 Back
BackPost-Transcriptional Processing and Splicing in Eukaryotic mRNA
Study Guide - Smart Notes

Post-Transcriptional Processing of Eukaryotic mRNA
Overview of Pre-mRNA Processing
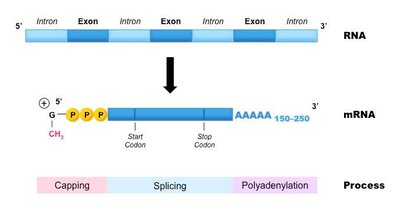
In eukaryotes, the initial RNA transcript (pre-mRNA) undergoes several modifications before becoming mature mRNA. These modifications are essential for mRNA stability, export from the nucleus, and translation efficiency. The three main processing steps are 5' capping, intron splicing, and 3' polyadenylation.
Pre-mRNA: The unprocessed transcript containing both exons (coding regions) and introns (non-coding regions).
Mature mRNA: The processed transcript, ready for translation, containing only exons, a 5' cap, and a 3' poly(A) tail.
5' Capping of mRNA
Biochemical Steps and Enzymes Involved
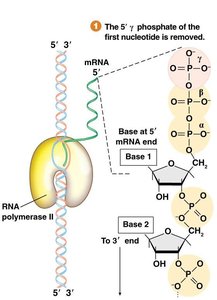
The 5' cap is a modified guanine nucleotide added to the 5' end of the pre-mRNA. This process occurs shortly after transcription initiation and involves several enzymatic steps:
RNA Triphosphatase removes the γ-phosphate from the 5' end of the nascent mRNA.
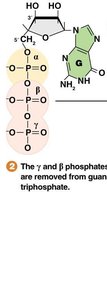
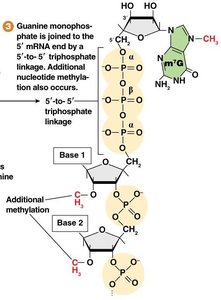
Guanylyl Transferase adds a GMP (guanosine monophosphate) in a 5'-to-5' triphosphate linkage.
Methyl Transferase methylates the guanine and sometimes the ribose sugars of the first few nucleotides.
Functions of the 5' Cap:
Protects mRNA from exonuclease degradation
Facilitates splicing of the first intron
Promotes export of mRNA from the nucleus
Essential for ribosome binding during translation initiation
3' Polyadenylation of mRNA
Steps and Enzymes Involved
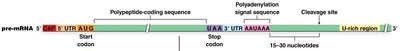
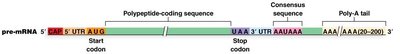
Polyadenylation is the addition of a poly(A) tail to the 3' end of the pre-mRNA. This process is directed by a conserved polyadenylation signal sequence (AAUAAA) located downstream of the coding region.
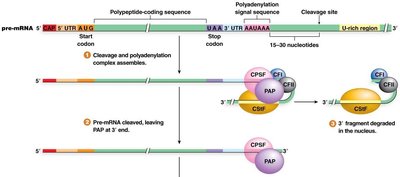
Recognition of the polyadenylation signal by protein factors (CPSF, CstF, CFI, CFII).
Cleavage of the pre-mRNA 15–30 nucleotides downstream of the signal.
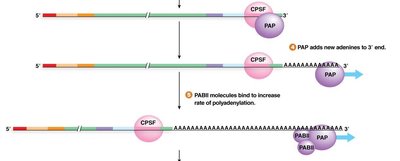
Addition of 20–200 adenine nucleotides by Poly(A) Polymerase (PAP).
Binding of Poly(A) Binding Proteins (PABP) to the tail, enhancing stability and translation.
Functions of Polyadenylation:
Enhances export of mRNA from the nucleus
Protects mRNA from degradation
Facilitates translation by aiding ribosome recruitment
Torpedo Model of Transcription Termination
Mechanism of Termination
After cleavage and polyadenylation, the residual RNA attached to RNA Polymerase II is degraded by a 5'→3' exonuclease (Torpedo RNase). This enzyme rapidly digests the leftover transcript, eventually causing RNA Polymerase II to dissociate from the DNA template, thus terminating transcription.
