 Back
BackIntroduction to Chromosomes, Genome Organization, and Repetitive DNA
Study Guide - Smart Notes

Unit 1: Introduction to Chromosomes & Nuclear Processes
Overview of Chromosomes and Genome Organization
Chromosomes are the fundamental units of genetic material, composed of DNA and associated proteins. Their structure, number, and organization vary widely among organisms, influencing genome size, gene density, and complexity. Understanding chromosomes is essential for studying inheritance, gene expression, and genome evolution.
Chromosome Number: The number of chromosomes varies by species. For example, humans have 46 chromosomes, while koalas have 16.
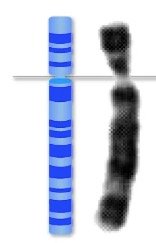
Chromosome Structure: Chromosomes can be linear (eukaryotes) or circular (prokaryotes and organelles like mitochondria and chloroplasts).
Key Features: Each chromosome contains a centromere, telomeres, and numerous genes.
The Cell Cycle and Chromosome Behavior
The cell cycle describes the sequence of events in a cell's life, including growth, DNA replication, and division. Chromosomes undergo specific changes during each phase, which are critical for accurate genetic transmission.
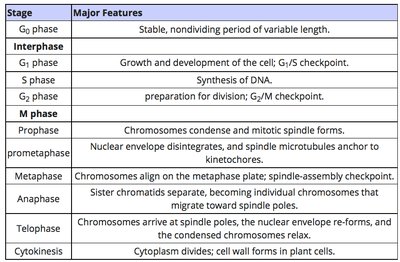
Interphase: Includes G1 (growth), S (DNA synthesis), and G2 (preparation for division).
M Phase: Mitosis and cytokinesis, where chromosomes condense, align, and segregate into daughter cells.
Stage | Major Features |
|---|---|
G0 phase | Stable, nondividing period of variable length. |
G1 phase | Growth and development of the cell; G1/S checkpoint. |
S phase | Synthesis of DNA. |
G2 phase | Preparation for division; G2/M checkpoint. |
M phase | Chromosomes condense and mitotic spindle forms. |
Prophase | Chromosomes condense and mitotic spindle forms. |
Prometaphase | Nuclear envelope disintegrates, spindle microtubules anchor to kinetochores. |
Metaphase | Chromosomes align on the metaphase plate; spindle-assembly checkpoint. |
Anaphase | Sister chromatids separate, becoming individual chromosomes that migrate toward spindle poles. |
Telophase | Chromosomes arrive at spindle poles, nuclear envelope re-forms, chromosomes relax. |
Cytokinesis | Cytoplasm divides; cell wall forms in plant cells. |
Meiosis: Reductional Division and Genetic Diversity
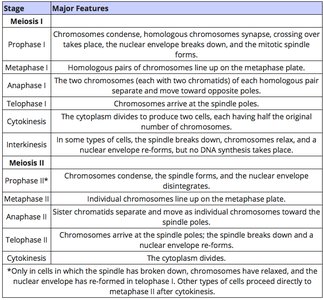
Meiosis is a specialized cell division that reduces chromosome number by half, producing gametes or spores. It consists of two sequential divisions (meiosis I and II) and introduces genetic variation through recombination and independent assortment.
Stage | Major Features |
|---|---|
Prophase I | Homologous chromosomes synapse, crossing over occurs, nuclear envelope breaks down, spindle forms. |
Metaphase I | Homologous pairs align on the metaphase plate. |
Anaphase I | Homologous chromosomes separate to opposite poles. |
Telophase I | Chromosomes arrive at spindle poles. |
Cytokinesis | Cytoplasm divides, producing two cells with half the original chromosome number. |
Meiosis II | Similar to mitosis; separates sister chromatids. |
Ploidy and Chromosome Number
Ploidy refers to the number of sets of chromosomes in a cell. Organisms can be haploid (1n), diploid (2n), triploid (3n), tetraploid (4n), or even higher. Chromosome number and ploidy vary widely among species and can influence genetic diversity and adaptation.
Ploidy: Number of complete chromosome sets (e.g., 2n for humans).
Chromosome Number (n): The actual number of chromosomes in one set (e.g., n=23 for humans).
C-value: The mass (in picograms) or number of base pairs of DNA in a haploid nucleus.



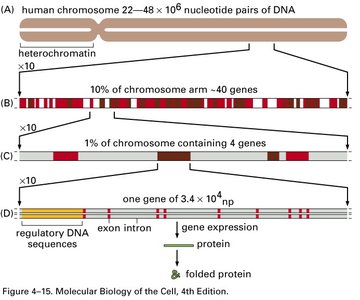
Human Genome Size and Gene Density
The human genome contains approximately 3.2 billion base pairs and about 20,000 protein-coding genes. Gene density (number of genes per megabase) varies among organisms and is not directly proportional to genome size or organismal complexity.
Cell | Chromosomes Description | Type | Ploidy | Base Pairs (bp) | GC Content (%) | Density (Mbp/pg) | Mass (pg) | C-Value |
|---|---|---|---|---|---|---|---|---|
Sperm or egg | 23 heterologous chromosomes | X Gamete | Haploid | 3,031,042,417 | 40.574607 | 977.9571 | 3.09831 | 3.09831 |
Sperm only | 23 heterologous chromosomes | Y Gamete | Haploid | 2,932,228,857 | 41.077476 | 977.9564 | 2.99833 | 2.99833 |
Zygote | 46 chromosomes (XX Female) | Diploid | 6,062,084,834 | 40.826541 | 977.9567 | 6.19664 | 3.09832 | |
Zygote | 46 chromosomes (XY Male) | Mostly diploid | 5,963,271,554 | 40.955875 | 977.9567 | 6.09784 | 3.15787 |
The C-Value Paradox
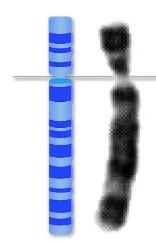
The C-value paradox refers to the observation that genome size (C-value) does not correlate with organismal complexity. For example, some plants and amphibians have much larger genomes than humans, despite being less complex in terms of cell types and functions.
Genome size can be measured in base pairs (bp) or DNA mass (picograms).
Organisms with similar complexity can have vastly different genome sizes.





