 Back
BackRecombinant DNA Technology and Its Applications: Study Notes
Study Guide - Smart Notes

Recombinant DNA Technology Basics
Introduction to Recombinant DNA Technology
Recombinant DNA technology encompasses a suite of molecular techniques for manipulating DNA sequences both in vitro (outside living organisms) and in vivo (within living organisms). These methods are foundational to modern genetics, biotechnology, and medicine, enabling the study and engineering of genes for research, therapeutic, and industrial purposes.
Key Principle: Combining DNA from different sources to create new genetic combinations.
Applications: Gene cloning, genetic modification of organisms, gene therapy, and the production of pharmaceuticals.
Foundational Tools: Restriction enzymes and plasmids are essential for cutting and pasting DNA.
Restriction Enzymes
Definition and Function
Restriction enzymes (restriction endonucleases) are proteins that recognize specific DNA sequences and make double-stranded cuts at or near these sites. They were first discovered in bacteria, where they serve as a defense mechanism against invading viral DNA.
Recognition Sites: Typically 4-8 base pairs, often palindromic (the sequence reads the same on both strands in the 5' to 3' direction).
Biological Role: Restrict the growth of bacteriophages by cleaving their DNA.
Laboratory Use: Generate DNA fragments for mapping, cloning, and analysis.
Types and Cutting Mechanisms
Type I: Cut far from recognition site (rarely used in labs).
Type II: Cut within the recognition site (most commonly used in molecular biology).
Type IIS: Cut at a defined distance from the recognition site.
Type III: Cut a short distance from the recognition site.
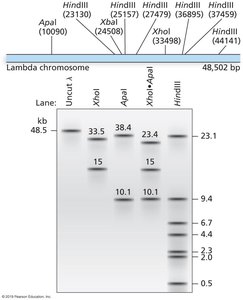
Restriction Mapping and Fragment Analysis
Restriction enzymes are used to generate restriction maps, which are diagrams showing the locations of cut sites within a DNA molecule. This is especially useful for small genomes, such as bacteriophage lambda.
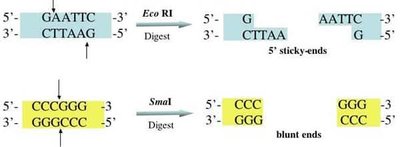
Sticky Ends vs. Blunt Ends
Restriction enzymes can produce two types of DNA ends:
Sticky ends: Single-stranded overhangs that can base-pair with complementary sequences, facilitating ligation.
Blunt ends: Straight cuts with no overhangs, which can be joined but are less efficient for ligation.
Molecular Cloning
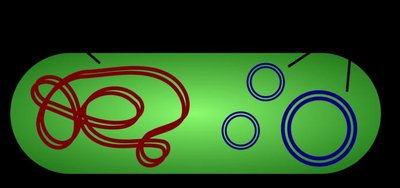
Plasmids as Cloning Vectors
Plasmids are small, circular DNA molecules found in bacteria and some eukaryotes. They replicate independently of the host genome and can be engineered to carry foreign DNA for cloning purposes.
Key Features: Origin of replication, selectable marker genes (e.g., antibiotic resistance), and multiple cloning sites (MCS).
Role as Vectors: Plasmids are used to introduce new DNA into host cells for replication and expression.

