 Back
BackCh 9: Molecular Structure of DNA and RNA: Foundations of Genetic Material
Study Guide - Smart Notes

Molecular Structure of DNA and RNA
Introduction to Molecular Genetics
Molecular genetics is the study of DNA and RNA structure and function at the molecular level. Recent advances in techniques have greatly expanded our understanding of how genetic information is stored, transmitted, and expressed. The molecular structure of DNA and RNA forms the foundation for all genetic processes, including inheritance, replication, and variation.
Discovery of Nucleic Acids
The discovery of nucleic acids began in 1869 when Friedrich Miescher identified a substance he called "nuclein" in cell nuclei, which was later recognized as DNA. This discovery was unique among cellular components and played a pivotal role in understanding fertilization and heredity.

Chromosomes and the Human Genome
Each human cell contains 23 sets of chromosomes, with each chromosome consisting of a single DNA molecule. DNA molecules can range from 500,000 to 2.5 million nucleotide pairs and, when uncoiled, can be several centimeters long. The total DNA in all human cells can stretch millions of miles, illustrating the vast amount of genetic information contained within the genome.
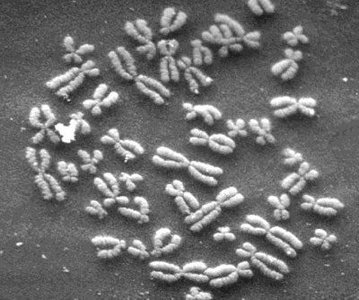
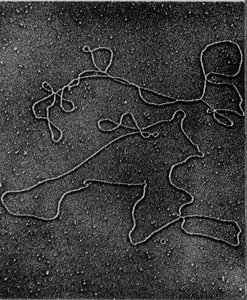
Components of the Human Genome
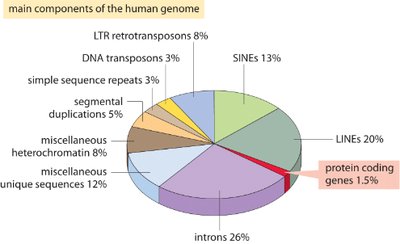
Only about 1.5% of the human genome codes for proteins, while the remaining 98.5% consists of non-coding regions, regulatory elements, and sequences with unknown functions. These components have accumulated and evolved over time, contributing to genetic diversity and complexity.
Protein-coding genes: 1.5%
Introns: 26%
Transposable elements: LINES (20%), SINES (13%), LTR retrotransposons (8%), DNA transposons (3%)
Miscellaneous heterochromatin: 8%
Miscellaneous unique sequences: 12%
Segmental duplications: 5%
Simple sequence repeats: 3%
Identification of DNA as the Genetic Material
Criteria for Genetic Material
To serve as genetic material, a molecule must:
Information: Contain instructions to build an organism
Transmission: Be passed from parent to offspring
Replication: Be copied accurately
Variation: Allow for changes to account for phenotypic diversity
Griffith's Transformation Experiment (1928)
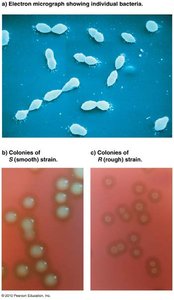
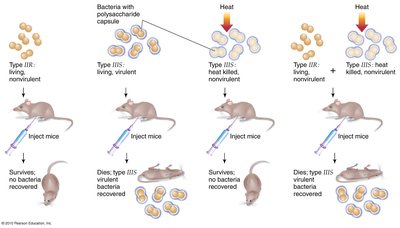
Griffith used two strains of Streptococcus pneumoniae (S and R) to demonstrate that a "transforming principle" could convert non-virulent bacteria into virulent forms. This experiment laid the groundwork for identifying DNA as the genetic material.
