 Back
BackNon-Mendelian Inheritance, Complementation, and Organelle Genetics
Study Guide - Smart Notes

Complementation and Gene Interaction
Complementation Analysis
Complementation analysis is a classical genetic approach used to determine whether mutations that produce similar phenotypes are in the same gene or in different genes. This method is essential for understanding genetic heterogeneity and the functional relationships between genes.
Genetic Heterogeneity: Occurs when mutations in different genes result in the same or similar mutant phenotypes.
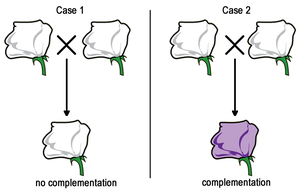
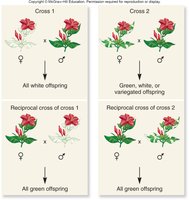
Complementation: When two organisms with similar mutant phenotypes are crossed, and their offspring display the wild-type phenotype, the mutations are said to complement each other. This indicates that the mutations are in different genes.
No Complementation: If the offspring display the mutant phenotype, the mutations are likely in the same gene.
Complementation Group: A set of mutations that fail to complement each other, defining a single gene.
Example: In a classic experiment, crossing two white-flowered plants can yield either white or purple offspring, depending on whether the mutations are in the same or different genes.
Non-Mendelian Inheritance
Introduction to Non-Mendelian Patterns
While Mendelian inheritance describes genes that directly influence traits and obey Mendel's laws, many genes in eukaryotes exhibit non-Mendelian inheritance patterns. These include maternal effect, dosage compensation, genomic imprinting, and extranuclear inheritance.
Maternal Effect: The genotype of the mother determines the phenotype of the offspring, regardless of the offspring's own genotype.
Dosage Compensation: Mechanisms that balance the expression of sex chromosome genes between males and females.
Genomic Imprinting: Only one parental allele is expressed, depending on epigenetic marks.
Extranuclear Inheritance: Inheritance of genes located outside the nucleus, typically in mitochondria or chloroplasts.
Maternal Effect
Definition and Mechanism
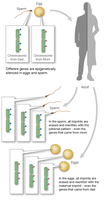
Maternal effect refers to a phenomenon where the phenotype of the offspring is determined by the genotype of the mother. This is due to gene products deposited in the egg during oogenesis, which influence early development.
Key Point: The father's genotype and the offspring's own genotype do not affect the phenotype in maternal effect traits.
Mechanism: Maternal gene products (mRNA or proteins) are present in the egg and direct early embryonic development.
Dosage Compensation
Mechanisms and Examples
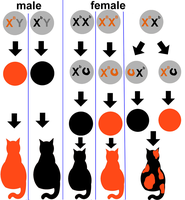
Dosage compensation ensures equal expression of X-linked genes in males (XY) and females (XX). Different species use different mechanisms to achieve this balance.
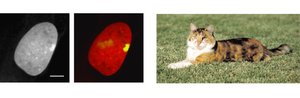
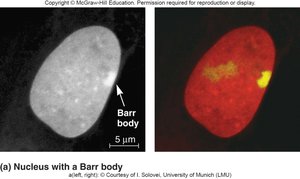
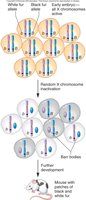
Mammals: One X chromosome in females is inactivated (Barr body formation).
Drosophila: The single X chromosome in males is hyperactivated.
C. elegans: Both X chromosomes in hermaphrodites are partially downregulated.
Example: In mammals, X inactivation leads to mosaic phenotypes, such as the coat color patterns in calico cats.

Genomic Imprinting
Definition and Mechanism
Genomic imprinting is an epigenetic phenomenon where only one allele of a gene is expressed, depending on its parental origin. The silenced allele is typically marked by DNA methylation during gamete formation.
Monoallelic Expression: Only the maternal or paternal allele is expressed, not both.
Resetting: Imprints are erased and re-established in each generation during gametogenesis.
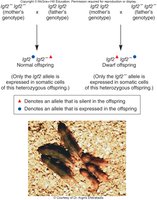
Example: Igf-2 Gene in Mice
The Igf-2 gene encodes insulin-like growth factor 2, necessary for normal growth. Only the paternal allele is expressed; the maternal allele is silenced. If the paternal allele is mutated, the offspring will be dwarfed, regardless of the maternal allele.
Imprinting and Human Disease
Genomic imprinting can influence the inheritance of certain human diseases, such as Prader-Willi syndrome (PWS) and Angelman syndrome (AS), both associated with deletions on chromosome 15. The phenotype depends on whether the deletion is inherited from the mother or the father.
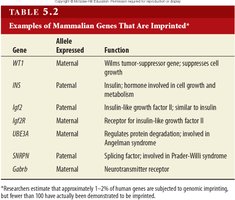
Gene | Allele Expressed | Function |
|---|---|---|
WT1 | Maternal | Wilms tumor-suppressor gene; suppresses cell growth |
INS | Maternal | Insulin; hormone involved in cell growth and metabolism |
Igf2 | Paternal | Insulin-like growth factor II; similar to insulin |
Igf2r | Maternal | Receptor for insulin-like growth factor II |
UBE3A | Maternal | Regulates protein degradation; involved in Angelman syndrome |
SNRPN | Paternal | Splicing factor; involved in Prader-Willi syndrome |
Gabrb3 | Maternal | Neurotransmitter receptor |
Extranuclear (Cytoplasmic) Inheritance
Organelle Genetics: Mitochondria and Chloroplasts
Extranuclear inheritance involves genes located in organelles such as mitochondria and chloroplasts. These organelles contain their own DNA, which is inherited independently of nuclear chromosomes, usually from the mother.
Mitochondrial DNA (mtDNA): Small, circular genome; codes for rRNAs, tRNAs, and proteins involved in oxidative phosphorylation.
Chloroplast DNA (cpDNA): Larger than mtDNA; codes for rRNAs, tRNAs, and proteins required for photosynthesis.
Maternal Inheritance: Offspring inherit organelle genomes almost exclusively from the mother, as the egg contributes most of the cytoplasm.
Maternal Inheritance vs. Maternal Effect
Maternal Inheritance: Transmission of organelle genes (e.g., mtDNA, cpDNA) from mother to offspring.
Maternal Effect: Phenotype determined by maternal nuclear genotype due to gene products in the egg.
Example: Pigmentation in Four O'Clock Plants
Carl Correns discovered that leaf pigmentation in Mirabilis jalapa is determined by the maternal parent, not the paternal parent. This is due to the inheritance of chloroplasts through the egg cytoplasm.
Human Mitochondrial Diseases
Mutations in mitochondrial DNA can cause a variety of chronic degenerative diseases, especially affecting tissues with high energy demands, such as nerves and muscles. These diseases are strictly maternally inherited.
Disease | Mitochondrial Gene Mutated |
|---|---|
Leber hereditary optic neuropathy | ND1, ND2, CO1, ND4, ND5, ND6 |
Neuropgenic muscle weakness | ATPase gene |
Mitochondrial encephalomyopathy | tRNA genes for leucine and lysine |
Mitochondrial myopathy | tRNA for leucine |
Maternally inherited diabetes and deafness | tRNA for leucine |
Myoclonic epilepsy with ragged-red muscle fibers | tRNA for lysine |

The Endosymbiosis Theory
Origin of Mitochondria and Chloroplasts
The endosymbiosis theory proposes that mitochondria and chloroplasts originated from free-living bacteria that were engulfed by ancestral eukaryotic cells. Over time, these bacteria became integrated as organelles.
Evidence: Organelle genomes are circular and resemble bacterial DNA; organelle genes are more similar to bacterial genes than to nuclear genes.
Gene Transfer: Many genes originally present in organelle genomes have been transferred to the nuclear genome.
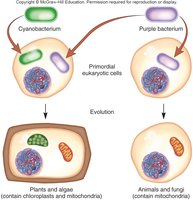
Additional info: The unidirectional transfer of genes from organelles to the nucleus has resulted in most organelle proteins now being encoded by nuclear genes and imported into the organelles after translation.
