 Back
BackMolecular Cloning, Transgenic Technology, and Gene Regulation: Study Notes for Genetics
Study Guide - Smart Notes

Regulation of Transcription and Chromatin Structure
Regulatory Transcription Factors (RTFs)
Regulatory transcription factors (RTFs) are proteins that modulate the rate of transcription of specific target genes, distinguishing them from basal transcription factors, which are required for the basic level of transcription. RTFs bind to specific DNA sequences known as regulatory elements and can act as activators or repressors.
Activators: Increase transcription by facilitating the binding of RNA polymerase to the promoter.
Repressors: Decrease transcription by hindering the assembly or activity of the transcriptional machinery.
Regulatory Elements: DNA sequences (such as enhancers or silencers) recognized by RTFs.
Chromatin Status and Transcription
The structure of chromatin influences the accessibility of DNA to RTFs and RNA polymerase, thereby affecting transcriptional activity.
Open Conformation: Chromatin is less condensed, allowing transcription factors and polymerase to access DNA, promoting transcription.
Closed Conformation: Chromatin is tightly packed, restricting access and repressing transcription.
Additional Factors: Nucleosome positioning, histone variants, histone modifications, and DNA methylation all contribute to chromatin status and gene regulation.
Histone Variants and Gene Expression
Histone variants are alternative protein forms that can replace standard histones in nucleosomes, influencing chromatin structure and gene expression.
Variants are often incorporated after DNA replication.
Functions include roles in centromere function, telomere maintenance, DNA repair, and X-chromosome inactivation.
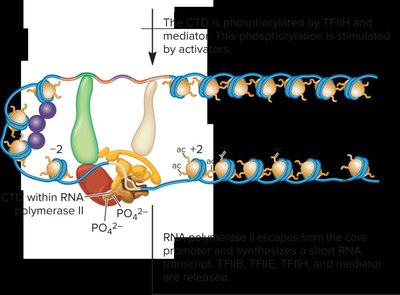
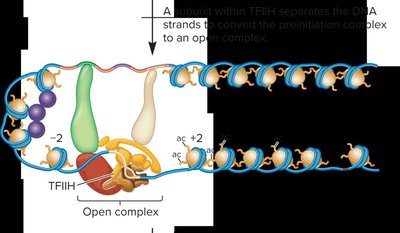
Formation of the Open Complex in Transcription Initiation
During transcription initiation, the DNA strands separate to form an open complex, allowing RNA polymerase II to begin RNA synthesis. Promoter escape occurs when RNA polymerase II is released from the core promoter and starts elongation.
Epigenetics and Noncoding RNAs
Epigenetics
Epigenetics is the study of heritable and reversible changes in gene expression that do not involve alterations to the DNA sequence itself.
Key Mechanisms: DNA methylation, chromatin remodeling, histone modification and variants.
Noncoding RNAs (ncRNAs)
Noncoding RNAs are RNA molecules that do not encode proteins but can bind to DNA, RNA, proteins, or small molecules to regulate gene expression.
Gene Silencing by Interfering RNAs
Small interfering RNAs (siRNAs) and microRNAs (miRNAs) are processed from double-stranded or stem-loop RNA precursors by the enzyme Dicer, generating ~25 nucleotide fragments.
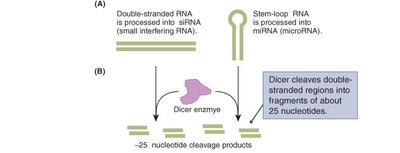
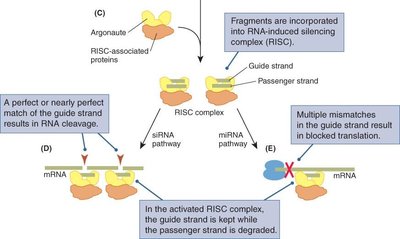
These fragments are loaded onto the RNA-induced silencing complex (RISC), which uses the guide strand to target complementary mRNA for degradation or translational repression.
Molecular Cloning Techniques
Overview of Genetic Engineering
Genetic engineering involves manipulating DNA to create new genotypes and phenotypes, often using recombinant DNA technology or gene cloning. Organisms with introduced recombinant DNA are called transgenic organisms or genetically modified organisms (GMOs).
Applications: Studying gene function, producing proteins, creating GMOs, and gene therapy.
Making Recombinant DNA
Recombinant DNA is created by combining DNA fragments of interest with vector DNA, which can be propagated in host cells such as bacteria.
Vector: A DNA molecule (e.g., plasmid or viral vector) used to carry foreign DNA into a host cell.
Selectable Marker: A gene (e.g., antibiotic resistance) that allows for identification of cells containing the vector.
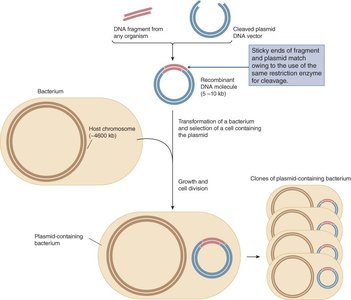
Restriction Enzymes and DNA Ligation
Restriction enzymes (endonucleases) recognize specific DNA sequences and cleave DNA, generating fragments with sticky or blunt ends. DNA ligase is used to join these fragments, forming recombinant DNA molecules.
