 Back
BackRecombinant DNA Technology: Tools, Methods, and Applications
Study Guide - Smart Notes

Recombinant DNA Technology
Introduction to Recombinant DNA Technology
Recombinant DNA technology refers to the artificial joining of DNA molecules from different biological sources, creating combinations not found in nature. This technology has revolutionized molecular biology and genetics, enabling the isolation, replication, and analysis of specific genes. It is foundational to the biotechnology industry and has numerous applications in research, medicine, and agriculture.
Restriction enzymes and cloning vectors are the two key tools that made recombinant DNA technology possible.
Restriction enzymes cut DNA at specific sequences, while vectors carry DNA fragments into host cells for cloning.
Cloning allows for the production of large quantities of identical DNA molecules (clones) for study.
Historical Perspective
Restriction enzymes were first isolated in the 1970s by Kathleen Danna and Daniel Nathans.
These enzymes were shown to cleave viral DNA at specific nucleotide sequences, providing a defense mechanism for bacteria against bacteriophages.
Scientists soon discovered how to use these enzymes to cut DNA from any organism at specific sites, enabling the creation of recombinant DNA molecules.
Key Tools in Recombinant DNA Technology
Restriction Enzymes
Restriction enzymes, also known as restriction endonucleases, are proteins produced by bacteria to protect against viral infection by cleaving foreign DNA. They recognize specific, short DNA sequences (restriction sites) and cut both strands of DNA, generating fragments with either sticky or blunt ends.
More than 800 restriction enzymes are known, each recognizing a unique sequence (usually 4–8 base pairs).
Restriction sites are often palindromic, meaning the sequence reads the same on both strands in the 5' to 3' direction.
Cuts can result in sticky ends (overhangs) or blunt ends (no overhangs).
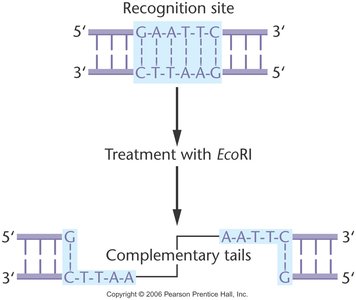
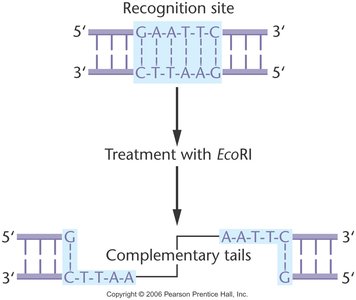
Example: EcoRI Restriction Enzyme
EcoRI recognizes the sequence 5'-GAATTC-3' and cuts between G and A, producing sticky ends.
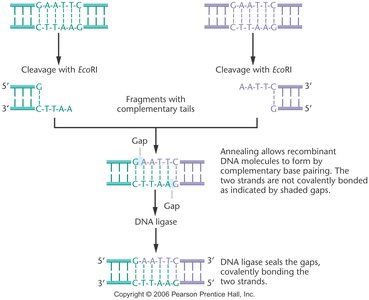
DNA Ligase
DNA ligase is an enzyme that joins DNA fragments by forming phosphodiester bonds between adjacent nucleotides. After restriction enzymes cut DNA, DNA ligase can seal the sticky or blunt ends, creating a stable recombinant DNA molecule.
Essential for covalently bonding recombinant DNA fragments.
Cloning Vectors
Vectors are carrier DNA molecules that allow foreign DNA fragments to be introduced into host cells for replication and analysis. The most common vectors are plasmids, but others include bacteriophage λ, BACs, YACs, and Ti plasmids for plants.
Vectors must replicate independently in the host cell.
They contain multiple restriction sites for DNA insertion and selectable marker genes for identifying successful clones.
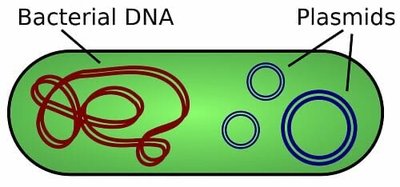
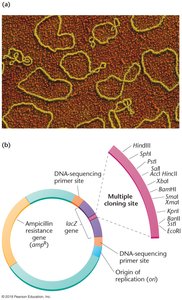
Bacterial Plasmids
Small, circular, double-stranded DNA molecules found in bacteria.
Replicate independently of the bacterial chromosome.
Engineered to include multiple cloning sites and selectable markers (e.g., antibiotic resistance genes).
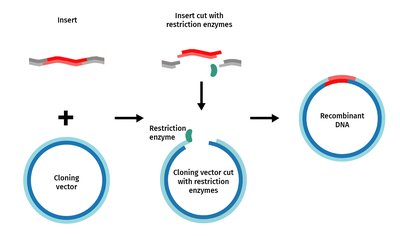
Steps in Creating Recombinant DNA
General Procedure
Cut both the vector and the DNA to be cloned with the same restriction enzyme to generate compatible ends.
Mix the DNA fragments and vector; DNA ligase is added to seal the nicks and form recombinant DNA.
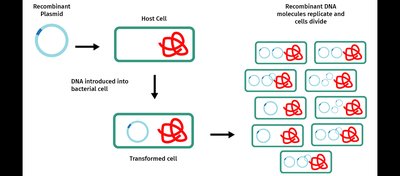
Introduce the recombinant DNA into a host cell (usually bacteria) by transformation.
Host cells replicate, producing many copies (clones) of the recombinant DNA.
Cloning and Clones
Definition and Examples
Cloning refers to the process of making exact genetic copies of DNA, cells, tissues, or whole organisms. Clones are genetically identical copies derived from a single original source.
Occurs naturally in bacteria (binary fission), some plants, and animals like Caenorhabditis elegans.
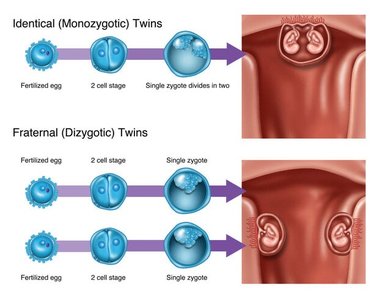
Examples include monozygotic twins, crop plants, and the famous cloned animal, Dolly the sheep.

