 Back
BackGenomics, Bioinformatics, and Proteomics: An Overview
Study Guide - Smart Notes

Genomics, Bioinformatics, and Proteomics
Introduction to Genomics and Bioinformatics
Genomics is the comprehensive study of genomes, encompassing the structure, function, evolution, and mapping of genetic material in organisms. Bioinformatics applies mathematical and computational tools to organize, analyze, and interpret vast amounts of genetic data, including gene structure, sequence, expression, and protein function.
Genomics: Focuses on the sequencing, assembly, and functional analysis of genomes.
Bioinformatics: Involves the use of software and algorithms to manage and analyze genetic and protein data.
Applications: Includes gene identification, sequence comparison, evolutionary studies, and prediction of protein structure and function.
Whole-Genome Sequencing (WGS)
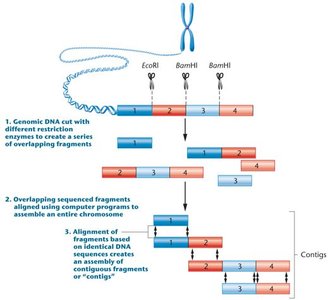
Whole-genome sequencing, also known as shotgun sequencing, is a primary strategy for determining the complete DNA sequence of an organism's genome. The process involves fragmenting DNA, sequencing the fragments, and assembling them into a continuous sequence using computational tools.
Steps in WGS:
Shearing or digestion of genomic DNA with restriction enzymes to create overlapping fragments.
Sequencing the fragments.
Alignment of sequenced fragments using computer programs.
Contig overlap: Identifying overlapping sequences to assemble the chromosome.
Gene identification by bioinformatics.
Contigs: Continuous DNA fragments formed by aligning overlapping sequences.
Bioinformatics Applications
Bioinformatics enables the alignment and comparison of DNA sequences, facilitating the reconstruction of chromosomes and identification of genes and regulatory elements.
DNA Sequence Alignment: Aligns similar sequences for comparison and assembly.
Gene Identification: Locates genes and regulatory regions such as promoters and enhancers.
Evolutionary Analysis: Deduces relationships between genes and species.
GenBank and BLAST
GenBank is the largest public database of DNA sequences, maintained by the National Center for Biotechnology Information (NCBI). BLAST (Basic Local Alignment Search Tool) is a software application used to compare new sequences to known sequences in databases, calculating identity values and E-values to assess similarity.
GenBank: Assigns accession numbers to sequences for retrieval and analysis.
BLAST: Identifies sequence similarity and potential gene function.

Gene Annotation and Hallmark Features
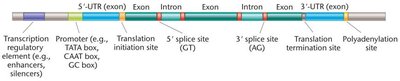
Gene annotation involves identifying the structural and functional elements of genes within a genome. Hallmark features include exons, introns, untranslated regions (UTRs), promoters, enhancers, silencers, and polyadenylation sites.
Exons and Introns: Coding and noncoding segments, respectively.
Regulatory Sequences: Promoters, enhancers, and silencers control gene expression.
UTRs: Untranslated regions at the 5' and 3' ends of mRNA.

Open Reading Frames (ORFs)
Open reading frames are sequences of DNA that can be translated into proteins. They typically begin with an initiation codon (ATG) and end with a stop codon (TAA, TAG, TGA).
Initiation Codon: ATG (codes for methionine).
Stop Codons: TAA, TAG, TGA (correspond to UAA, UAG, UGA in mRNA).
Exon-Intron Structure: Eukaryotic ORFs may contain both exons and introns.
Predicting Gene and Protein Functions
Functional genomics interprets DNA sequences to establish gene functions, often using similarity searches and experimental validation. BLAST searches can infer gene function based on sequence similarity to known genes.
Similarity Searches: Genes with similar sequences are likely to have similar functions.
Experimental Validation: Confirms computational predictions of gene function.
Homologous Genes and Orthologs
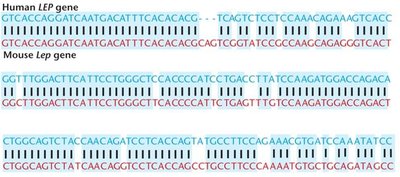
Homologous genes are evolutionarily related genes found in different species. Orthologs are homologous genes in different species that originated from a common ancestor.
Example: Comparison of the human LEP gene and mouse Lep gene shows high sequence similarity, indicating evolutionary conservation.
Protein Domains and Motifs
Protein domains and motifs are specific structural or functional regions within a protein. Analysis of gene sequences can predict the presence of domains (e.g., ion channels, membrane-spanning regions) and motifs (e.g., helix-turn-helix, leucine zipper, zinc-finger), which help infer protein function.
Major Features of the Human Genome
The human genome contains approximately 3.1 billion nucleotides, with protein-coding sequences comprising only about 2%. The genome is highly dynamic, with significant portions derived from transposable elements and repetitive DNA. Alternative splicing allows a relatively small number of genes (~20,000) to produce a much larger number of proteins.
Genome Diversity: Single-nucleotide polymorphisms (SNPs) and copy number variations (CNVs) contribute to genetic diversity.
Gene Distribution: Genes are unevenly distributed across chromosomes, with gene-rich and gene-poor regions.
Alternative Splicing: Over 50% of genes produce multiple proteins via alternative splicing.
Accessing and Utilizing the Human Genome Project (HGP)
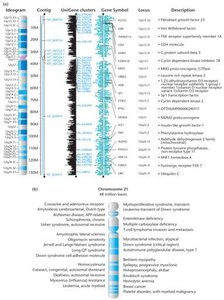
The HGP provides extensive maps and databases for all human chromosomes, facilitating the identification of disease genes and the development of new treatment strategies.
Gene Maps: Visual representations of gene locations and disease associations on chromosomes.