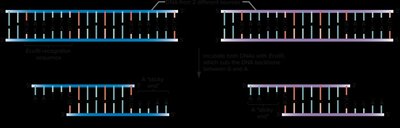
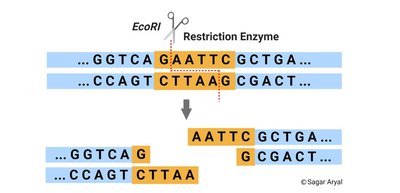
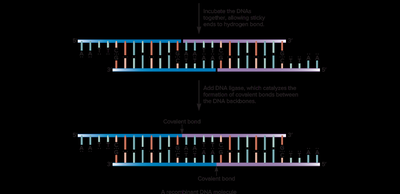
Cloning Workflow
Restriction enzymes and ligase are used to link together DNA from different sources, which is then transformed into a bacterial host for propagation.
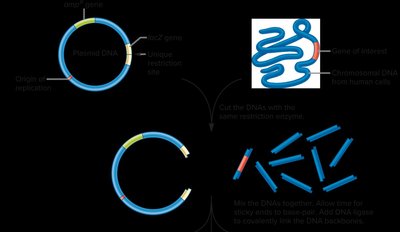
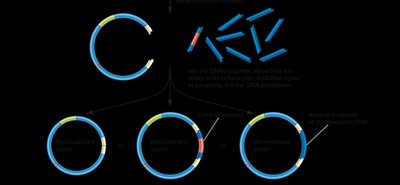
Selection of Recombinant Clones
Vectors often contain antibiotic resistance genes for selection. Only bacteria that have taken up the plasmid can grow on antibiotic-containing media.
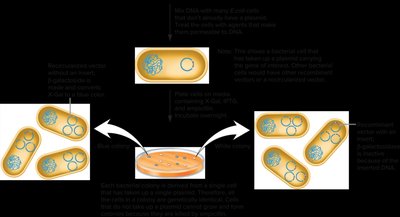
Blue-White Screening for Recombinant DNA
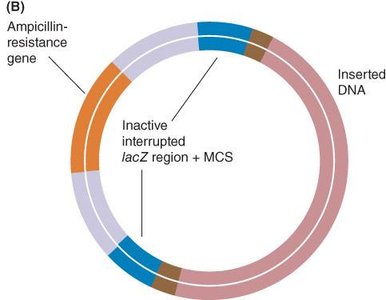
Blue-white selection uses the lacZ gene and X-gal substrate to distinguish between recombinant and non-recombinant clones. Insertion of foreign DNA disrupts lacZ, resulting in white colonies (recombinant), while blue colonies indicate functional lacZ (non-recombinant).
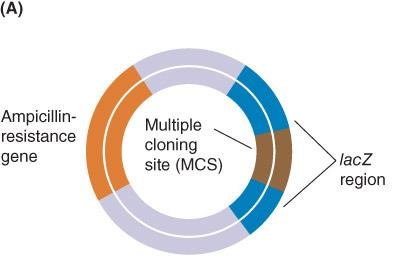
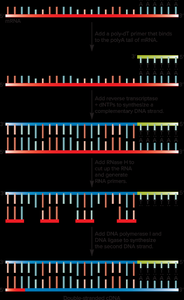
cDNA Cloning
Complementary DNA (cDNA) is synthesized from mRNA using reverse transcriptase, allowing for cloning of eukaryotic genes without introns.
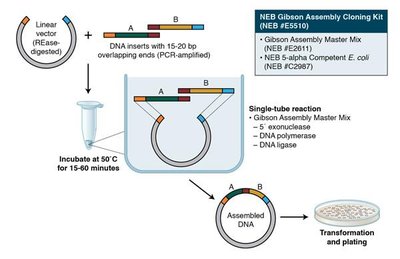
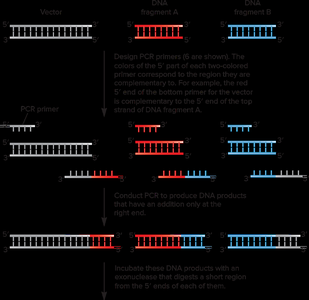
Gibson Assembly
Gibson Assembly is an exonuclease-based method for joining multiple DNA fragments in a single reaction, independent of restriction enzymes. It uses overlapping sequences at fragment ends and three enzymes: a 5' exonuclease, a polymerase, and a ligase.
Transgenic Technology and Reproductive Cloning
Reproductive Cloning
Reproductive cloning refers to the creation of an entire organism genetically identical to the donor. The classic example is Dolly the sheep, produced by transferring a somatic cell nucleus into an enucleated egg.
