Back
BackCH17&18: Regulation of Gene Expression in Eukaryotes: Mechanisms and Molecular Features
Study Guide - Smart Notes

Regulation of Gene Expression in Eukaryotes
Overview of Eukaryotic Gene Regulation
Gene expression in eukaryotes is a highly regulated process, allowing cells to produce gene products at specific times, in specific cell types, and in response to environmental changes. Unlike prokaryotes, eukaryotic gene regulation is more complex due to the larger amount of DNA, multiple chromosomes, spatial separation of transcription and translation, mRNA processing, RNA stability, and cellular differentiation.
Temporal regulation: Genes are expressed at particular times during development or in response to signals.
Spatial regulation: Expression occurs in specific cell types or tissues.
Environmental response: Gene expression adapts to external stimuli.
Chromosome Organization and Chromatin Modifications
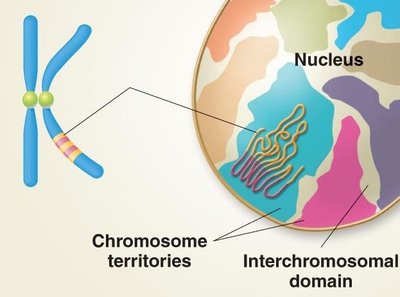
Eukaryotic gene expression is influenced by the structural organization of chromosomes and chromatin modifications. Chromosomes occupy distinct regions within the nucleus, known as chromosome territories, separated by interchromosomal domains.
Chromosome territories: Each chromosome resides in a discrete domain within the nucleus, maintaining separation from others.
Interchromosomal domains: Channels between chromosome territories with little or no DNA, facilitating molecular trafficking.
Chromatin Remodeling
Chromatin remodeling is essential for regulating gene expression, as it determines the accessibility of DNA to transcription factors and RNA polymerase. Remodeling can occur via modification of nucleosomes or DNA.
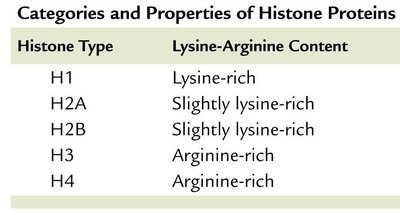
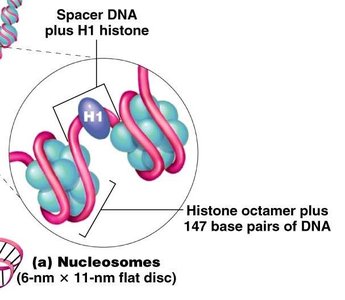
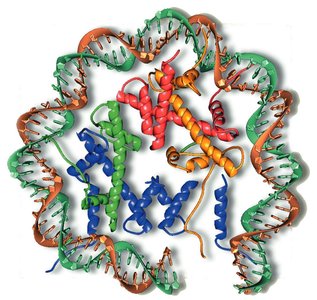
Nucleosome composition: Variant histones (e.g., H2A.Z, H3.3) are found in nucleosomes flanking active promoters.
Histone modifications: Covalent modifications such as acetylation, phosphorylation, and methylation alter chromatin structure and gene activity.
Nucleosome repositioning: Remodeling complexes (e.g., SWI/SNF) shift nucleosomes to expose regulatory regions.
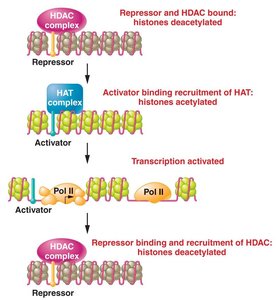
Histone Modification
Acetylation: Addition of acetyl groups by histone acetyltransferase (HAT) neutralizes lysine charge, loosening chromatin and promoting transcription.
Deacetylation: Histone deacetylases (HDACs) remove acetyl groups, condensing chromatin and repressing transcription.
Phosphorylation: Addition of phosphate groups by kinases introduces negative charge, affecting chromatin structure.
Methylation: Addition of methyl groups by methyltransferases, often linked to gene repression.
Histone code: The pattern of histone modifications determines gene activation or silencing.
