 Back
BackRegulation of Gene Expression in Bacteria: The lac Operon
Study Guide - Smart Notes

Regulation of Gene Expression in Bacteria
Introduction to Gene Regulation
Gene regulation is the process by which cells control the transcription of genetic information from DNA to RNA, determining when, where, and how much gene product is made. This regulation is essential for cellular efficiency, differentiation, and adaptation to environmental changes.
Gene regulation allows cells to conserve energy and resources by producing proteins only when needed.
It enables cell differentiation in multicellular organisms and allows cells to respond to environmental signals.
Differences in gene expression, rather than gene content, can explain significant phenotypic differences between closely related species, such as humans and chimpanzees.

Types of Prokaryotic Gene Regulation
Prokaryotes regulate gene expression primarily in response to environmental conditions, such as nutrient availability and metabolic needs. This regulation is highly efficient and ensures that only necessary genes are expressed at any given time.
Inducible systems: Genes are transcribed only in the presence of a specific inducer (e.g., enzymes for lactose metabolism are produced only when lactose is present).
Repressible systems: Genes are transcribed unless a specific corepressor is present, which stops transcription (e.g., enzymes for amino acid synthesis are repressed when the amino acid is abundant).
Mechanisms of Gene Control
Negative control: Transcription occurs unless it is shut off by a repressor protein binding to the DNA.
Positive control: Transcription occurs only if an activator protein stimulates RNA polymerase activity.
Key DNA Sequences in Gene Regulation
Several DNA elements are critical for the regulation of gene expression:
Promoter: The DNA sequence where RNA polymerase binds to initiate transcription.
Operator: A regulatory DNA sequence that interacts with repressor or activator proteins to control transcription.
Coding sequence: The region of DNA that is transcribed into mRNA and translated into protein.
Terminator: The DNA sequence signaling the end of transcription.
The lac Operon: A Model for Gene Regulation in Bacteria
Structure and Function of the lac Operon
The lac operon in Escherichia coli is a classic example of gene regulation. It enables the bacterium to metabolize lactose only when it is available and glucose is absent.
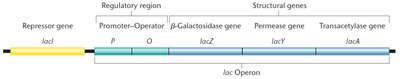
The operon consists of a promoter, operator, and three structural genes: lacZ (β-galactosidase), lacY (permease), and lacA (transacetylase).
These genes are transcribed together as a single polycistronic mRNA.
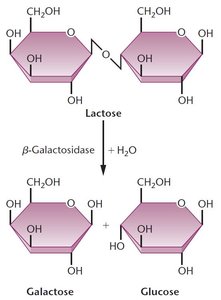
Lactose Metabolism and the Role of β-Galactosidase
Lactose, a disaccharide, is hydrolyzed into glucose and galactose by the enzyme β-galactosidase. Glucose can then enter glycolysis for energy production.
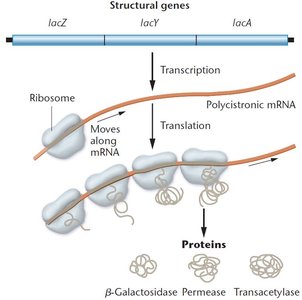
Transcription and Translation of the lac Operon
When the lac operon is active, all three enzymes required for lactose metabolism are synthesized from a single mRNA transcript.
β-galactosidase (lacZ): Cleaves lactose into glucose and galactose.
Permease (lacY): Facilitates lactose entry into the cell.
Transacetylase (lacA): Function is less well understood.
Negative Control: The lac Repressor
The lac operon is primarily regulated by a repressor protein encoded by the lacI gene. In the absence of lactose, the repressor binds to the operator, blocking RNA polymerase and preventing transcription.
The operator is a cis-acting element (must be adjacent to the promoter).
The repressor is a trans-acting factor (diffusible protein).
When lactose is present, it binds to the repressor, causing an allosteric change that prevents the repressor from binding the operator, allowing transcription to proceed.
Mutations Affecting the lac Operon
I- mutations: Result in a nonfunctional repressor, leading to constitutive expression of the operon.
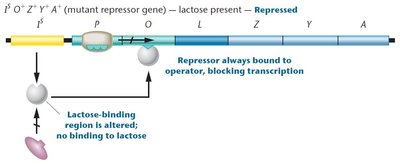
IS mutations: Produce a super-repressor that cannot bind lactose, so the operon is always repressed.
OC mutations: Alter the operator so the repressor cannot bind, resulting in constitutive expression.
Positive Control: Catabolite Repression and the Role of CAP
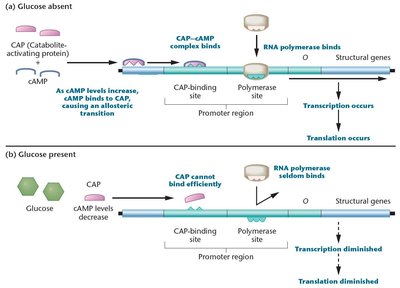
When both glucose and lactose are present, the cell preferentially metabolizes glucose. The presence of glucose inhibits the lac operon through a mechanism called catabolite repression, mediated by the catabolite-activating protein (CAP).
CAP requires cyclic AMP (cAMP) to bind DNA and activate transcription.
Glucose inhibits adenyl cyclase, reducing cAMP levels and preventing CAP from activating the operon.
Only when glucose is absent do cAMP levels rise, allowing CAP to bind and stimulate transcription of the lac operon.
Summary Table: Key Components of the lac Operon
Component | Type | Function |
|---|---|---|
lacI | Regulatory gene | Encodes the lac repressor protein |
Promoter (P) | Regulatory region | Binding site for RNA polymerase |
Operator (O) | Regulatory region | Binding site for the repressor protein |
lacZ | Structural gene | Encodes β-galactosidase |
lacY | Structural gene | Encodes permease |
lacA | Structural gene | Encodes transacetylase |
CAP | Regulatory protein | Activates transcription in the absence of glucose |
Key Equations
Lactose hydrolysis by β-galactosidase:
Additional info: The lac operon is a foundational model for understanding gene regulation in prokaryotes and has informed the study of more complex regulatory systems in eukaryotes.
