 Back
BackMolecular Structure and Organization of Chromosomes
Study Guide - Smart Notes

Chapter 11: Molecular Structure of Chromosomes
Introduction to Chromosome Structure
The genome encompasses all the genetic material of an organism. In bacteria, this is typically a single circular chromosome, while in eukaryotes, it refers to a complete set of nuclear chromosomes. Eukaryotes also possess mitochondrial genomes, and plants have an additional chloroplast genome.
Genome: The total genetic content in a cell or organism.
Mitochondrial genome: Circular DNA found in mitochondria, inherited maternally in most species.
Chloroplast genome: Circular DNA found in plant chloroplasts, essential for photosynthesis.
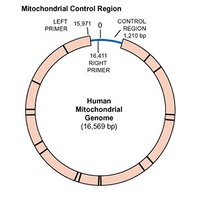

Structure of Bacterial Chromosomes
Bacterial chromosomes are typically circular and located in the nucleoid region, which is not membrane-bound. The DNA is in direct contact with the cytoplasm, and most of the DNA encodes proteins, with intergenic regions present.
Nucleoid: Region in bacteria where the chromosome is located.
Supercoiling: Additional twisting of the DNA helix to compact the chromosome further.
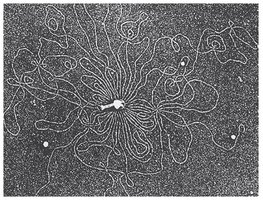
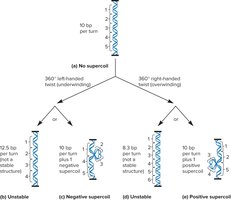
Structure of Eukaryotic Chromosomes
Eukaryotic DNA is highly organized and packaged to prevent tangling and to allow regulated access for replication, repair, and gene expression. DNA is packaged into multiple chromosomes, each consisting of a long DNA molecule wrapped around proteins.
Chromatin: The complex of DNA and proteins (mainly histones) that forms chromosomes.
Origins of replication: Sites where DNA replication begins; eukaryotic chromosomes have many origins.
Centromeres: Regions important for chromosome segregation during cell division.
Telomeres: Specialized ends of chromosomes, important for stability and replication.



Chromosome Number and Genome Size
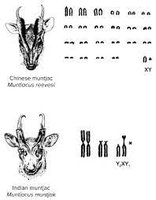
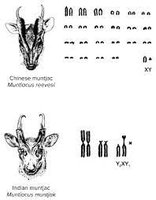
Each species has a characteristic number of chromosomes, but there is no direct correlation between chromosome number, genome size, and organismal complexity. For example, some ferns have over 1,000 chromosomes, while humans have 46.
Homologous chromosomes: Chromosome pairs with the same genes but possibly different alleles.
Sex chromosomes: Chromosomes that determine sex (XX in females, XY in males in humans).





Genes and Intergenic DNA
Chromosomes are long chains of nucleotides, with specific sequences (genes) encoding proteins. Not all DNA codes for proteins; much of it is intergenic, sometimes called "junk DNA," though some is conserved and may have regulatory or structural roles.
Gene: Functional unit of heredity; any DNA sequence transcribed into RNA.
Intergenic DNA: Non-coding regions between genes.
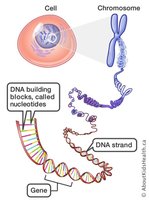
DNA Compaction in Chromosomes
Human chromosomes are highly compacted to fit within the nucleus. For example, chromosome 22 is compacted 10,000-fold during mitosis. Compaction must balance DNA accessibility for gene expression and replication.
Nucleosomes: The Basic Units of Eukaryotic Chromosomes
The nucleosome is the fundamental repeating unit of chromatin, consisting of DNA wrapped around a histone octamer. This structure is essential for the first level of DNA compaction.
Histone octamer: Composed of two copies each of H2A, H2B, H3, and H4.
Linker DNA: DNA between nucleosomes, not bound by histones.
Histone H1: Linker histone that helps compact nucleosomes into higher-order structures.