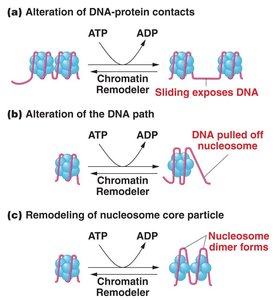
Nucleosome Remodeling Complexes
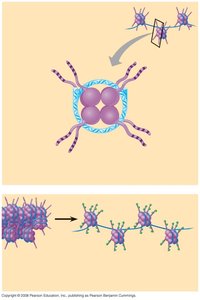
Remodeling complexes use ATP to alter nucleosome structure, exposing DNA for transcription.
Alter DNA-protein contacts
Change DNA path around nucleosome
Remodel nucleosome core particle
DNA Methylation
DNA methylation involves the addition of methyl groups to cytosine residues, particularly in CpG islands. Heavily methylated promoter regions are associated with gene silencing.
CpG islands: CG-rich regions in promoters; methylation reduces transcription.
Inactive X chromosome: More heavily methylated than active X.
5-azacytidine: Base analog that prevents methylation, activating silent genes.
Transcriptional Regulation: Cis-Acting Elements and Transcription Factors
Cis-Acting Elements
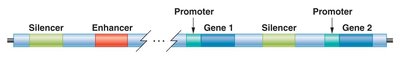
Cis-acting elements are DNA sequences that regulate gene expression on the same chromosome. Key types include promoters, enhancers, and silencers.
Promoters: Recognition sites for transcription machinery, located adjacent to regulated genes.
Enhancers: Modular DNA sequences that increase transcription rates, can be located far from the gene.
Silencers: DNA elements that repress transcription initiation.
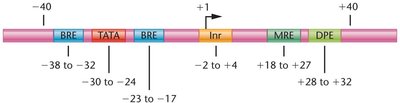
Promoter Structure
Promoters consist of core elements and proximal promoter elements, such as TATA boxes, CAAT boxes, and GC boxes, which bind transcription factors and enhance basal transcription.
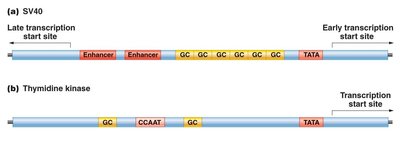
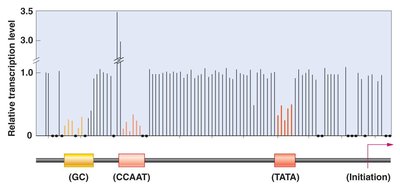
Transcription Factors
Transcription factors are proteins that regulate gene expression by binding to cis-acting sites. They can act as activators or repressors, and their effects are finely tuned according to cell type, environmental cues, or developmental stage.
Activators: Increase transcription initiation.
Repressors: Decrease transcription initiation.
Functional domains: DNA-binding domain and trans-activating domain.
DNA-Binding Motifs
Helix-turn-helix: Two adjacent α-helices separated by a turn, enabling DNA binding.
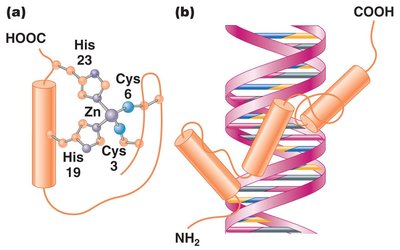
Zinc finger: Clusters of cysteines and histidines bind zinc atoms, forming finger-like projections that interact with DNA.
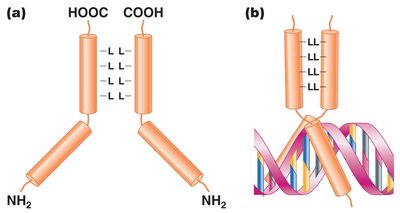
Leucine zipper: Leucine-rich region allows dimerization; α-helical regions bind DNA.
