 Back
BackMutations, DNA Repair, and Their Impact on Gene Expression
Study Guide - Smart Notes

Mutations and DNA Repair
Introduction to Mutations
Mutations are heritable changes in the DNA sequence or chromosome structure that generate new alleles, drive genetic diversity, and provide the raw material for evolution. They can occur spontaneously or be induced by external factors, and their effects range from silent to deleterious or even lethal.
Somatic mutations: Occur in non-germ cells; not heritable.
Germ-line mutations: Occur in gametes; heritable and contribute to evolution and inherited diseases.
Mutations can affect coding regions (exons) or noncoding regions (introns, promoters, enhancers).
Classification of Gene Mutations
Gene mutations are classified by the type of nucleotide change and their effect on protein function.
Point mutations (base substitutions): Change of one base pair to another.
Silent (synonymous) mutation: Alters a codon but not the encoded amino acid.
Missense mutation: Changes a codon to encode a different amino acid.
Nonsense mutation: Converts a codon to a stop codon, terminating translation prematurely.
Frameshift mutation: Insertion or deletion of nucleotides shifts the reading frame, altering downstream amino acids.
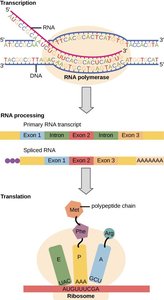
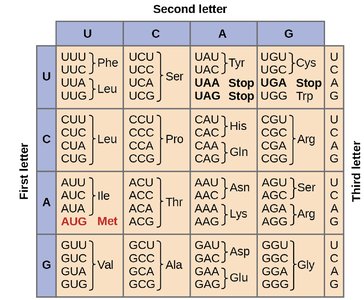
Wobble Hypothesis and Synonymous Mutations
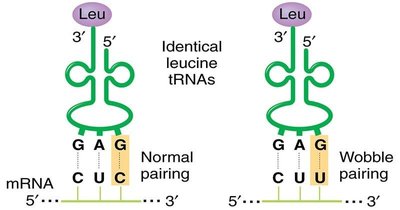
The genetic code is degenerate, meaning multiple codons can encode the same amino acid. The third base of the codon (wobble position) often tolerates changes without altering the amino acid, leading to silent mutations.
Wobble pairing: Allows one tRNA to recognize multiple codons.
Reduces the impact of mutations at the third codon position.

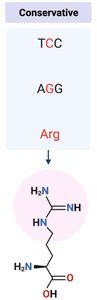
Types of Missense Mutations
Missense mutations can be further classified based on the properties of the substituted amino acid:
Conservative mutation: New amino acid has similar properties; protein function often preserved.
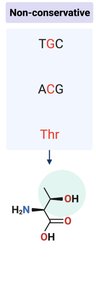
Non-conservative mutation: New amino acid has different properties; may disrupt protein structure or function.
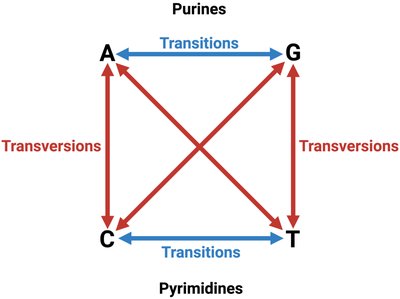
Base Substitutions: Transitions and Transversions
Base substitutions are categorized as transitions or transversions:
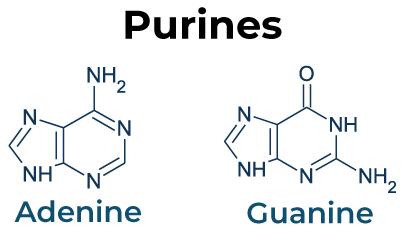
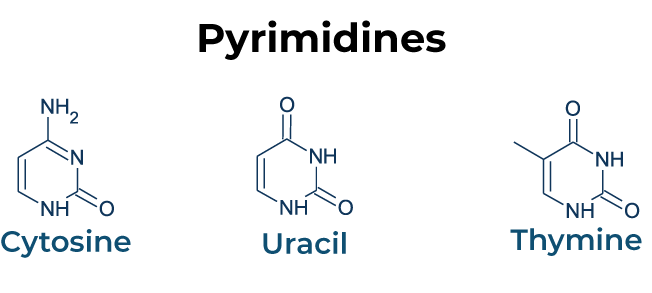
Transition: Purine replaces purine (A ↔ G) or pyrimidine replaces pyrimidine (C ↔ T).
