 Back
BackLesson 5: Telomerase, DNA Replication, and DNA Damage
Study Guide - Smart Notes

DNA Replication: Mechanisms and Challenges
Replication Bubble and Bidirectional Replication
DNA replication in eukaryotes initiates at multiple origins of replication, forming structures known as replication bubbles. Replication proceeds bidirectionally from each origin, creating leading and lagging strands at each fork.
Replication Bubble: A region where the double helix is unwound and replication occurs in both directions.
Leading Strand: Synthesized continuously in the 5' to 3' direction toward the replication fork.
Lagging Strand: Synthesized discontinuously in short segments called Okazaki fragments, also in the 5' to 3' direction but away from the fork.
Key Proteins: DNA polymerase, primase, helicase, ligase, and single-stranded binding proteins are essential for replication.
Okazaki Fragments: Short DNA segments synthesized on the lagging strand, later joined by DNA ligase.
Example: In a replication bubble, the leading strand is synthesized continuously, while the lagging strand is synthesized in fragments that are later joined together.
Directionality and Enzyme Function
DNA polymerase can only add nucleotides to the 3' end of a growing DNA strand, resulting in the need for primers and discontinuous synthesis on the lagging strand.
Polarity: DNA strands are antiparallel, with one running 5' to 3' and the other 3' to 5'.
Primase: Synthesizes short RNA primers to provide a 3' OH group for DNA polymerase.
Helicase: Unwinds the DNA double helix ahead of the replication fork.
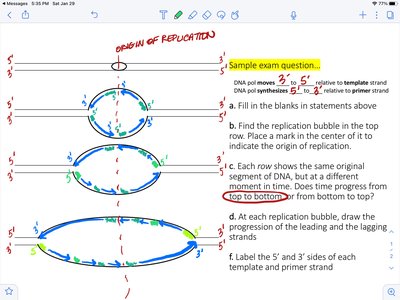
Additional info: The first dNMP incorporated into the daughter strand is adjacent to the primer, and the first phosphodiester bond forms between the primer and the first dNMP.
End-Replication Problem and Telomerase
Chromosome Shortening in Eukaryotes
During replication, the lagging strand cannot be fully replicated at the chromosome ends, leading to progressive shortening with each cell division. This is known as the end-replication problem.
Leading Strand: Can be replicated completely to the end.
Lagging Strand: After removal of the final RNA primer, there is no upstream 3' OH for DNA polymerase to extend, resulting in a shortened daughter strand.
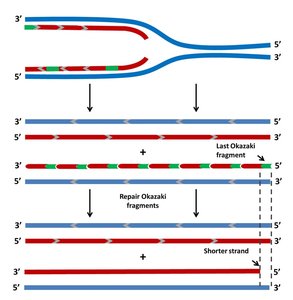
Example: In human somatic cells, telomeres shorten with each division, eventually leading to cell senescence.
Telomerase: Structure and Function
Telomerase is a ribonucleoprotein enzyme that extends the 3' ends of linear chromosomes, preventing loss of genetic information.
Components: Telomerase consists of a protein component (TERT, a reverse transcriptase) and an RNA template (TERC).
Mechanism: Telomerase uses its RNA template to add telomeric repeats to the 3' overhang, allowing primase and DNA polymerase to fill in the lagging strand.
Reverse Transcription: Telomerase synthesizes DNA from an RNA template, a process called reverse transcription.
Species Specificity: The sequence of the telomerase RNA template is species-specific, matching the telomere repeat sequence.
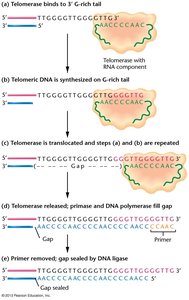
Example: In Tetrahymena, the telomerase RNA template is 9 ribonucleotides long and matches the telomere repeat sequence.
Regulation and Cancer Connection
Telomerase activity is tightly regulated in normal cells but is often reactivated in cancer cells, contributing to their immortality.
Normal Regulation: Telomerase is active in embryonic, stem, and germline cells but inactive in most somatic cells.
Cancer: Over 90% of human tumors reactivate telomerase (hTERT), enabling unlimited cell division.
Therapeutic Target: Telomerase is a promising target for cancer immunotherapy due to its restricted expression in normal tissues.
Additional info: Telomere shortening acts as a tumor suppressor mechanism by limiting cell proliferation.
DNA Damage and Repair Mechanisms
Types of DNA Damage
DNA is constantly exposed to damaging agents, both endogenous and exogenous, which can lead to mutations if unrepaired.
Induced Damage: Caused by external factors such as UV radiation, chemicals, and ionizing radiation (e.g., thymidine dimers from UV light).
Spontaneous Damage: Results from intrinsic instability, including base tautomers, depurination, and deamination.
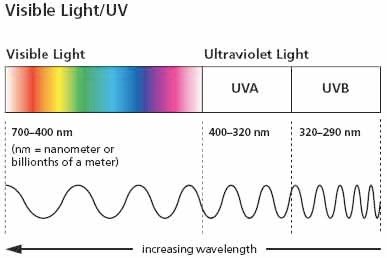
Example: UV light induces cyclobutane pyrimidine dimers, distorting the DNA helix and blocking replication/transcription.
DNA Proofreading and Repair Pathways
Cells have evolved multiple mechanisms to maintain genome integrity by correcting DNA lesions before they become permanent mutations.
Proofreading: DNA polymerase has 3' to 5' exonuclease activity, removing misincorporated nucleotides and reducing error rates.
Mismatch Repair (MMR): Detects and repairs mismatches missed by proofreading. In bacteria, strand discrimination is based on methylation status.
Base Excision Repair (BER): Removes and replaces damaged bases.
Nucleotide Excision Repair (NER): Removes bulky lesions, such as thymidine dimers.
Double-Strand Break Repair: Includes homologous recombination (HR) and non-homologous end joining (NHEJ).
Example: MMR proteins (MutS, MutL, MutH in bacteria) recognize mismatches, excise the incorrect strand, and fill in the correct sequence.
Consequences of Unrepaired DNA Damage
If DNA lesions are not repaired before the next round of replication, they become permanent mutations, which can contribute to disease, including cancer.
Lesion: A temporary chemical change in DNA that can be repaired.
Mutation: A permanent change in the DNA sequence.
Additional info: The high fidelity of DNA replication and repair is essential for maintaining genetic stability across generations.
