 Back
BackRegulation of Gene Expression in Bacteria: trp Operon, Attenuation, Riboswitches, sRNAs, and CRISPR/Cas
Study Guide - Smart Notes

Regulation of Gene Expression in Bacteria
The trp Operon: A Repressible System
The trp operon is a classic example of a repressible system in bacteria, specifically Escherichia coli. It regulates the synthesis of tryptophan, an essential amino acid, through negative control mechanisms.
Operon Structure: The trp operon consists of five structural genes encoding enzymes for tryptophan biosynthesis, a promoter, an operator, and a regulatory gene (trpR).
Negative Control: The repressor protein, encoded by trpR, requires tryptophan (corepressor) to bind the operator and inhibit transcription.
Regulation: When tryptophan is absent, the repressor cannot bind DNA, and the operon is transcribed. When tryptophan is present, it binds the repressor, enabling it to bind the operator and block transcription.
Mutations:
trpR-: Mutations in the repressor gene prevent repression, leading to constitutive expression.
trpOC: Mutations in the operator prevent repressor binding, also resulting in constitutive expression.
Example: The trp operon allows bacteria to conserve energy by only producing tryptophan-synthesizing enzymes when tryptophan is scarce.
Attenuation: Regulation via the 5' UTR
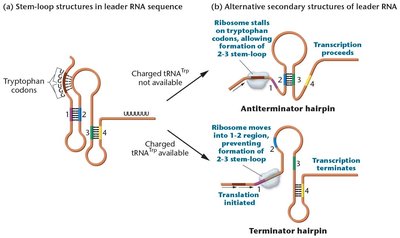
Attenuation is a regulatory mechanism involving the leader sequence (5' UTR) of the trp operon. It fine-tunes gene expression based on tryptophan availability.
Leader Sequence: Contains an attenuator region that can form alternative RNA secondary structures.
Mechanism:
If tryptophan is scarce, ribosomes stall, forming an antiterminator hairpin, allowing transcription to proceed.
If tryptophan is abundant, ribosomes do not stall, forming a terminator hairpin, causing transcription termination.
Example: Attenuation ensures that the operon is not transcribed when tryptophan is plentiful, even if the repressor is inactive.
Riboswitches: Metabolite-Sensing mRNA Structures
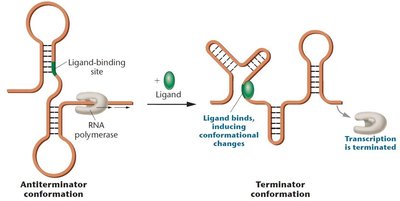
Riboswitches are regulatory segments within mRNA that bind small molecules (ligands) and alter mRNA secondary structure, affecting transcription or translation.
Structure: Consists of an aptamer (ligand-binding site) and an expression platform (switching region).
Function: Ligand binding induces conformational changes, switching between antiterminator and terminator forms.
Regulation:
Antiterminator conformation allows transcription to proceed.
Terminator conformation halts transcription.
Example: Riboswitches regulate genes involved in metabolite biosynthesis, such as vitamins and amino acids.
sRNAs: Small Noncoding RNAs in Bacterial Gene Regulation
sRNAs (small noncoding RNAs) are regulatory RNAs that modulate gene expression post-transcriptionally, often in response to environmental changes.
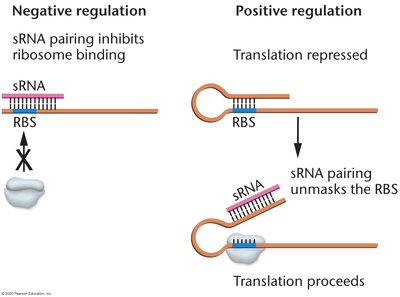
Mechanism: sRNAs are complementary to target mRNAs and can either inhibit or promote translation.
Negative Regulation: sRNA pairing blocks ribosome binding, repressing translation.
Positive Regulation: sRNA pairing exposes the ribosome binding site, activating translation.
Example: sRNAs are involved in stress responses, iron metabolism, and quorum sensing.
CRISPR/Cas: Adaptive Immunity in Prokaryotes
The CRISPR/Cas system is a prokaryotic adaptive immune mechanism that provides sequence-specific defense against invading phages and plasmids.
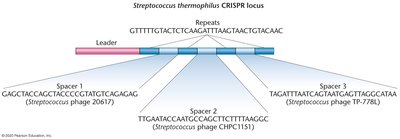
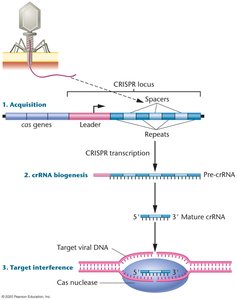
CRISPR Locus: Contains repeats and spacers derived from foreign DNA.
Mechanism:
Spacer Acquisition: Invading DNA is cleaved and integrated as spacers in the CRISPR locus.
crRNA Biogenesis: CRISPR locus is transcribed and processed into crRNAs, each containing a spacer.
Target Interference: crRNAs guide Cas nucleases to complementary sequences in invading DNA, leading to cleavage and neutralization.
Example: CRISPR/Cas is used in biotechnology for genome editing.
Summary Table: Regulation Mechanisms in Bacteria
Mechanism | Regulatory Element | Mode of Action | Example |
|---|---|---|---|
trp Operon | Repressor, Operator | Negative control, repressible | Tryptophan biosynthesis |
Attenuation | Leader sequence, attenuator | Transcription termination | trp operon |
Riboswitch | Aptamer, expression platform | Ligand-induced conformational change | Vitamin biosynthesis |
sRNA | sRNA, mRNA | Translation regulation | Stress response |
CRISPR/Cas | CRISPR locus, Cas proteins | Adaptive immunity | Phage defense |
Key Terms and Definitions
Operon: A cluster of genes under the control of a single promoter and regulatory elements.
Repressible System: A gene regulatory system that is turned off in the presence of a specific substance.
Attenuation: Regulation of transcription termination via RNA secondary structure.
Riboswitch: mRNA element that binds metabolites and regulates gene expression.
sRNA: Small noncoding RNA that regulates translation.
CRISPR/Cas: Prokaryotic adaptive immune system using RNA-guided nucleases.
Relevant Equations and Concepts
Polycistronic mRNA: Single mRNA molecule encoding multiple proteins.
Transcription Termination:
CRISPR Spacer Integration:
Applications
Biotechnology: CRISPR/Cas is widely used for genome editing in research and medicine.
Microbial Genetics: Understanding operon regulation aids in metabolic engineering and antibiotic development.
