 Back
BackRegulation of Gene Expression in Bacteria: The lac Operon Model
Study Guide - Smart Notes

Regulation of Gene Expression in Bacteria
Introduction to Gene Regulation
Gene expression in bacteria is tightly regulated to ensure that proteins are produced only when needed, optimizing energy and resource use. This regulation allows bacteria to adapt rapidly to changing environmental conditions by turning genes on or off in response to specific signals.
Central Dogma and Gene Expression
The central dogma of molecular genetics describes the directional flow of genetic information from DNA to RNA to protein. Gene expression refers to the process by which information from a gene is used to synthesize RNA and protein products. Regulation of gene expression primarily occurs at the level of transcription and translation.
Basal Components of Transcription and Translation
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
RNA Polymerase: Enzyme that synthesizes RNA from a DNA template.
Ribosome: Cellular machinery for protein synthesis during translation.
These are required for all genes, but additional regulatory mechanisms determine when and how much a gene is expressed.
Types of Gene Expression
Constitutive vs. Regulated Genes
Constitutive genes: Expressed continuously, regardless of environmental conditions. Examples include genes for rRNA, tRNA, and RNA polymerase.
Regulated genes: Expressed only under certain conditions, such as in response to environmental changes (e.g., nutrient availability).
Constitutive expression is essential for basic cellular functions, while regulated expression allows adaptation to environmental changes.
Mechanisms of Gene Regulation
Positive and Negative Control
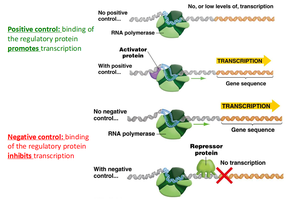
Gene expression can be regulated by proteins that either promote or inhibit transcription:
Positive control: Regulatory protein (activator) increases transcription when bound to DNA.
Negative control: Regulatory protein (repressor) decreases or prevents transcription when bound to DNA.
Operons in Prokaryotes
Structure and Function of Operons
An operon is a cluster of genes under the control of a single promoter and regulatory region, allowing coordinated expression of genes with related functions. Operons are a hallmark of prokaryotic gene regulation.
Promoter: Site where RNA polymerase binds.
Operator: DNA region where regulatory proteins bind to influence transcription.
Structural genes: Genes encoding proteins with related functions, transcribed as a single polycistronic mRNA.
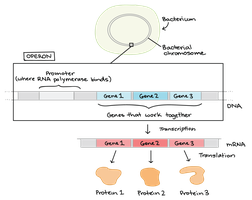
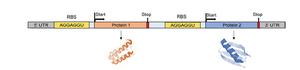
Polycistronic mRNA
In bacteria, operons are transcribed into a single mRNA molecule that encodes multiple proteins. This allows for efficient and coordinated regulation of gene clusters.
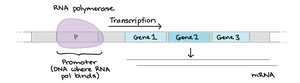
Regulatory DNA Elements: Operator, Promoter, and Regulatory Proteins
Operator and Promoter Regions
The operator is a regulatory DNA sequence where repressors or activators bind to control transcription. The promoter is the site for RNA polymerase binding. Together, these regions integrate signals to modulate gene expression.
Repressors and Activators
Repressors: Proteins that bind to the operator to block RNA polymerase, inhibiting transcription.
Activators: Proteins that enhance RNA polymerase binding to the promoter, increasing transcription.
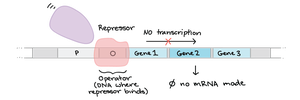

Cis- and Trans-Acting Elements
Cis-acting elements: DNA sequences (e.g., promoters, operators) that regulate genes on the same DNA molecule.
Trans-acting elements: Diffusible molecules (usually proteins) that bind to cis-acting sites to regulate gene expression.
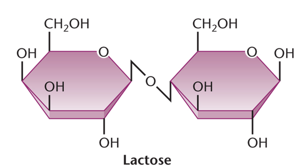
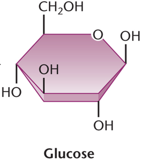
Lactose and Glucose Metabolism in E. coli
Inducible and Repressible Systems
E. coli prefers glucose as an energy source but can metabolize lactose when glucose is unavailable. The enzymes for lactose metabolism are produced only when lactose is present, demonstrating an inducible system.
Inducible enzymes: Synthesized in response to an inducer (e.g., lactose).
Repressible enzymes: Synthesized unless a corepressor is present.
The lac Operon: A Model of Inducible Gene Regulation
Structure of the lac Operon
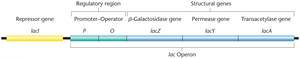
The lac operon in E. coli consists of three structural genes (lacZ, lacY, lacA) and regulatory regions (promoter, operator). The operon is regulated by the presence or absence of lactose.
lacZ: Encodes β-galactosidase, which cleaves lactose into glucose and galactose.
lacY: Encodes permease, which facilitates lactose entry into the cell.
lacA: Encodes transacetylase, involved in detoxification.
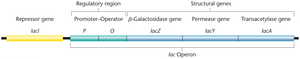
Regulation by the lacI Repressor
The lacI gene encodes a repressor protein that binds to the operator to inhibit transcription in the absence of lactose. When lactose is present, it binds to the repressor, causing a conformational change that prevents the repressor from binding the operator, thus allowing transcription.
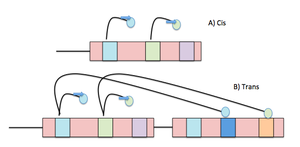
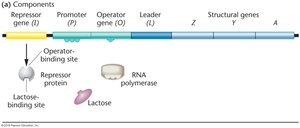
Negative Control of the lac Operon
The lac operon is an example of negative control. Transcription occurs only when the repressor is not bound to the operator. The presence of lactose (inducer) inactivates the repressor, permitting gene expression.
Summary Table: Components of the lac Operon
Component | Function |
|---|---|
lacZ | Encodes β-galactosidase (lactose hydrolysis) |
lacY | Encodes permease (lactose transport) |
lacA | Encodes transacetylase (detoxification) |
lacI | Encodes repressor protein (regulation) |
Operator | DNA site for repressor binding |
Promoter | RNA polymerase binding site |
Regulation Scenarios for the lac Operon
Lactose Absent
lacI repressor binds operator, blocking RNA polymerase.
No transcription of lac genes.
Lactose Present
Lactose binds repressor, causing it to release from the operator.
RNA polymerase transcribes lac genes, enabling lactose metabolism.
Example: Predicting lac Operon Activity
Glucose present, lactose absent: lac operon is OFF (repressed).
Glucose absent, lactose present: lac operon is ON (expressed).
Additional info: The lac operon is also subject to positive control by the catabolite activator protein (CAP), which further enhances transcription when glucose is absent. This is covered in more detail in advanced sections.
