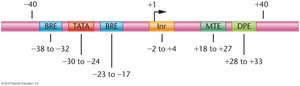
Step-by-Step Guidance
Recall the order and function of promoter elements in eukaryotic genes.
Identify where RNA polymerase binds and where transcription starts relative to these elements.
Distinguish between the binding site and the transcription start site.
Try solving on your own before revealing the answer!