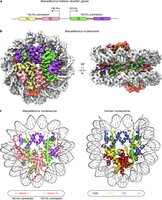
Experimental Evidence for Nucleosome Structure
The nucleosome model was proposed by Roger Kornberg and tested by Markus Noll using DNase I digestion and gel electrophoresis. The results supported the "beads-on-a-string" model, where nucleosomes are separated by linker DNA.



Higher-Order Chromatin Structure
Nucleosomes further associate to form a 30 nm fiber, a more compact structure. Histone H1 is important for this compaction. At low salt concentrations, H1 remains bound, and nucleosomes are more compact.
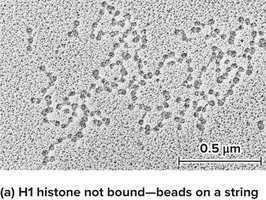
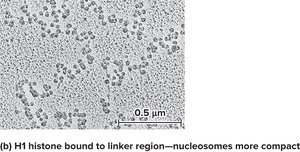


Chromatin Remodeling and Chemical Modifications
Chromatin structure is dynamic and can be modified by chemical changes to histone tails, affecting gene expression and DNA accessibility.
Acetylation: Addition of acetyl groups by histone acetyltransferase (HAT), neutralizing positive charges and increasing gene activity.
Methylation: Addition of methyl groups by methyltransferase, which can increase or decrease transcription.
Phosphorylation: Addition of phosphate groups by kinases, important in cell cycle and DNA replication.
CpG islands: Regions where cytosine is methylated, often associated with gene silencing.
Further Compaction: Loop Domains and SMC Proteins
The 30 nm fiber folds into loop domains, further compacting the chromosome. CTCF and SMC proteins (Structural Maintenance of Chromosomes) are key in forming and stabilizing these loops.
Heterochromatin vs. Euchromatin
Chromatin exists in two main forms during interphase:
Heterochromatin: Highly compacted, transcriptionally inactive, often contains repetitive sequences.
Euchromatin: Less condensed, transcriptionally active, contains most genes.
Heterochromatin can be constitutive (always inactive) or facultative (can switch between active and inactive states).
Chromosome Structure During Cell Division
During mitosis, chromosomes become highly condensed, forming metaphase chromosomes. Condensin and cohesin, both containing SMC proteins, are essential for chromosome condensation and sister chromatid cohesion, respectively.
Transposable Elements (TEs)
Transposable elements are DNA sequences that can move within the genome, discovered by Barbara McClintock. They contribute to genetic diversity and genome evolution.
Autonomous elements: Contain all necessary sequences for movement, including transposase.
Non-autonomous elements: Lack some sequences and rely on autonomous elements for movement.
Transposase: Enzyme that catalyzes the movement of TEs.
Terminal inverted repeats (TIRs): Short, repeated sequences at the ends of TEs recognized by transposase.
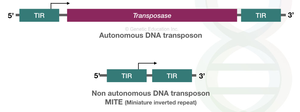
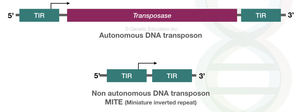
Types and Mechanisms of Transposition
Simple transposition: TE moves directly to a new site.
Retrotransposition: TE moves via an RNA intermediate, requiring reverse transcriptase and integrase.
SINEs and LINEs: Short and long interspersed nuclear elements, common retrotransposons in eukaryotes.
Abundance and Significance of TEs
TEs are major components of eukaryotic genomes, making up a significant portion of DNA in many species. Their biological significance is debated, but they are thought to persist as "selfish DNA," proliferating as long as they do not severely harm the host.
Organism | Percentage of Genome as TEs |
|---|---|
Human, chimp, mouse, ape | ~50% |
Maize | ~75% |
Barley | ~85% |
Iris | ~98% |
Example: In maize, TEs cause the spotted kernel phenotype by inserting into and excising from pigment genes, leading to mosaic expression.
Additional info: The discovery of TEs revolutionized our understanding of genome plasticity and regulation, and TEs are now recognized as important drivers of evolution and genetic diversity.