Transversion: Purine replaces pyrimidine or vice versa (A/G ↔ C/T).
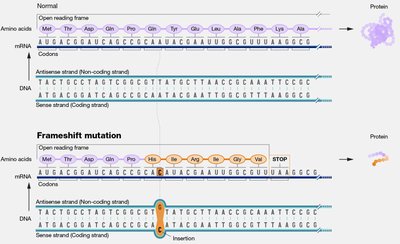
Frameshift Mutations
Frameshift mutations result from insertions or deletions that are not multiples of three nucleotides, altering the reading frame and usually resulting in nonfunctional proteins.
Mutation Classification by Location and Phenotype
Autosomal mutations: Affect genes on autosomes (chromosomes 1–22).
X- and Y-linked mutations: Affect genes on sex chromosomes.
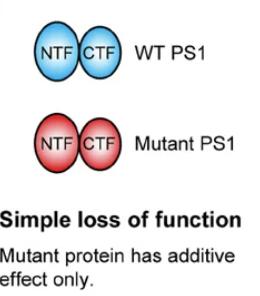
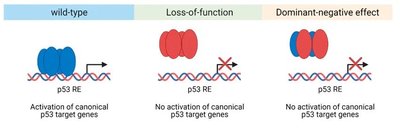
Loss-of-function mutation: Decreased or abolished gene function.
Null mutation: Complete loss of gene function.
Dominant mutation: Phenotype appears in heterozygotes.
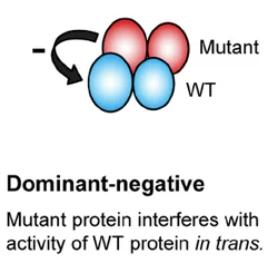
Dominant negative mutation: Mutant protein interferes with normal protein function.
Haploinsufficiency: One functional allele is insufficient for normal function (e.g., Marfan syndrome).
Molecular Causes of Mutations
DNA replication errors: Mismatches during replication can lead to mutations if not corrected.
Spontaneous chemical changes: Depurination (loss of purine base) and deamination (removal of amino group) can alter base pairing.
Mutagens: Physical (UV, X-rays), chemical (base analogs, intercalating agents), or biological agents that increase mutation rates.
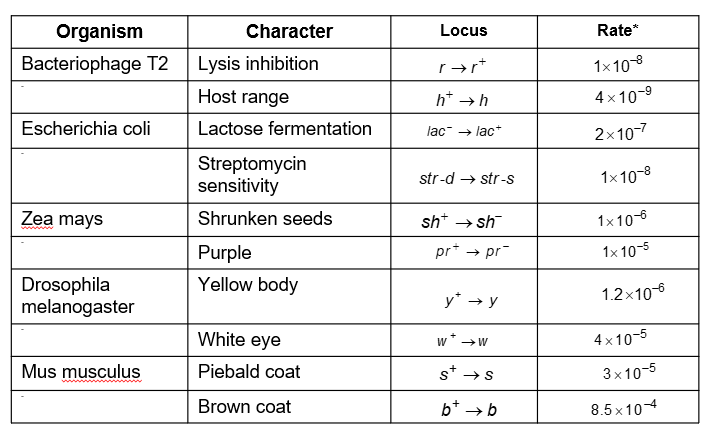
Mutation Rates and Genome Size
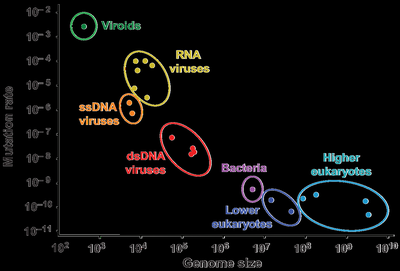
Mutation rates vary among organisms and are influenced by genome size and the fidelity of DNA replication and repair mechanisms.
Viruses (especially RNA viruses) have high mutation rates due to error-prone replication.
Bacteria and eukaryotes have lower mutation rates due to proofreading and repair systems.
Per base mutation rate decreases with genome size, but total mutations per genome increase in larger genomes.
DNA Repair Mechanisms
Cells possess multiple mechanisms to correct DNA errors and maintain genomic integrity:
Proofreading by DNA polymerase: 3′ → 5′ exonuclease activity removes mismatched bases during replication.
Mismatch repair: Detects and repairs mismatches missed by proofreading.