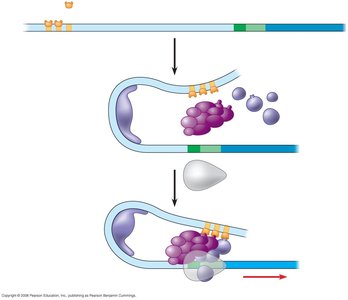
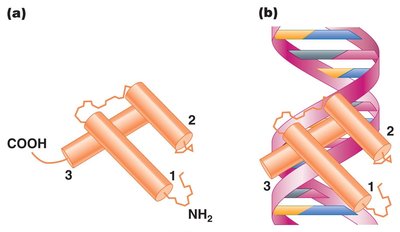
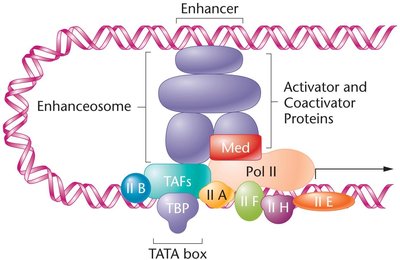
Transcription Initiation Complex
Basal transcription factors assemble at the promoter, facilitating RNA polymerase II binding. TFIID binds the TATA box, recruiting additional factors and RNA polymerase II. Activators and coactivators form enhanceosomes, interacting with general transcription factors to modulate transcription rates.
Posttranscriptional Regulation
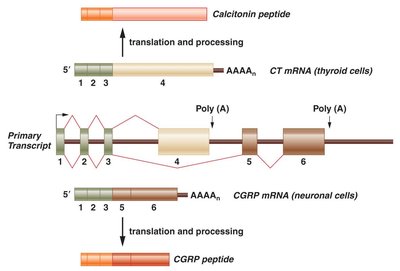
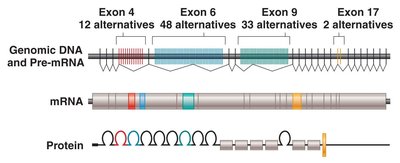
Alternative RNA Splicing
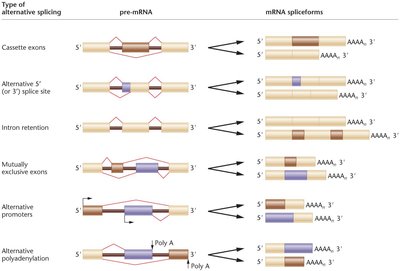
Alternative splicing generates multiple protein isoforms from a single gene, increasing proteomic diversity. Types include cassette exons, alternative splice sites, intron retention, mutually exclusive exons, alternative promoters, and polyadenylation.
Control of mRNA Stability
The steady-state level of mRNA is determined by transcription rate and degradation rate. mRNA stability is influenced by sequences in the 5' and 3' untranslated regions (UTRs), poly-A tail length, and cap structure.
Poly-A tail shortening destabilizes mRNA.
Decapping enzymes remove the 5' cap, leading to degradation.
Endonucleases cleave mRNA internally, especially during nonsense-mediated decay.

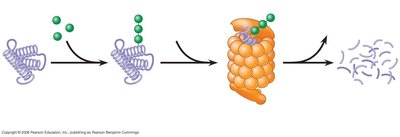
Protein Stability and Posttranslational Modification
After translation, proteins undergo modifications such as cleavage, phosphorylation, methylation, acetylation, glycosylation, and complexing with metals. Proteasomes degrade proteins tagged with ubiquitin, regulating protein levels and function.
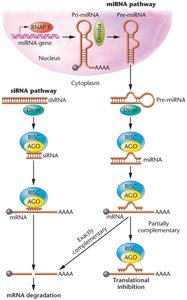
RNA Silencing and RNA Interference (RNAi)
RNA-Induced Gene Silencing
Short RNA molecules, including siRNAs and miRNAs, regulate gene expression by repressing translation or triggering mRNA degradation. RNAi is a sequence-specific posttranscriptional regulatory mechanism.
siRNAs: Arise from viral infection or exogenous sources, processed by Dicer.
miRNAs: Endogenous noncoding RNAs that negatively regulate gene expression.
RISC: RNA-induced silencing complex, containing Argonaut proteins with endonuclease activity.
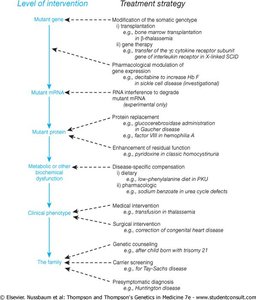
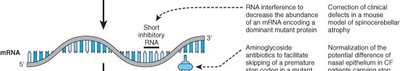
Therapeutic Applications of RNAi
RNAi technology is being developed to treat diseases caused by overexpression or abnormal expression of specific genes. RNAi can selectively degrade mutant mRNAs, reducing the production of harmful proteins.