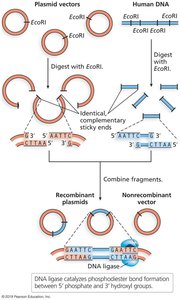
Steps in Molecular Cloning
Isolation of target gene and plasmid DNA.
Restriction digestion of both DNA sources to create compatible ends.
Ligation of the target gene into the plasmid using DNA ligase.
Transformation of recombinant plasmid into host bacterial cells.
Selection and screening for successful transformants.
Selection and Screening
Engineered plasmids often contain selectable markers (e.g., antibiotic resistance) and reporter genes (e.g., lacZ for blue/white screening) to identify successful recombinants.
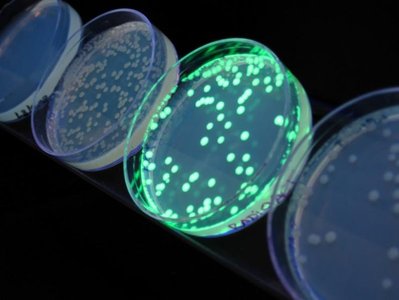
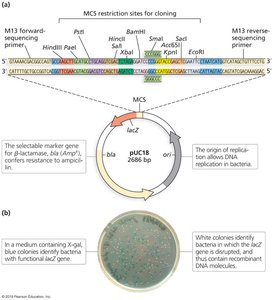
Amplification of Recombinant DNA
Once inside the host cell, recombinant plasmids are amplified by:
Replication of the plasmid within the cell.
Cell division, producing many copies of the recombinant DNA.
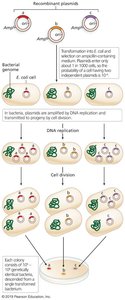
Cloning with Larger Vectors
For larger DNA fragments, alternative vectors such as bacteriophage lambda, cosmids, BACs (bacterial artificial chromosomes), and YACs (yeast artificial chromosomes) are used. These allow the cloning of larger genomic regions.
Recombinant DNA Libraries
Genomic and cDNA Libraries
A DNA library is a collection of cloned DNA fragments from a single source, stored in host cells. There are two main types:
Genomic libraries: Contain fragments representing the entire genome.
cDNA libraries: Contain DNA copies of mRNA, representing expressed genes.
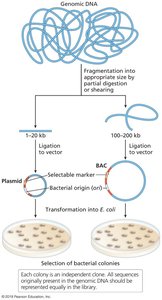
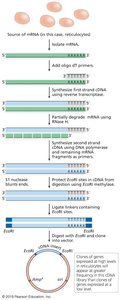
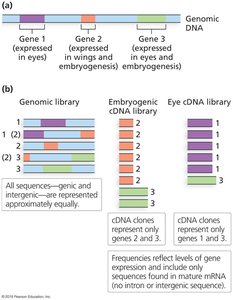
Applications of Libraries
Genomic libraries: Used for sequencing, mapping, and gene discovery.
cDNA libraries: Used to study gene expression in different tissues or developmental stages.
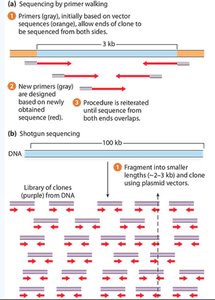
Sequencing Strategies
Primer walking: Sequential sequencing from each end of a clone using newly designed primers.
Shotgun sequencing: Random fragmentation and sequencing of DNA, followed by computational assembly.
Transgenic Organisms
Definition and Creation
A transgenic organism contains a gene (transgene) from another species, integrated into its genome. This process requires integration of foreign DNA and appropriate regulatory sequences for expression.
Transgenic Bacteria
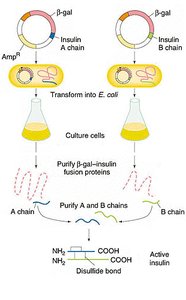
Escherichia coli is widely used to produce recombinant proteins, such as human insulin. The gene for insulin is inserted into a plasmid and expressed in bacteria, which can then be harvested for the protein product.
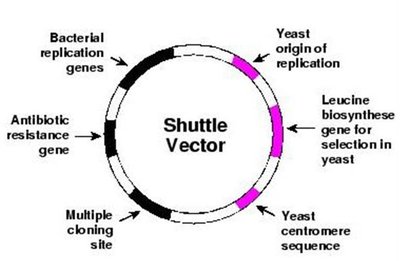
Transgenic Fungi and Shuttle Vectors
Yeast, such as Saccharomyces cerevisiae, can also be engineered to produce recombinant proteins. Shuttle vectors allow DNA to be transferred and expressed in both yeast and bacteria.
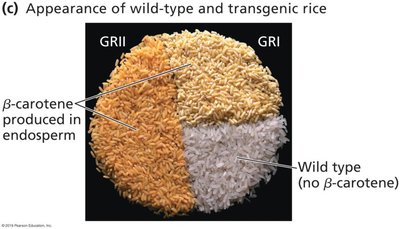
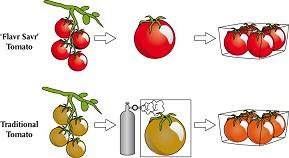
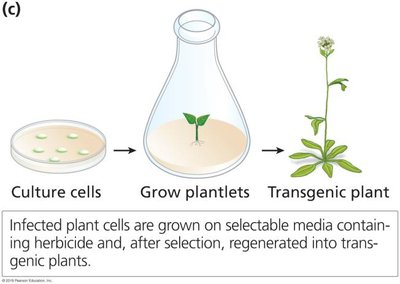
Transgenic Plants
Genetically modified plants (GMOs) are engineered for traits such as herbicide resistance, pest resistance, and improved nutrition. The 'Flavr Savr' tomato and 'Golden Rice' are notable examples.

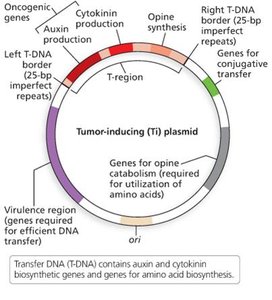
Plant Transformation via Agrobacterium
The Agrobacterium tumefaciens Ti plasmid is commonly used to introduce genes into plants. The T-DNA region is engineered to carry the gene of interest, and plant cells are transformed using methods such as electroporation or gene gun.



Transgenic Animals
Transgenic animals are less common than plants but are used for research and biotechnology. Examples include GM salmon with enhanced growth and transgenic mice for gene function studies.

Targeted DNA Modifications and Gene Therapy
CRISPR-Cas9 Gene Editing
The CRISPR-Cas9 system is a revolutionary gene-editing tool derived from bacterial immune systems. It uses a guide RNA to direct the Cas9 nuclease to a specific DNA sequence, where it introduces a double-stranded break. Repair can result in gene disruption (NHEJ) or precise editing (HDR).
Applications: Gene knockout, gene correction, and insertion of new sequences.
Ethical Considerations: Use in human embryos is controversial and highly regulated.
Gene Drives
Gene drives use CRISPR to ensure a genetic trait is inherited at a rate higher than Mendelian inheritance, rapidly spreading the trait through a population. This has potential applications in disease control (e.g., malaria eradication).
Gene Therapy
Gene therapy aims to treat genetic diseases by introducing, removing, or altering genes within an individual's cells. Delivery methods include viral vectors, liposomes, and nanoparticles. Major challenges include targeted delivery, control of gene expression, and immune responses.
Genetic Cloning
Cloning in Plants and Animals
Plant cloning is facilitated by the totipotency of plant cells, allowing regeneration from single cells in tissue culture. Animal cloning is more complex, often involving somatic cell nuclear transfer into enucleated egg cells.
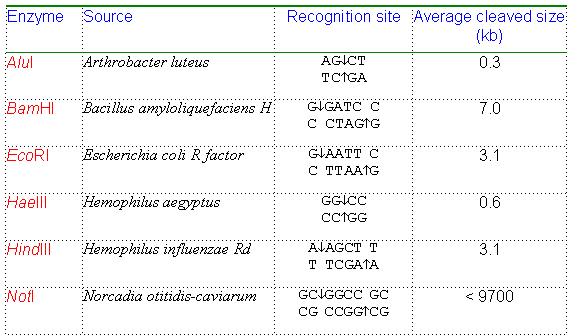
Summary Table: Common Restriction Enzymes
Enzyme | Source | Recognition Site | Average Cleaved Size (kb) |
|---|---|---|---|
AluI | Arthrobacter luteus | AGCT TCGA | 0.3 |
BamHI | Bacillus amyloliquefaciens H | GGATCC CCTAGG | 7.0 |
EcoRI | Escherichia coli R factor | GAATTC CTTAAG | 3.1 |
HaeIII | Hemophilus aegyptus | GGCC CCGG | 0.6 |
HindIII | Hemophilus influenzae Rd | AAGCTT TTCGAA | 3.1 |
NotI | Nocardia otitidis-caviarum | GCGGCCGC CGCCGGCG | <9700 |
