 Back
BackRegulation of Gene Expression in Bacteria: The lac and trp Operons
Study Guide - Smart Notes

Regulation of Gene Expression in Bacteria
Introduction to Bacterial Gene Regulation
Gene expression in bacteria is tightly regulated to ensure that proteins are produced only when needed. This regulation allows bacteria to adapt quickly to changes in their environment, such as the availability of nutrients. Two classic examples of gene regulation in Escherichia coli are the lac operon and the trp operon, which illustrate inducible and repressible systems, respectively.
The lac Operon: Negative and Positive Control
Structure and Function of the lac Operon
The lac operon is an inducible operon responsible for the metabolism of lactose in E. coli. It consists of structural genes (lacZ, lacY, lacA), a promoter, an operator, and the regulatory gene lacI, which encodes the repressor protein.
Inducible system: The operon is usually off but can be turned on (induced) in the presence of lactose.
Negative control: The lacI repressor binds to the operator to block transcription when lactose is absent.
Inducer: Allolactose (a lactose derivative) binds to the repressor, causing it to release from the operator, allowing transcription.
Mutations Affecting the lac Operon
lacI DNA-binding domain mutation: If the repressor cannot bind DNA, the operon is constitutively expressed (always on).
Operator mutation: If the operator sequence is altered so the repressor cannot bind, the operon is also constitutively expressed.
Lactose-binding site mutation in lacI: If the repressor cannot bind lactose, it will always bind the operator, and the operon will never be expressed, even if lactose is present.
Cis-acting vs. Trans-acting Elements
Cis-acting elements: DNA sequences (e.g., operator, promoter) that affect expression of genes on the same DNA molecule.
Trans-acting elements: Diffusible molecules (e.g., lacI repressor protein) that can regulate genes on different DNA molecules.
lacI is trans-acting because the protein product can diffuse and regulate multiple operons.
Summary of the lac Operon Model
No lactose: Repressor binds operator, blocking transcription.
Lactose present: Lactose binds repressor, repressor releases operator, transcription occurs.
All lactose metabolized: Repressor binds operator again, transcription stops.
Catabolite Repression and Positive Control by CAP
When both glucose and lactose are present, E. coli prefers glucose. The presence of glucose inhibits the lac operon through a mechanism called catabolite repression.
Catabolite-activating protein (CAP): Exerts positive control over the lac operon.
cAMP: Cyclic AMP is produced from ATP by adenyl cyclase. Glucose inhibits adenyl cyclase, so cAMP levels are low when glucose is present.
CAP-cAMP complex: When glucose is absent, cAMP levels rise, cAMP binds to CAP, and the complex binds to the promoter, enhancing RNA polymerase binding and transcription.
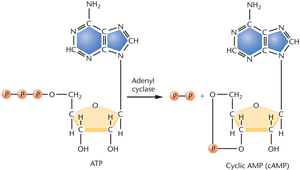
Mechanism of Catabolite Repression
High glucose: Low cAMP, CAP cannot bind, lac operon transcription is diminished.
Low glucose: High cAMP, CAP-cAMP binds promoter, lac operon transcription is activated (if lactose is present).
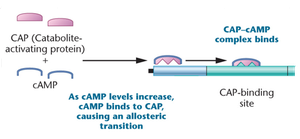
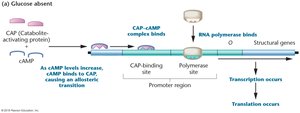
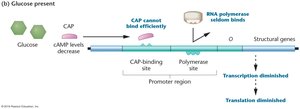
Summary Table: Regulation of the lac Operon
Condition | cAMP Level | CAP Binding | lac Operon Expression |
|---|---|---|---|
Glucose present, lactose absent | Low | No | Off |
Glucose absent, lactose present | High | Yes | On |
Glucose present, lactose present | Low | No | Very low |
Glucose absent, lactose absent | High | Yes | Off |
The trp Operon: A Repressible Gene System
Structure and Function of the trp Operon
The trp operon in E. coli is a repressible operon responsible for the biosynthesis of tryptophan. It consists of five structural genes (trpE, trpD, trpC, trpB, trpA), a promoter (trpP), and an operator (trpO).
Repressible system: The operon is usually on but can be turned off (repressed) when tryptophan is present.
Corepressor: Tryptophan acts as a corepressor by binding to the trp repressor protein, enabling it to bind the operator and block transcription.
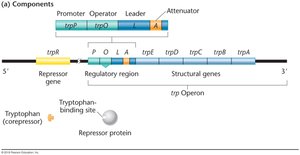
Regulation of the trp Operon
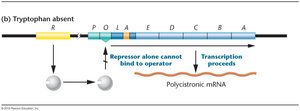
Tryptophan absent: Repressor cannot bind operator, transcription proceeds, tryptophan is synthesized.
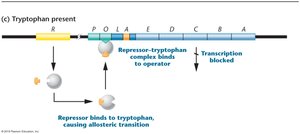
Tryptophan present: Tryptophan binds repressor, repressor binds operator, transcription is blocked.
Mutations Affecting the trp Operon
Repressor cannot bind DNA: Transcription of the trp operon occurs all the time, regardless of tryptophan presence.
Repressor cannot bind tryptophan: The operon is always on, as the repressor cannot be activated to bind the operator.
RNA-Based Regulation in Bacteria
Small Noncoding RNAs (sRNAs)
Bacterial small noncoding RNAs (sRNAs) play important roles in regulating gene expression post-transcriptionally.
Negative regulation: sRNAs can bind to mRNAs and block the ribosome-binding site, preventing translation.
Positive regulation: sRNAs can bind to mRNAs and prevent the formation of inhibitory secondary structures, enabling translation.
Additional info: sRNAs are also important in eukaryotic gene regulation, such as microRNAs (miRNAs) and small interfering RNAs (siRNAs).
Summary
Bacterial gene expression is regulated at the transcriptional level by proteins that respond to environmental signals.
The lac operon is an inducible system controlled by both negative (repressor) and positive (CAP-cAMP) mechanisms.
The trp operon is a repressible system, turned off when tryptophan is abundant.
Small RNAs provide additional layers of regulation, fine-tuning gene expression in response to cellular needs.
