 Back
BackRegulation of Gene Expression and Cell Specialization: Prokaryotic and Eukaryotic Mechanisms
Study Guide - Smart Notes

Regulation of Gene Expression and Cell Specialization
Overview of Gene Expression
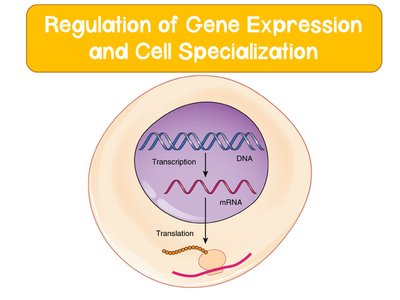
Gene expression is the process by which information from a gene is used to synthesize functional gene products, such as proteins. Both prokaryotic and eukaryotic cells regulate gene expression to respond to environmental and internal cues, enabling cell specialization and adaptation.
Gene regulation allows cells to turn genes "on" or "off" depending on need.
Transcription is the first step, where DNA is converted to mRNA.
Translation follows, producing proteins from mRNA.
Bacterial Gene Expression
Operons and Their Regulation
In bacteria, gene expression is often controlled by operons, which are clusters of genes regulated together. Operons consist of three main parts and can be repressible or inducible.
Promoter: Site where RNA polymerase binds to initiate transcription.
Operator: Acts as an on/off switch for transcription.
Genes: Code for enzymes involved in a metabolic pathway.
Operon Type | Transcription State | Regulation Mechanism |
|---|---|---|
Repressible | Usually ON | Can be turned OFF by a repressor |
Inducible | Usually OFF | Can be turned ON by an inducer |
Regulatory genes produce repressor proteins that bind to the operator, blocking transcription. This binding is reversible and allows dynamic control of gene expression.
Allosteric Regulation of Enzymes
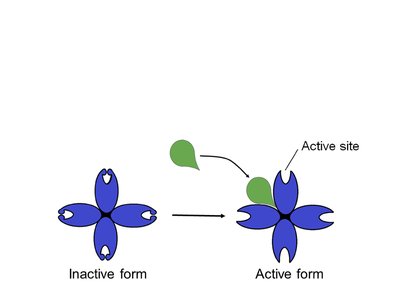
Allosteric regulation involves molecules binding to an enzyme at a site other than the active site, affecting enzyme activity. This is important in operon regulation.
Allosteric activator: Stabilizes the enzyme's active form, keeping active sites open.
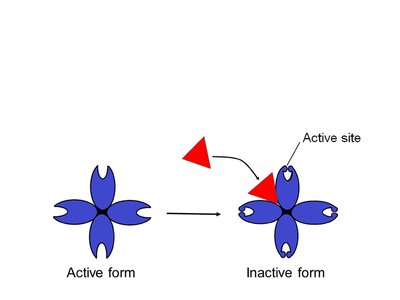
Allosteric inhibitor: Stabilizes the enzyme's inactive form, closing active sites.
Repressible Operons: The trp Operon
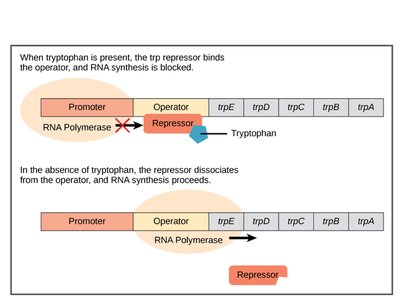
The trp operon controls the synthesis of tryptophan in bacteria. It is a repressible operon, meaning transcription is usually active but can be shut off when tryptophan levels are high.
When tryptophan accumulates, it binds to the trp repressor, activating it.
The active repressor binds to the operator, blocking RNA polymerase and halting transcription.
When tryptophan is scarce, the repressor is inactive, and transcription proceeds.
Inducible Operons: The lac Operon
The lac operon regulates the synthesis of lactase, which digests lactose. It is inducible, meaning transcription is usually off but can be turned on in the presence of lactose.
The lac repressor is normally bound to the operator, preventing transcription.
When allolactose (an inducer) is present, it binds to the repressor, inactivating it.
This allows RNA polymerase to transcribe the genes for lactose metabolism.
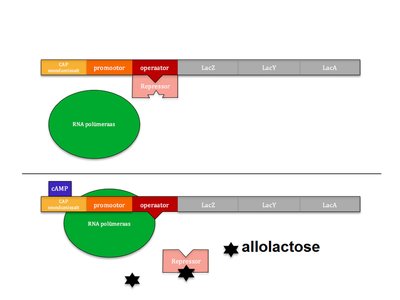
Comparison of trp and lac Operons
Both regulate transcription of genes in bacteria.
trp operon is repressible (usually ON, can be turned OFF).
lac operon is inducible (usually OFF, can be turned ON).