Gene Density and Noncoding DNA
Gene density is the number of genes per megabase of DNA. Organisms with similar genome sizes can have different gene densities due to varying amounts of noncoding DNA, such as introns, regulatory sequences, and repetitive elements.
Example: Yeast has a much higher gene density than humans, meaning more genes per megabase of DNA.
Less complex organisms often have less intergenic (noncoding) DNA.
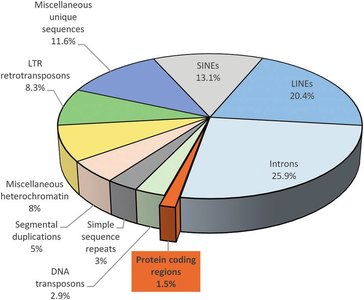
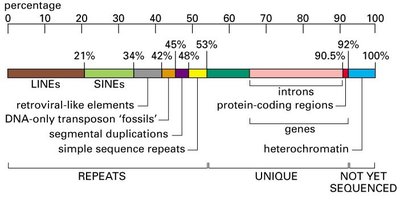
Repetitive DNA and Genome Composition
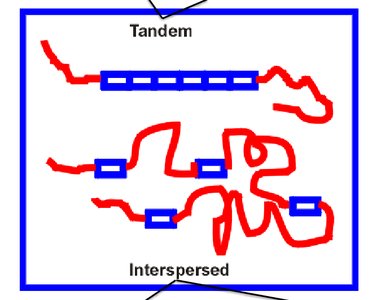
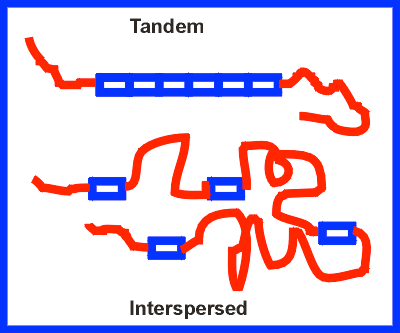
More than half of the human genome consists of repetitive DNA, which can be classified as tandem repeats or interspersed repeats. These sequences play roles in genome structure, evolution, and sometimes gene regulation.
Tandem Repeats: Short sequences repeated in series (e.g., microsatellites, minisatellites).
Interspersed Repeats: Scattered throughout the genome (e.g., LINEs, SINEs).
Transposable Elements (TEs): Mobile DNA sequences that can move within the genome, including LINEs and SINEs.
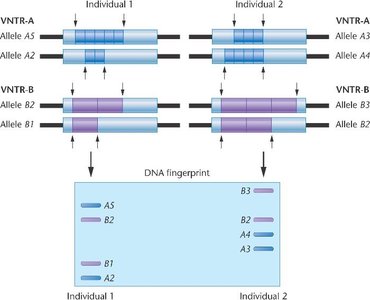
DNA Fingerprinting and Variable Number Tandem Repeats (VNTRs)
DNA fingerprinting exploits the variability in the number of tandem repeats (VNTRs) among individuals. This technique is widely used in forensics and genetic identification.
VNTRs: Regions where a short DNA sequence is repeated a variable number of times.
Individuals differ in the number of repeats, producing unique DNA profiles.
Transposable Elements: Types and Effects
Transposable elements (TEs) are DNA sequences that can move within the genome. They are classified into two main types:
DNA Transposons: Move by a cut-and-paste mechanism.
Retrotransposons: Move by a copy-and-paste mechanism via an RNA intermediate, increasing their copy number.
TEs can disrupt gene function, alter gene expression, and cause chromosomal rearrangements. They are a major source of genetic variation and evolution.
Historical Perspective: Discovery of Transposable Elements
The discovery of transposable elements revolutionized our understanding of the genome. Barbara McClintock first described these 'jumping genes' in maize, demonstrating that the genome is dynamic and capable of rearrangement.
Her work was initially dismissed but later recognized with a Nobel Prize in Physiology or Medicine in 1983.
TEs are now known to play roles in gene regulation and genome evolution.


Summary and Key Takeaways
Chromosome number, structure, and ploidy vary widely among organisms and are not always correlated with complexity.
The human genome is mostly noncoding and contains a large proportion of repetitive DNA, including transposable elements.
Transposable elements are a major source of genetic variation and can impact gene function and genome structure.
DNA fingerprinting utilizes variable tandem repeats for individual identification.
