 Back
BackGenomic Technologies: DNA Sequencing, Genomics, and Multi-Omics Applications
Study Guide - Smart Notes

Recombinant DNA Technology and Genetically Modified Organisms (GMOs)
Introduction to Recombinant DNA and GMOs
Recombinant DNA technology enables the manipulation and combination of DNA from different sources to create genetically modified organisms (GMOs). This technology is foundational for modern genetics, biotechnology, and genomics research.
Genetically Modified Organisms (GMOs): Organisms whose genetic material has been altered using recombinant DNA methods to express desired traits.
Recombinant DNA: DNA molecules formed by laboratory methods of genetic recombination, such as molecular cloning, to bring together genetic material from multiple sources.
Applications: Agriculture (e.g., pest-resistant crops), medicine (e.g., insulin production), and research (e.g., gene function studies).

Polymerase Chain Reaction (PCR) and DNA Amplification
Principles and Applications of PCR
The Polymerase Chain Reaction (PCR) is a technique used to amplify specific DNA segments, making millions of copies from a small initial sample. PCR is essential for genetic analysis, cloning, and sequencing.
Key Steps: Denaturation, annealing, and extension using DNA polymerase.
Applications: Genetic testing, cloning, forensic analysis, and preparation for sequencing.
DNA Sequencing Technologies
Overview of DNA Sequencing
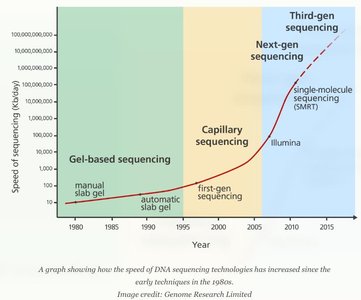
DNA sequencing determines the precise order of nucleotides in a DNA molecule. Sequencing technologies have evolved through three generations, each with distinct features and applications.
Goal: Identify the complete sequence of nucleotide bases (A, T, C, G) in a DNA sample.
Applications: Genome mapping, disease gene identification, evolutionary studies.
First Generation: Sanger Sequencing
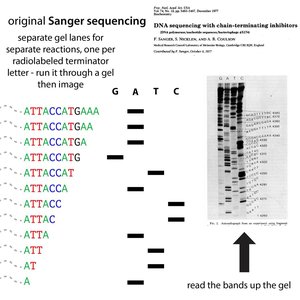
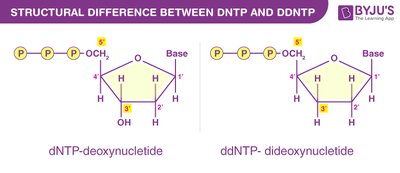
Sanger sequencing, also known as chain-termination sequencing, uses dideoxynucleotides (ddNTPs) to terminate DNA synthesis at specific bases, allowing the sequence to be determined by fragment size analysis.
Key Principle: Incorporation of ddNTPs halts DNA synthesis due to the absence of a 3'-OH group, preventing further elongation.
Detection: Fragments are separated by size using gel electrophoresis, and the sequence is read from the pattern of terminated fragments.
Limitations: Low throughput, suitable for sequencing single genes or small DNA fragments.
Second Generation: Next Generation Sequencing (NGS)
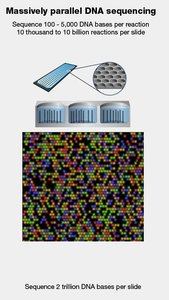
NGS technologies, such as Illumina sequencing, enable massively parallel sequencing of millions of short DNA fragments, dramatically increasing throughput and reducing cost per base.
Sequencing by Synthesis: DNA fragments are amplified and sequenced in parallel, with each nucleotide addition detected by fluorescence.
Applications: Whole genome sequencing, transcriptomics, epigenomics, and large-scale genetic studies.
Advantages: High throughput, automation, and cost-effectiveness for large-scale projects.


Third Generation: Single-Molecule Sequencing
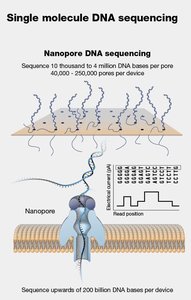
Third-generation sequencing technologies, such as PacBio and Oxford Nanopore, sequence single DNA molecules in real time, producing long reads that facilitate genome assembly and structural variant detection.
Key Features: Long read lengths (up to tens of kilobases), real-time sequencing, and the ability to detect epigenetic modifications.
Applications: De novo genome assembly, detection of structural variants, and analysis of complex genomic regions.
Limitations: Higher error rates and cost compared to NGS, but rapidly improving.