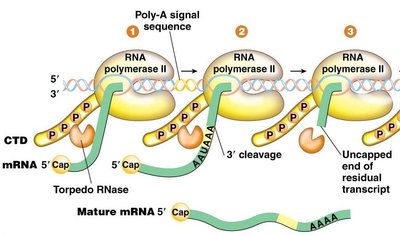
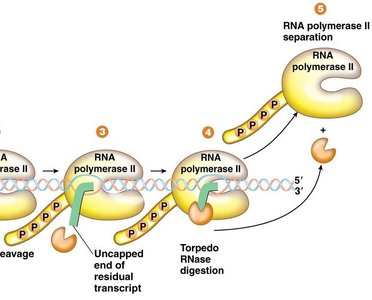
Intron Splicing in Pre-mRNA
Discovery and Importance
Splicing removes non-coding introns from pre-mRNA, joining exons to form mature mRNA. This process is highly precise, as errors can lead to frameshifts or nonfunctional proteins. The discovery of split genes (exons and introns) was a major milestone in molecular biology.
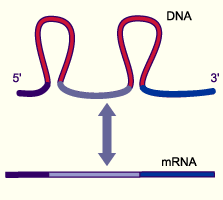
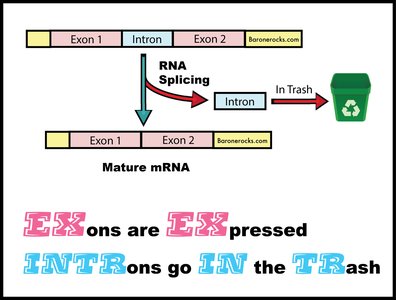
Splicing Signal Sequences
Splicing is guided by conserved sequences at the exon-intron boundaries:
5' Splice Site: GU dinucleotide at the 5' end of the intron
3' Splice Site: AG dinucleotide at the 3' end of the intron
Branch Site: An adenine (A) residue within a pyrimidine-rich region, 20–40 nucleotides upstream of the 3' splice site
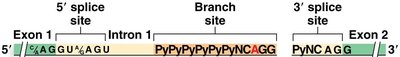
The Spliceosome and the Splicing Reaction
Spliceosome Composition and Function
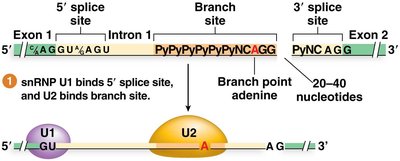
The spliceosome is a large ribonucleoprotein complex composed of five small nuclear ribonucleoproteins (snRNPs: U1, U2, U4, U5, U6). It catalyzes the two-step splicing reaction:
U1 snRNP binds the 5' splice site; U2 snRNP binds the branch site.
U4, U5, and U6 snRNPs join to form the inactive spliceosome.
Spliceosome rearranges, U1 and U4 leave, and the 5' splice site is cleaved, forming a lariat structure.
The 3' splice site is cleaved, and exons are ligated together.
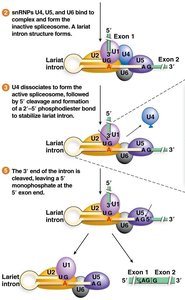
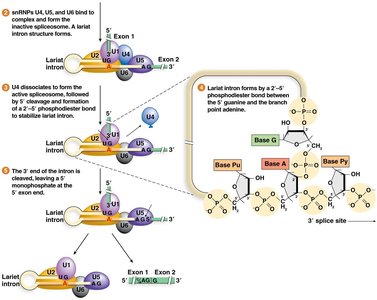
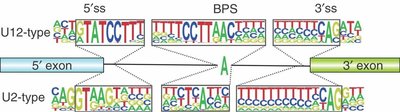
Types of Spliceosomes
Major Spliceosome (U2-type): Removes 99.5% of introns
Minor Spliceosome (U12-type): Removes rare U12-type introns
Coupling of Pre-mRNA Processing Steps
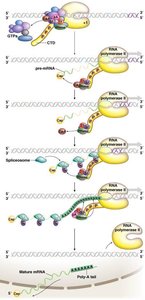
Coordination of Capping, Splicing, and Polyadenylation
The carboxyl terminal domain (CTD) of RNA Polymerase II acts as a platform for the assembly of processing factors. Capping, splicing, and polyadenylation are coordinated with transcription, ensuring efficient and accurate mRNA maturation.