S strain: Virulent, has polysaccharide coat
R strain: Non-virulent, lacks coat
Transformation occurred when heat-killed S strain was mixed with live R strain, resulting in virulent bacteria
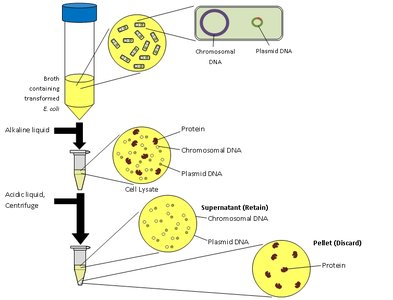
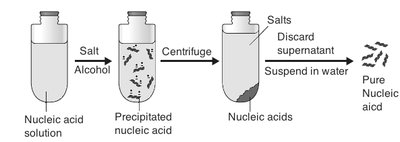
Avery, MacLeod, and McCarty (1944)
These researchers isolated DNA and demonstrated that it was the transforming principle responsible for genetic change, not protein or RNA. Their experiments used chemical methods to purify DNA and showed that only DNA could transform bacteria.
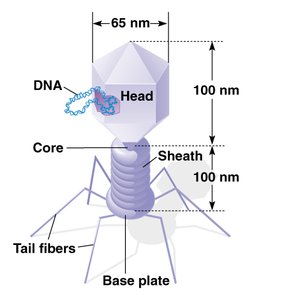
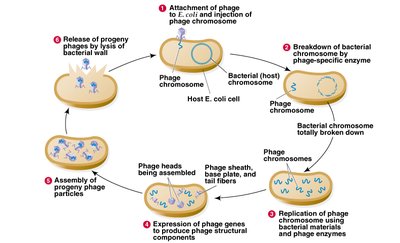
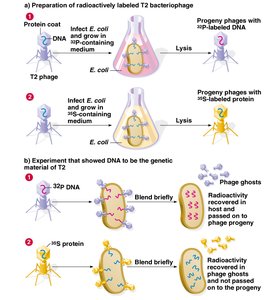
Hershey-Chase Experiment (1952)
Using bacteriophage T2, Hershey and Chase conclusively showed that DNA, not protein, is the genetic material. Radioactive labeling of DNA and protein allowed them to track which component entered bacterial cells and directed viral replication.
DNA labeled with 32P entered cells and directed viral replication
Protein labeled with 35S did not enter cells
Structure of DNA and RNA
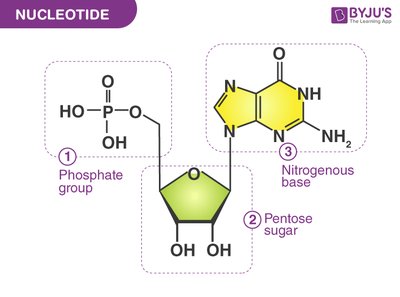
Nucleotide Structure
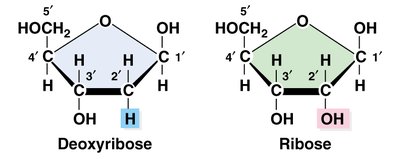
Nucleotides are the repeating units of DNA and RNA, each consisting of a phosphate group, a pentose sugar (ribose in RNA, deoxyribose in DNA), and a nitrogenous base. Nucleotides are linked by phosphodiester bonds to form strands.
Phosphate group
Pentose sugar: Ribose (RNA), Deoxyribose (DNA)
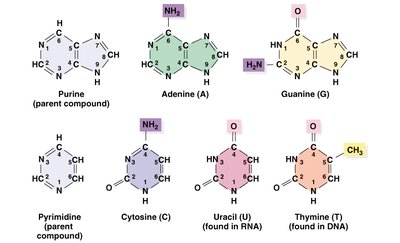
Nitrogenous base: Purines (A, G), Pyrimidines (C, T, U)
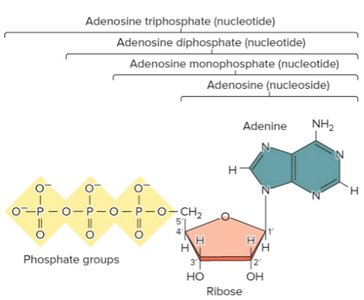
Nucleosides and Nucleotides
A nucleoside consists of a base and a sugar, while a nucleotide includes one or more phosphate groups. Examples include adenosine (nucleoside), adenosine monophosphate (AMP), adenosine diphosphate (ADP), and adenosine triphosphate (ATP).