Summary Table: DNA Sequencing Technologies
Generation | Technology | Read Length | Throughput | Key Features |
|---|---|---|---|---|
First | Sanger Sequencing | 500–1,000 bp | Low | Accurate, single gene, slow |
Second | Illumina (NGS) | 50–500 bp | High | Massively parallel, short reads, cost-effective |
Third | PacBio, Nanopore | 10,000+ bp | Very High | Long reads, real-time, single molecule |
Genomic Libraries and Whole Genome Sequencing
Genomic DNA Libraries
Genomic libraries are collections of DNA fragments that represent the entire genome of an organism. These libraries were essential for early genome projects and are constructed using cloning vectors such as BACs and YACs.
Construction: Genomic DNA is fragmented, inserted into vectors, and cloned in host cells.
Applications: Genome mapping, gene discovery, and functional studies.
cDNA Libraries
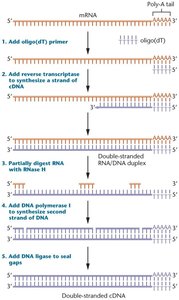
cDNA libraries are collections of complementary DNA (cDNA) synthesized from mRNA, representing only the expressed genes in a cell or tissue. They are used to study gene expression and identify genes involved in specific biological processes.
Construction: mRNA is reverse transcribed into cDNA, which is then cloned into vectors.
Applications: Expression analysis, identification of disease-related genes, and functional genomics.
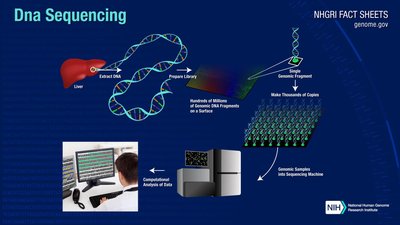
Whole Genome Sequencing (WGS)
WGS involves sequencing the entire genome of an organism, typically using NGS technologies. This approach provides comprehensive information about genetic variation, structure, and function.
Steps: DNA extraction, fragmentation, library preparation, sequencing, and bioinformatics analysis for assembly and annotation.
Applications: Human Genome Project, disease gene discovery, evolutionary studies.
Genome Assembly and Mapping
Genome Assembly
Genome assembly is the process of reconstructing the original genome sequence from short sequencing reads. This involves aligning overlapping reads to form contiguous sequences (contigs) and scaffolds.
Contig: A continuous sequence assembled from overlapping reads.
Scaffold: A series of contigs joined together using additional information, such as genetic or physical maps.
Challenges: Repetitive sequences, gaps, and sequencing errors.
Genetic vs. Physical Mapping
Genetic maps are based on recombination frequencies, while physical maps are based on actual DNA distances measured in base pairs.
Genetic Maps: Measure distances in centimorgans (cM), based on crossing over during meiosis.
Physical Maps: Measure distances in base pairs (bp), using sequencing, restriction mapping, or cytogenetics.
The Human Genome Project (HGP)
Overview and Achievements
The Human Genome Project was an international effort to sequence and map all human genes. It provided the first comprehensive reference genome and revolutionized genetics and biomedical research.
Timeline: Draft released in 2000, declared complete in 2003, gapless assembly in 2022.
Key Outcomes: Identification of ~20,000 protein-coding genes, discovery of genetic variation (SNPs, CNVs), and comparative genomics insights.
Reference Genome: Assembled from pooled DNA of multiple individuals, not a single person.
Multi-Omics Technologies
Genomics, Transcriptomics, Epigenomics, Proteomics, and Metabolomics
Multi-omics integrates various high-throughput technologies to analyze the genome, transcriptome, epigenome, proteome, and metabolome, providing a comprehensive view of biological systems.
Genomics: Study of the complete DNA sequence of an organism.
Transcriptomics: Analysis of all expressed RNA molecules (mRNA, ncRNA) in a cell or tissue.
Epigenomics: Study of DNA methylation, histone modifications, and chromatin structure affecting gene expression.
Proteomics: Analysis of all proteins, including their modifications and interactions.
Metabolomics: Study of small molecules and metabolites involved in cellular metabolism.
Applications of Multi-Omics
Genetic Testing: Identification of disease-associated variants.
Genome-Wide Association Studies (GWAS): Linking genetic variants to traits and diseases.
Synthetic Biology: Engineering organisms with novel functions.
Cancer Research: Multi-omics approaches for diagnosis, prognosis, and treatment strategies.
Comparative Genomics and Metagenomics
Comparative Genomics
Comparative genomics analyzes and compares genome sequences from different species to identify conserved and unique genetic elements, study evolutionary relationships, and understand gene function.
Applications: Disease gene discovery, evolutionary biology, and functional annotation.
Metagenomics
Metagenomics involves sequencing DNA from entire microbial communities in natural environments, bypassing the need for culturing individual species.
Applications: Discovery of novel microbes, understanding microbial diversity, and studying the human microbiome.
Functional Genomics
Defining Gene Function
Functional genomics uses genome-wide approaches to determine the roles of genes and regulatory elements, integrating data from transcriptomics, epigenomics, and proteomics.
Transcriptome: All RNA molecules transcribed from the genome.
Epigenome: All chemical modifications to DNA and histones.
Proteome: All proteins encoded by the genome.
Transcriptomics
Transcriptomics studies gene expression patterns in different tissues or conditions, using technologies like RNA sequencing (RNA-seq).
Bulk RNA-Seq: Measures average gene expression across many cells.
Single-Cell RNA-Seq: Resolves gene expression at the single-cell level, revealing cellular heterogeneity.
Epigenomics
Epigenomics analyzes genome-wide epigenetic modifications, such as DNA methylation and histone modifications, which regulate gene expression without altering the DNA sequence.
Methods: Whole genome bisulfite sequencing (WGBS), ATAC-seq, ChIP-seq.
Applications: Cancer research, developmental biology, environmental studies.
Proteomics
Proteomics investigates the structure, function, and interactions of proteins in a cell or tissue, often using mass spectrometry-based approaches.
Applications: Disease biomarker discovery, drug development, and understanding cellular processes.
Summary Table: Multi-Omics Technologies
Omics Field | Analyte | Key Methods | Applications |
|---|---|---|---|
Genomics | DNA | WGS, NGS | Genome mapping, variant discovery |
Transcriptomics | RNA | RNA-seq | Gene expression analysis |
Epigenomics | DNA/histones | WGBS, ChIP-seq, ATAC-seq | Epigenetic regulation |
Proteomics | Proteins | LC-MS/MS | Protein identification, function |
Metabolomics | Metabolites | MS, NMR | Metabolic profiling |