Alternative Splicing and mRNA Diversity
Mechanisms and Biological Significance
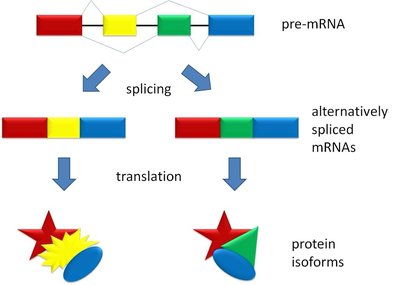
Alternative splicing allows a single gene to produce multiple mRNA and protein isoforms by varying the combination of exons included in the mature transcript. This mechanism greatly expands proteomic diversity and enables tissue-specific gene expression.
Alternative promoters, splicing, and polyadenylation sites contribute to transcript diversity.
Approximately 95% of human genes undergo alternative splicing.
Dysregulation of alternative splicing can lead to disease.
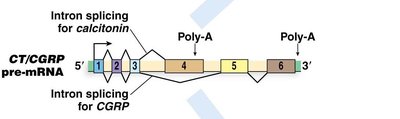
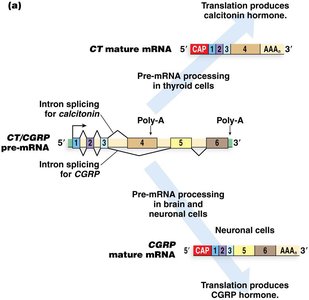
Example: CT/CGRP Gene
The human calcitonin/calcitonin gene-related peptide (CT/CGRP) gene uses alternative splicing and polyadenylation to produce two distinct hormones in different tissues.
Thyroid cells use exon 4 polyadenylation to produce calcitonin.
Neuronal cells splice out exon 4 and use exon 6 polyadenylation to produce CGRP.
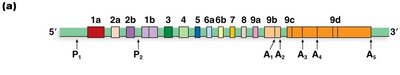
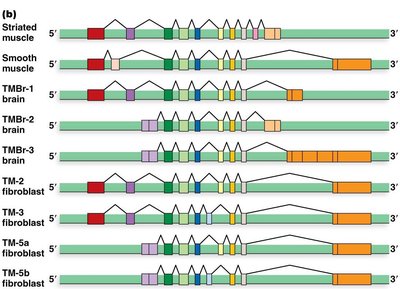
Example: Tropomyosin Gene
The rat α-tropomyosin gene undergoes alternative splicing to produce different isoforms in muscle and non-muscle cells.
Intron Self-Splicing and RNA Editing
Group I Introns and Ribozymes
Some introns, such as Group I introns, can catalyze their own excision without the spliceosome. These self-splicing introns are found in some mRNAs, tRNAs, and rRNAs of lower eukaryotes and plants.
RNA Editing
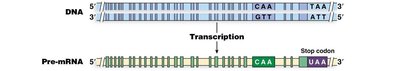
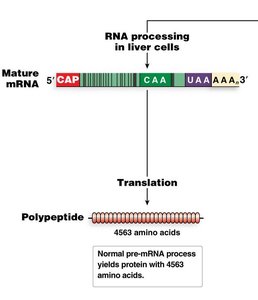
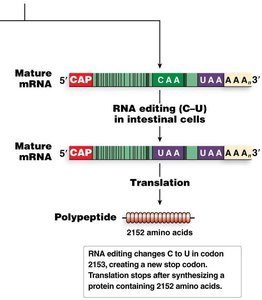
RNA editing involves post-transcriptional changes to nucleotide sequences, such as cytosine-to-uracil (C-to-U) conversion. An example is the editing of the human apolipoprotein B mRNA, which results in two different protein products in liver and intestine.
Summary Table: Key Steps in Eukaryotic mRNA Processing
Step | Enzyme/Complex | Key Sequence/Signal | Main Function |
|---|---|---|---|
5' Capping | RNA Triphosphatase, Guanylyl Transferase, Methyl Transferase | 5' end of pre-mRNA | Stability, ribosome binding, nuclear export |
Splicing | Spliceosome (snRNPs) | 5' GU, 3' AG, branch site A | Removes introns, joins exons |
Polyadenylation | Poly(A) Polymerase, CPSF, CstF, CFI, CFII | AAUAAA signal | Stability, translation, nuclear export |