Direct repair: Reverses specific base modifications.
Excision repair: Removes damaged bases or nucleotides and fills the gap with new DNA.
Induced Mutations and Experimental Mutagenesis
Mutations can be experimentally induced using chemicals, radiation, or gene-editing techniques:
Base analogs: Incorporated into DNA, causing mispairing.
Deaminating agents: Change base identity (e.g., cytosine to uracil).
Alkylating agents: Add alkyl groups to bases, altering pairing.
Intercalating agents: Insert between bases, causing frameshifts.
Site-directed mutagenesis: Introduces specific mutations at chosen sites for research or biotechnology.
Distribution of Fitness Effects (DFE) of Mutations
The DFE describes how new mutations affect an organism's fitness, influencing evolution, disease, and population viability. Most mutations are neutral or deleterious; only a small fraction are beneficial.
Regulation of Gene Expression and Post-Transcriptional Control
Post-Translational Regulation
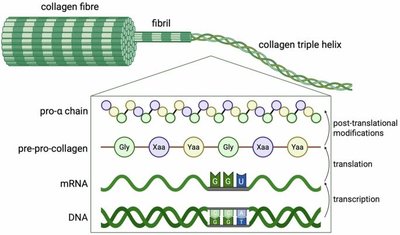
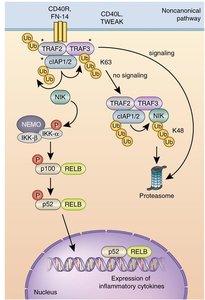
Gene expression can be regulated at multiple levels, including post-translational modifications such as ubiquitination, phosphorylation, cleavage, and translocation. These modifications control protein stability, activity, and localization.
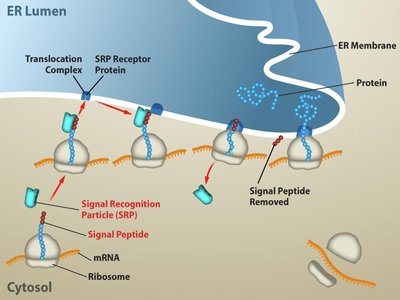
Protein Targeting and Secretion
Proteins destined for secretion contain an N-terminal signal sequence that directs them to the endoplasmic reticulum (ER) during translation. The signal recognition particle (SRP) guides the ribosome to the ER membrane, where the protein is translocated into the ER lumen for further processing and secretion.
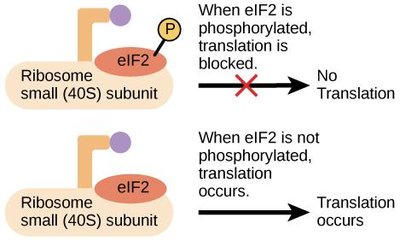
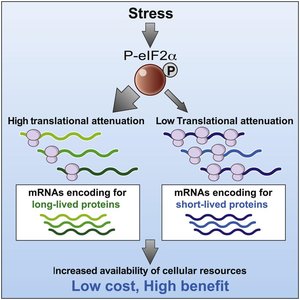
Translational Control: eIF2 Phosphorylation
Translation initiation can be regulated by phosphorylation of the eukaryotic initiation factor 2 (eIF2). Phosphorylated eIF2 inhibits translation, a mechanism activated during cellular stress and observed in neurodegenerative diseases.
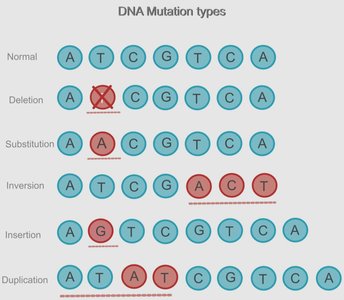
Summary Table: Types of DNA Mutations
Type | Description |
|---|---|
Normal | Reference DNA sequence; no mutation |
Deletion | Removal of one or more nucleotides; may cause frameshift |
Substitution | Replacement of one nucleotide with another; may alter protein function |
Inversion | Segment of DNA is reversed within the sequence |
Insertion | Addition of one or more nucleotides; may cause frameshift |
Duplication | Segment of DNA is copied and repeated |
Key Equations
Mutation rate per base per generation:
Additional info: This guide integrates foundational concepts from "Concepts of Genetics" (Ch. 15) and related biology texts, providing a comprehensive overview of mutation types, mechanisms, and their biological significance.