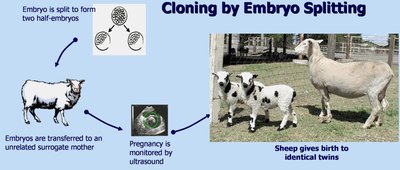
Methods of Cloning
Embryo Splitting: Early embryo is split into two, and each half is implanted into a surrogate mother, resulting in genetically identical offspring.
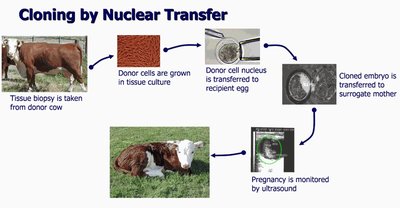
Nuclear Transfer: The nucleus from a donor cell is transferred into an enucleated egg cell, which is then implanted into a surrogate. This method was used to create Dolly the sheep.
Transformation and Selection
Transformation
Transformation is the process by which foreign DNA is introduced into a host cell. In bacteria, cells are made competent to take up DNA using chemical (calcium chloride and heat shock) or physical (electroporation) methods.
Transformed cells incorporate recombinant plasmids and replicate them, producing many copies.
Selectable Markers and Blue-White Screening
Selectable marker genes (e.g., antibiotic resistance) and reporter genes (e.g., lacZ) are used to identify cells that have taken up recombinant DNA. Blue-white screening distinguishes recombinant from nonrecombinant clones based on disruption of the lacZ gene, which encodes β-galactosidase.
Cells with nonrecombinant plasmids produce blue colonies on X-gal media; recombinant clones (with disrupted lacZ) produce white colonies.

Other Types of Cloning Vectors
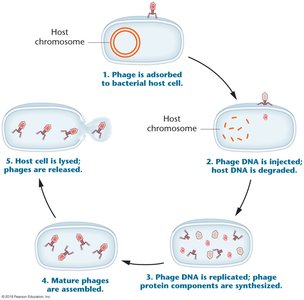
Phage Vectors
Bacteriophage λ vectors can carry larger DNA fragments (up to 45 kb) than plasmids and are used to infect bacterial hosts, forming plaques on bacterial lawns.
Central third of λ DNA can be replaced with foreign DNA without affecting infectivity.
BACs and YACs
Bacterial Artificial Chromosomes (BACs): Large, low-copy plasmids used for cloning very large DNA fragments (100–300 kb), important in genome projects.
Yeast Artificial Chromosomes (YACs): Linear vectors with telomeres, centromeres, and origins of replication, used for cloning up to 2 Mb of DNA.
Expression Vectors
Expression vectors are designed to ensure that the cloned gene is transcribed and translated in the host cell, producing large quantities of protein. They contain regulatory sequences for both transcription and translation.
Ti Plasmid Vectors for Plants
The Ti plasmid from Rhizobium radiobacter (formerly Agrobacterium tumefaciens) is used to introduce foreign genes into plant genomes. The tumor-inducing genes are removed, and the T-DNA region is replaced with the gene of interest.
DNA Libraries
Genomic and cDNA Libraries
DNA libraries are collections of cloned DNA fragments representing the genetic material of an organism. There are two main types:
Genomic Library: Contains at least one copy of every sequence in the genome, constructed by fragmenting genomic DNA and cloning into vectors.
cDNA Library: Contains DNA copies synthesized from mRNA, representing only the genes expressed in a particular cell type at a specific time.
Screening DNA Libraries
To isolate a specific gene from a library, researchers use labeled DNA or RNA probes that are complementary to the gene of interest. Colonies or plaques containing the desired gene are identified by hybridization with the probe.
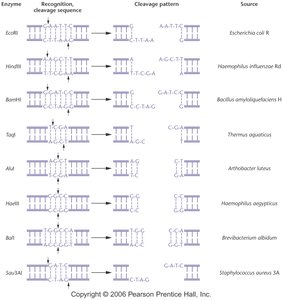
Summary Table: Common Restriction Enzymes
Enzyme | Recognition Sequence | Cleavage Pattern | Source |
|---|---|---|---|
EcoRI | GAATTC | Sticky ends | Escherichia coli R |
HindIII | AAGCTT | Sticky ends | Haemophilus influenzae Rd |
BamHI | GGATCC | Sticky ends | Bacillus amyloliquefaciens H |
AluI | AGCT | Blunt ends | Arthrobacter luteus |
HaeIII | GGCC | Blunt ends | Haemophilus aegyptius |
Key Equations
Palindromic sequence example:
Restriction enzyme cleavage:
