 Back
BackRecombinant DNA Technology: Tools, Methods, and Applications
Study Guide - Smart Notes

Recombinant DNA Technology
Introduction to Recombinant DNA Technology
Recombinant DNA technology refers to the artificial joining of DNA molecules from different biological sources, creating combinations not found in nature. This technology has revolutionized molecular biology and genetics, enabling the isolation, replication, and analysis of specific genes. It is foundational to the biotechnology industry and has numerous applications in research, medicine, and agriculture.
Restriction enzymes and cloning vectors are the two key tools that made recombinant DNA technology possible.
Restriction enzymes cut DNA at specific sequences, while vectors carry DNA fragments into host cells for cloning.
Cloning allows for the production of exact genetic copies of genes, cells, tissues, or whole organisms.
Restriction Enzymes
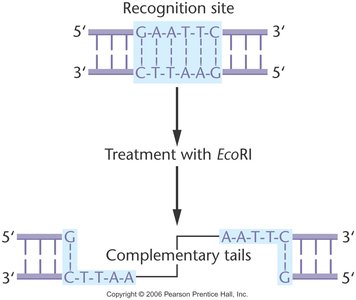
Restriction enzymes, also known as restriction endonucleases, are produced by bacteria as a defense mechanism against bacteriophage infection. They recognize and cleave DNA at specific nucleotide sequences called restriction sites, which are typically palindromic.
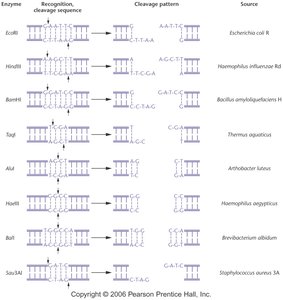
Recognition sequence: Usually 4–8 base pairs long and palindromic (reads the same on both strands).
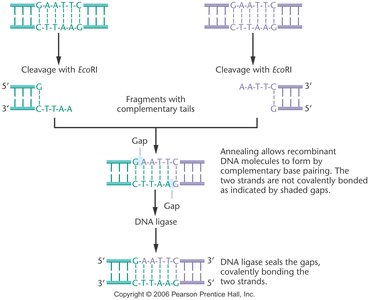
Cleavage pattern: Enzymes can produce sticky (cohesive) ends with overhangs or blunt ends with no overhangs.
DNA ligase: Enzyme that covalently joins restriction fragments, sealing the phosphodiester backbone to produce intact recombinant DNA molecules.
Cloning Vectors
Vectors are carrier DNA molecules that allow DNA restriction fragments to enter host cells for cloning. They must be able to replicate independently, contain multiple restriction sites, and carry selectable marker genes or reporter genes for identification.
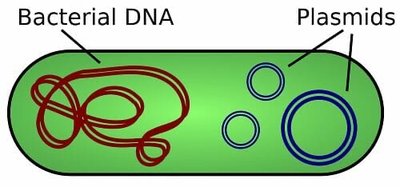
Bacterial plasmids: Small, circular, double-stranded DNA molecules that replicate independently of the bacterial chromosome. They are the most commonly used vectors for DNA cloning.
Phage vectors: Derived from bacteriophage λ, these vectors can carry larger DNA fragments than plasmids and infect bacterial cells efficiently.
BACs and YACs: Bacterial artificial chromosomes (BACs) and yeast artificial chromosomes (YACs) are used to clone very large DNA fragments, useful in genome projects.
Expression vectors: Designed to ensure transcription and translation of the cloned gene, allowing for protein production in host cells.
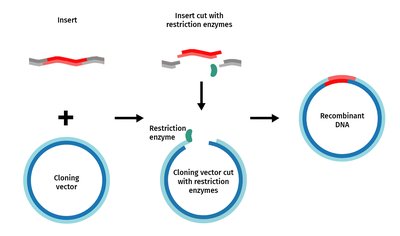
Transformation and Cloning Process
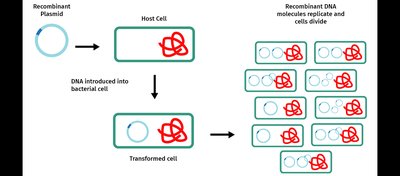
Transformation is the process by which plasmids are introduced into bacterial cells. Cells are made competent to take up DNA using calcium ions and heat shock or electroporation. Once inside, recombinant DNA molecules replicate as the host cells divide, producing many copies (clones) of the DNA fragment of interest.
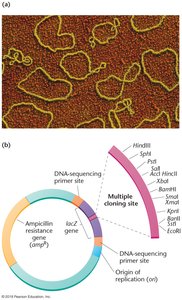
Selectable markers: Genes that confer antibiotic resistance or produce a visible effect, allowing identification of cells that have taken up the vector.
Reporter genes: Genes such as lacZ that encode easily detectable proteins, used in blue-white screening to distinguish recombinant from nonrecombinant clones.
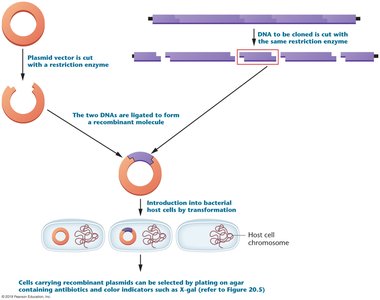
Blue-White Screening
Blue-white screening is a common method for identifying recombinant bacterial colonies. It utilizes the lacZ gene, which encodes β-galactosidase. When X-gal is present in the medium, colonies with functional lacZ turn blue, while those with disrupted lacZ (due to insertion of foreign DNA) remain white.
Blue colonies: Nonrecombinant plasmids with intact lacZ.
White colonies: Recombinant plasmids with disrupted lacZ due to DNA insertion.