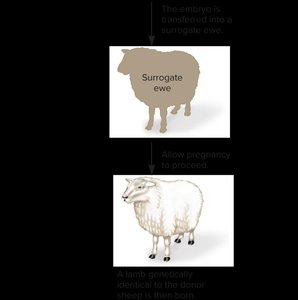
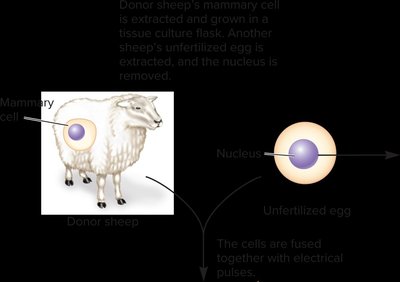
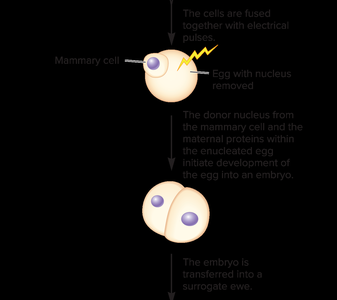

Issues: Cloned animals may have shortened telomeres and abnormal gene expression.
Making Transgenic Animals
Transgenic technology introduces foreign genes into animals for research or biotechnology. Methods vary by organism:
C. elegans: Extrachromosomal arrays and MosSCI-based single-copy insertion.
Drosophila: P element-mediated gene insertion using transposable elements.
Mice: Germline transformation using embryonic stem (ES) cells.
Extrachromosomal Arrays in C. elegans
Injected DNA forms extrachromosomal arrays, which can be inherited but may be unstable or silenced in the germline.
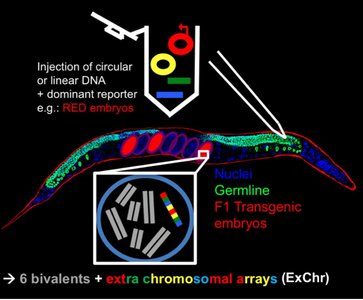
MosSCI-Based Insertion
MosSCI allows for single-copy insertion of transgenes into the genome, supporting stable germline expression.
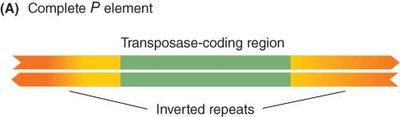
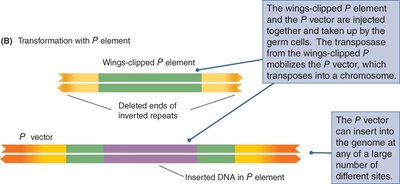
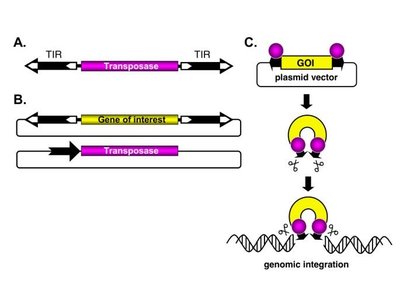
P Element-Mediated Gene Insertion in Drosophila
P elements are transposable elements with inverted repeats and a transposase-coding region. A two-component system is used: one plasmid carries the transposase, and another carries the gene of interest. The transposase mediates insertion of the gene into the genome.
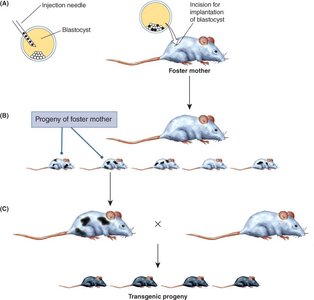
Germline Transformation of Mice
DNA is introduced into mouse ES cells by electroporation. Transformed cells are injected into a blastocyst, which is implanted into a foster mother. Mosaic offspring can transmit the transgene to progeny.
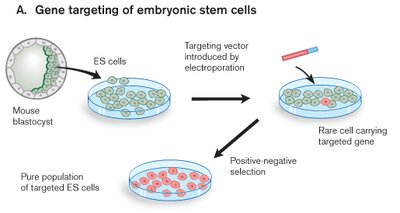
Gene Targeting in Embryonic Stem Cells
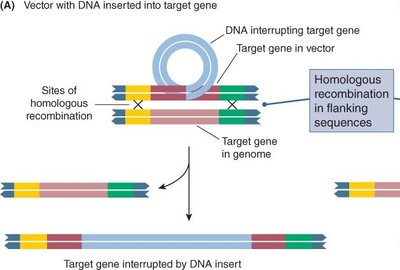
Gene targeting introduces specific changes into a gene using homologous recombination. Vectors with sequences homologous to the target gene facilitate precise insertion or replacement.
Method | Organism | Key Feature |
|---|---|---|
Extrachromosomal Array | C. elegans | Multiple copies, unstable, often silenced in germline |
MosSCI | C. elegans | Single-copy, stable, germline expression |
P Element | Drosophila | Transposase-mediated insertion |
ES Cell Transformation | Mouse | Germline transmission, gene targeting possible |
Additional info: These methods are foundational for creating genetically modified organisms for research, agriculture, and medicine.