Omics Disciplines
Omics refers to various fields that analyze different biological molecules on a genome-wide scale, including proteomics, metabolomics, glycomics, toxicogenomics, metagenomics, pharmacogenomics, and transcriptomics.
Proteomics: Study of the complete set of proteins (proteome) encoded by the genome.
Metagenomics: Analysis of genetic material from environmental samples.
Whole-Exome Sequencing (WES)
Whole-exome sequencing focuses on sequencing only the exons (protein-coding regions) of the genome, which are more likely to contain disease-related mutations. However, WES does not capture regulatory regions that influence gene expression.
ENCODE Project
The Encyclopedia of DNA Elements (ENCODE) project aims to identify and analyze all functional elements in the human genome, including transcription start sites, promoters, and enhancers, using experimental and computational approaches.
Nutrigenomics
Nutrigenomics studies the interaction between nutrition and genes, providing personalized dietary recommendations based on genetic analysis to optimize health and prevent disease.
Stone-Age Genomics
Stone-age genomics involves sequencing ancient DNA from fossils and preserved tissues to study evolutionary relationships among extinct and extant species.
Comparative Genomics
Comparative genomics compares the genomes of different organisms to understand gene function, evolution, and the relationship between organisms and their environments. It is essential for gene discovery and the development of model organisms for human disease research.
Organism (Scientific Name) | Genome Size | Chromosome Number | Number of Genes | % Genes Shared with Humans |
|---|---|---|---|---|
Human (Homo sapiens) | 3.1 Gb | 46 | ~20,000 | 100 |
Mouse (Mus musculus) | ~2.5 Gb | 40 | ~30,000 | 80 |
Chimpanzee (Pan troglodytes) | 3 Gb | 48 | ~20,000–24,000 | 98 |
Yeast (Saccharomyces cerevisiae) | 12 Mb | 32 | ~5,700 | 30 |
Fruit fly (Drosophila melanogaster) | 165 Mb | 8 | ~13,600 | 50 |
Rice (Oryza sativa) | 389 Mb | 24 | ~41,000 | Not determined |
The Neanderthal Genome and Modern Humans
Sequencing of Neanderthal DNA has revealed that modern humans and Neanderthals share about 99% of their genome, with evidence of interbreeding. Non-African human genomes contain approximately 1–4% Neanderthal DNA.
Metagenomics and the Human Microbiome Project
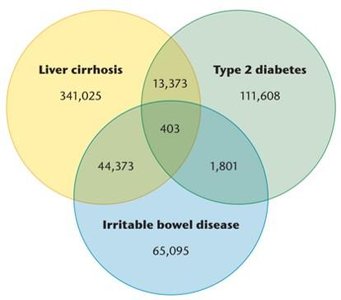
Metagenomics uses WGS to analyze genetic material from environmental samples, revealing the diversity of microbial communities. The Human Microbiome Project (HMP) aims to sequence the genomes of microorganisms living in and on humans, providing insights into health and disease.
Venn Diagram Analysis: Shows overlap and uniqueness of microbial genes associated with different diseases.
Proteomics
Proteomics is the large-scale study of proteins, including their identification, characterization, and quantification. It provides insights into protein structure, function, interactions, and modifications, and is crucial for understanding cellular processes and disease mechanisms.
Proteome: The complete set of proteins expressed by a genome.
Applications: Identification of disease biomarkers, study of protein-protein interactions, and analysis of post-translational modifications.
Proteomics Technologies
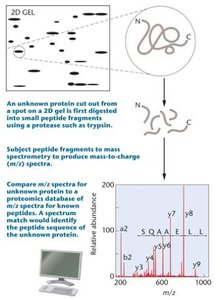
Two-dimensional gel electrophoresis (2DGE) and mass spectrometry (MS) are key technologies in proteomics.
2D Gel Electrophoresis: Separates proteins based on isoelectric point and molecular weight.
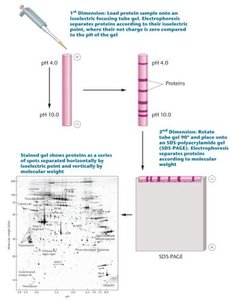
Mass Spectrometry: Identifies proteins by measuring the mass-to-charge ratio of ionized peptides, often following 2DGE.

Additional info: This guide summarizes key concepts from Chapter 18 of "Essentials of Genetics," focusing on genomics, bioinformatics, and proteomics, and their applications in modern genetics research.