Cloning in Nature and Applications
Cloning occurs naturally in single-celled organisms and some multicellular organisms. In biotechnology, cloning is used to produce genetically identical organisms or cells for research and agriculture.
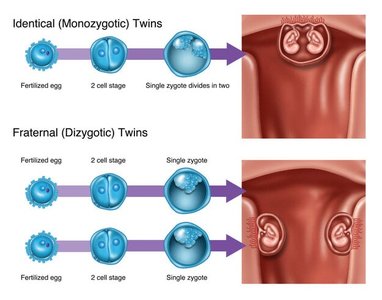
Monozygotic twins: Naturally occurring clones in humans and animals.
Crop plants and trees: Many are propagated by cloning to maintain desirable traits.
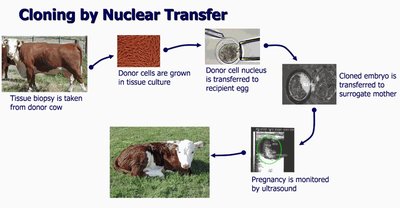
Dolly the Sheep: The first mammal cloned from an adult somatic cell using nuclear transfer.

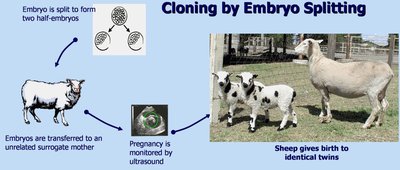
Cloning Methods
There are several methods for cloning animals, including embryo splitting and nuclear transfer.
Embryo splitting: Early embryo is divided to produce genetically identical offspring.
Nuclear transfer: Nucleus from a donor cell is transferred to an enucleated egg, which is then implanted into a surrogate mother.
Other Types of Cloning Vectors
Besides plasmids, other vectors are used for specialized applications:
Phage vectors: Used for cloning larger DNA fragments and efficient infection of bacterial hosts.
BACs and YACs: Used for cloning very large DNA fragments, essential for genome mapping and sequencing projects.
Expression vectors: Designed for gene expression in prokaryotic or eukaryotic hosts, enabling protein production.
Ti plasmid vectors: Used for transforming plant cells, derived from Rhizobium radiobacter (formerly Agrobacterium tumefaciens).
DNA Libraries
DNA libraries are collections of cloned DNA fragments representing the genetic material of an organism. There are two main types:
Genomic libraries: Contain at least one copy of all sequences in the genome.
cDNA libraries: Contain complementary DNA copies made from mRNA, representing genes actively transcribed at the time of mRNA isolation.
Screening DNA Libraries
To isolate a specific gene from a DNA library, researchers use probes—short DNA or RNA sequences complementary to the gene of interest. The probe hybridizes with the target sequence, allowing identification and recovery of the desired clone.
Probes: Can be labeled with radioactive or fluorescent tags for detection.
Library screening: Involves transferring colonies to a filter, hybridizing with the probe, and identifying positive clones.