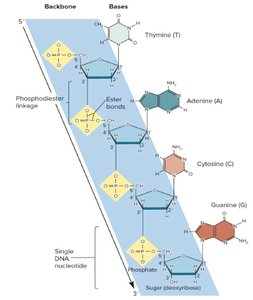
DNA Strand Structure
Nucleotides are covalently linked by phosphodiester bonds, connecting the 5' carbon of one nucleotide to the 3' carbon of another. This gives DNA strands directionality (5' to 3'). The sugar-phosphate backbone forms the structural framework, with bases projecting outward.
Discovery of the Double Helix
James Watson and Francis Crick elucidated the double helical structure of DNA in 1953, building on data from Linus Pauling, Rosalind Franklin, Maurice Wilkins, and Erwin Chargaff. Franklin's X-ray diffraction studies revealed DNA's helical nature and provided key measurements.
Chargaff's Rules
Erwin Chargaff found that DNA from any organism has a 1:1 ratio of purines to pyrimidines, specifically that the amount of adenine equals thymine and the amount of guanine equals cytosine. This base pairing is fundamental to the double helix structure.
A = T
G = C
Physical Structure of DNA: The Double Helix
The double helix consists of two antiparallel strands twisted around a common axis. There are 10 base pairs and 3.4 nm per complete turn. The helix is right-handed, and the strands are stabilized by hydrogen bonding and base stacking.
Hydrogen bonds: A-T (2 bonds), C-G (3 bonds)
Base stacking: Flat regions of bases face each other
Grooves in the DNA Helix
DNA has two asymmetrical grooves: the major groove and the minor groove. Certain proteins bind within these grooves to interact with specific DNA sequences, influencing gene regulation and expression.
Alternative DNA Structures
DNA can adopt alternative secondary structures, such as B DNA (the predominant form) and Z DNA (left-handed helix, 12 bp per turn). Z DNA formation is favored by alternating purine/pyrimidine sequences, cytosine methylation, and negative supercoiling. Z DNA may play roles in transcription and chromosome structure.
Triplex DNA
Triplex DNA can form when a synthetic DNA strand binds to the major groove of double-stranded DNA. This structure is sequence-specific and has been implicated in recombination, gene inactivation, and potential therapeutic applications.
RNA Structure
Primary and Secondary Structure of RNA
RNA strands are similar to DNA but use uracil instead of thymine and ribose instead of deoxyribose. RNA is usually single-stranded but can form short double-stranded regions through complementary base pairing (A-U, C-G). RNA double helices are right-handed and typically have the A form with 11-12 base pairs per turn.
Bulge loop
Internal loop
Multibranched loop
Stem loop (hairpin loop)
Tertiary Structure of RNA
The tertiary structure of RNA is determined by base pairing, base stacking, and interactions with ions, small molecules, and proteins. Transfer RNA (tRNA) is a classic example of a complex RNA tertiary structure.
Long Noncoding RNA (lncRNA): Xist
Xist RNA is a large transcript expressed on the inactive X chromosome in humans. It remains in the nucleus and coats the inactive X, playing a crucial role in X chromosome inactivation.
Summary Table: DNA vs. RNA
Feature | DNA | RNA |
|---|---|---|
Sugar | Deoxyribose | Ribose |
Bases | A, T, G, C | A, U, G, C |
Strandedness | Double-stranded (usually) | Single-stranded (usually) |
Helix Form | B DNA, Z DNA, Triplex DNA | A form (double helix in short regions) |
Additional info: Some content was expanded for clarity and completeness, including definitions, examples, and summary tables.