Eukaryotic Gene Expression
Differential Gene Expression
Eukaryotic cells regulate gene expression at multiple levels, resulting in cell specialization. The combination and level of gene expression determine the phenotype of a cell or organism.
Differential gene expression: Differences in gene expression between cell types.
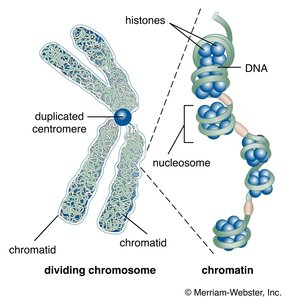
Chromatin Structure and Epigenetic Regulation
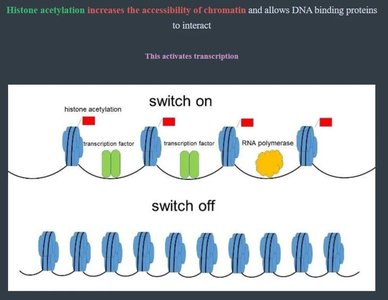
Chromatin structure affects gene accessibility. Modifications such as histone acetylation and DNA methylation regulate gene expression without altering the DNA sequence.
Histone acetylation: Adds acetyl groups to histones, loosening DNA and increasing transcription.
DNA methylation: Adds methyl groups to DNA, condensing chromatin and silencing genes.
These modifications are dynamic, heritable, and crucial for cell differentiation and development.
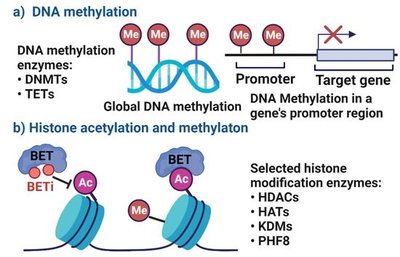
Epigenetic Inheritance
Epigenetic modifications can be inherited by future generations without changing the DNA sequence. These changes are reversible and explain phenomena such as differences in disease inheritance between identical twins.
Epigenetics acts as "sticky notes" on the DNA, guiding gene usage.
Unlike mutations, epigenetic changes can be reversed.
Transcription Initiation in Eukaryotes
Transcription initiation is regulated by transcription factors binding to control elements in non-coding DNA. The binding of activators or repressors to these elements can increase or decrease gene expression.
Control elements: Non-coding DNA sequences that serve as binding sites for transcription factors.
Gene expression is modulated by the presence of specific transcription factors.
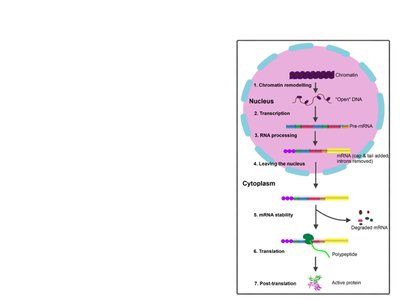
RNA Processing and Translation Regulation
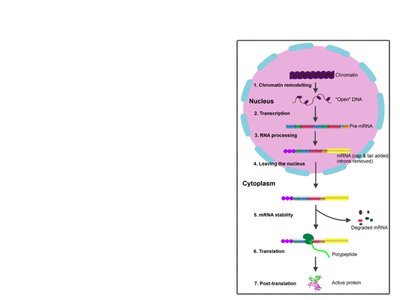
Gene expression in eukaryotes is further regulated during RNA processing and translation.
Alternative splicing: Pre-mRNA can be spliced in different ways to produce multiple protein variants.
Translation initiation: Can be activated or repressed by initiation factors.
MicroRNAs and small interfering RNAs: Bind to mRNA to degrade it or block translation.
Eukaryotic Development and Cell Differentiation
Cell Division, Differentiation, and Morphogenesis
During embryonic development, cells undergo division and differentiation, becoming specialized in structure and function. Morphogenesis is the process that shapes the organism.
Cytoplasmic determinants: Substances in the maternal egg that influence cell fate.
Induction: Cell-to-cell signals that alter gene expression.
Both mechanisms contribute to pattern formation and the establishment of the body plan.
Homeotic genes: Map out body structures during development.
Role of Apoptosis in Development
Apoptosis, or programmed cell death, is essential for proper development and formation of structures. For example, apoptosis shapes fingers and toes by removing webbing during human development.
Ensures correct formation of tissues and organs.
Prevents malformations by eliminating unnecessary cells.
