Back
BackPopulation and Evolutionary Genetics: Forces Shaping Genetic Variation
Study Guide - Smart Notes

Population and Evolutionary Genetics
Hardy–Weinberg Law and Population Gene Pools
The Hardy–Weinberg law describes how allele and genotype frequencies remain constant from generation to generation in an ideal population, provided that certain assumptions are met. These assumptions include random mating, no selection, no mutation, no migration, and an infinitely large population size. Deviations from these assumptions can lead to changes in allele frequencies over time, which is the basis for evolutionary change.
Allele frequency: The proportion of a specific allele among all alleles at a genetic locus in a population.
Genotype frequency: The proportion of a specific genotype among all individuals in a population.
Hardy–Weinberg equilibrium provides a null model for detecting evolutionary forces.
Forces Affecting Allele Frequency
Several evolutionary forces can alter allele frequencies in populations:
Natural Selection: Differential survival and reproduction of individuals due to differences in phenotype. Selection can be natural (environment-driven) or artificial (human-imposed).
Genetic Drift: Random fluctuations in allele frequencies, especially significant in small populations. Includes founder effect and genetic bottleneck.
Migration (Gene Flow): Movement of individuals between populations, introducing new alleles and increasing genetic variation.
Mutation: The ultimate source of new genetic variation, creating new alleles.
Fitness and Selection
Fitness (w) is a measure of an individual's genetic contribution to future generations. Hardy–Weinberg analysis allows fitness to be quantified for different genotypes:
Fitness values: , ,
Example: , ,
Selection against deleterious alleles can be modeled mathematically. For a homozygous lethal allele ():
where is the frequency of allele a in generation g, is the initial frequency, and g is the number of generations.
Types of Selection
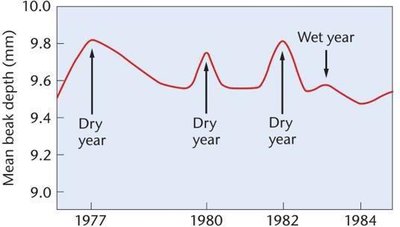
Directional Selection: Favors one extreme phenotype, shifting the population mean. Example: Beak size in finches increases during dry years due to selection for larger beaks.
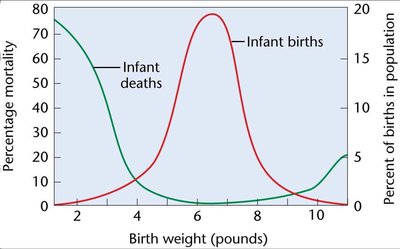
Stabilizing Selection: Favors intermediate phenotypes, reducing variance but not changing the mean. Example: Human birth weight, where both very low and very high weights have higher mortality.
Disruptive Selection: Favors both extremes over intermediates, leading to a bimodal distribution. Example: Selection for high and low bristle number in Drosophila.
Migration and Gene Flow
Migration introduces new alleles into a population, reducing genetic differences between populations and increasing genetic diversity. The change in allele frequency due to migration can be calculated as:
where is the new allele frequency on the island, is the original frequency, is the frequency in migrants, and m is the proportion of migrants.
Genetic Drift
Genetic drift refers to random changes in allele frequencies, especially in small populations. Two important forms are:
Founder Effect: A new population is established by a small number of individuals, carrying only a subset of the genetic variation.
Genetic Bottleneck: A large population undergoes a drastic reduction in size, losing genetic diversity.
Mutation
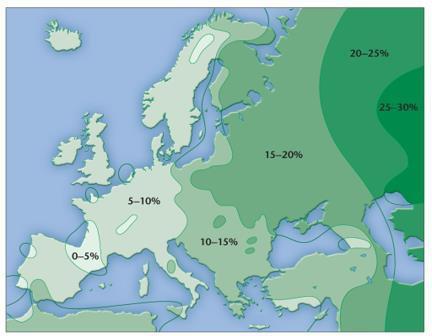
Mutation creates new alleles and is the ultimate source of genetic variation. The Hardy–Weinberg law allows estimation of allele and genotype frequencies, including the frequency of heterozygotes and carriers for recessive traits (e.g., cystic fibrosis).
Detecting Genetic Variation
Genetic variation can be detected by comparing DNA sequences among individuals. For example, the Adh gene in Drosophila melanogaster shows nucleotide differences among strains.
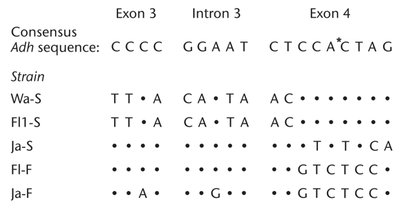
Molecular techniques such as PCR can identify carriers of genetic traits, as shown in studies of albinism in the Navajo population.
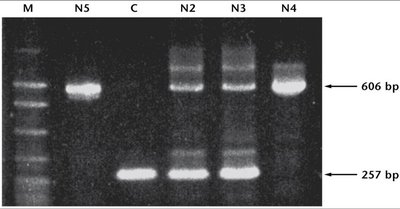
Neutral Theory of Molecular Evolution
The neutral theory proposes that most evolutionary changes at the molecular level are due to genetic drift of neutral mutations, rather than selection. The frequency of neutral alleles is determined by mutation rates and drift.
Microevolution and Macroevolution
Microevolution: Evolutionary changes within populations or species.
Macroevolution: Evolutionary events leading to the formation of new species and higher taxonomic groups.
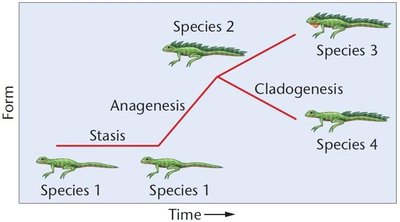
Speciation and Reproductive Isolation
Speciation is the formation of new species, often driven by reproductive isolating mechanisms that prevent gene flow between populations. These mechanisms can be ecological, behavioral, seasonal, mechanical, or physiological.

Geographic barriers, such as the formation of the Isthmus of Panama, can lead to speciation by dividing populations and preventing gene flow.
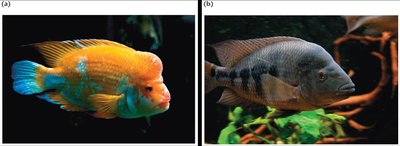

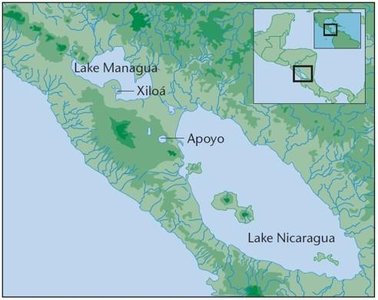
Phylogeny and Evolutionary History
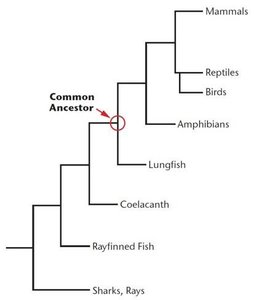
Phylogeny is the evolutionary history of a group of organisms. Phylogenetic trees are constructed using genetic data to infer relationships among species. Branches represent lineages, and nodes indicate common ancestors.
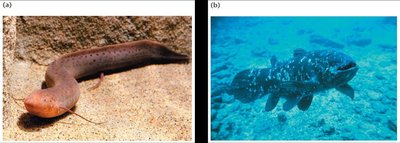

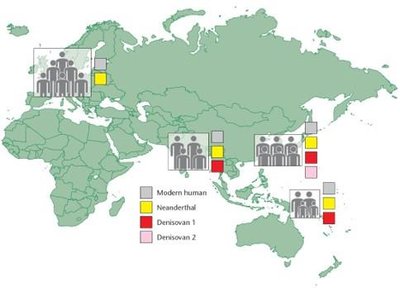
Complex Origins of the Human Genome
Modern humans (Homo sapiens) originated in Africa and migrated to other continents. Genomic evidence shows interbreeding with Neanderthals and Denisovans, with non-African populations carrying DNA from these extinct groups.