 Back
BackRegulation of Gene Expression in Eukaryotes – Study Notes
Study Guide - Smart Notes

Regulation of Gene Expression in Eukaryotes
Introduction to Eukaryotic Gene Regulation
Gene expression in eukaryotes is a highly regulated process, ensuring that genes are expressed in the right cell type, at the right time, and in the correct amount. This regulation is essential for cellular differentiation, response to environmental signals, and prevention of diseases such as cancer.
Cell-type specific expression: Different cell types express unique sets of genes (e.g., keratin in skin cells, myosin in muscle cells).
Conditional expression: Genes can be upregulated or downregulated in response to physiological conditions (e.g., erythropoietin in response to low oxygen).
Misregulation: Incorrect gene expression can lead to developmental defects, cancer, or cell death.
Cis-acting and trans-acting factors: These are fundamental for regulating transcription initiation.
Levels of Gene Regulation in Eukaryotes
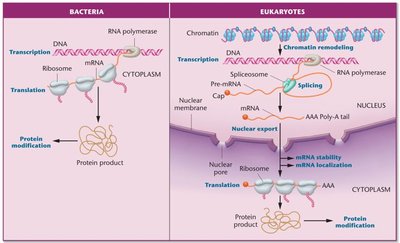
Gene expression in eukaryotes is regulated at multiple levels, including transcription, RNA processing, mRNA stability, translation, and post-translational modifications.
Transcriptional regulation
RNA splicing and processing
mRNA degradation
Translational regulation
Post-translational modifications
Chromatin Structure and Its Role in Gene Regulation
Chromatin Organization
Eukaryotic DNA is packaged with histones and nonhistone proteins to form chromatin. The basic unit of chromatin is the nucleosome, which consists of DNA wrapped around histone proteins. Chromatin structure can regulate access to DNA for processes such as replication, repair, and transcription.
Nucleosomes: Repeating DNA-histone complexes that compact DNA.
Chromatin compaction: Inhibits DNA replication, repair, and transcription.
Chromatin Alterations: Nucleosome and Histone Modifications
Chromatin can switch between open (euchromatin) and closed (heterochromatin) states, influenced by histone composition and covalent modifications.
Histone variants: Some nucleosomes contain H2A.Z, which is less stable and can affect transcription.
Histone modifications: Covalent addition of acetyl, methyl, or phosphate groups to histone tails alters chromatin structure and gene expression.
Acetylation: Decreases histone-DNA affinity, generally associated with increased transcription.
Histone acetyltransferases (HATs): Enzymes that add acetyl groups, promoting transcription.
Histone deacetylases (HDACs): Remove acetyl groups, often repressing transcription.
DNA Methylation
DNA methylation is a key epigenetic modification that typically represses gene expression. It occurs mainly at cytosines in CpG dinucleotides, often clustered in CpG islands near gene promoters.
Function: Silences transcription, plays a role in X-chromosome inactivation and genomic imprinting.
Location: Most common at the 5-position of cytosine in CpG doublets.
Transcriptional Regulation
Cis-acting DNA Elements and Trans-acting Factors
Transcription in eukaryotes requires the interaction of regulatory proteins (trans-acting factors) with specific DNA sequences (cis-acting elements) near or within genes.
Promoters: DNA regions recognized by RNA polymerase and transcription factors, required for transcription initiation.
Core promoter: Specifies the transcription start site and is essential for accurate initiation.
Proximal-promoter elements: Located upstream of the core promoter, enhance basal transcription levels (e.g., CAAT box, GC box).
Classification of Core Promoters
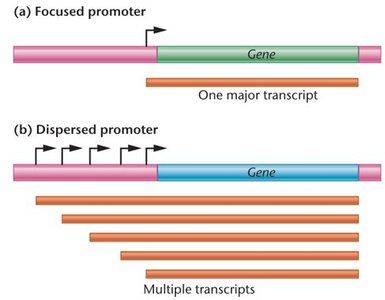
Core promoters can be classified as focused or dispersed, affecting the pattern of transcription initiation.
Focused promoters: Direct transcription from a single start site; common in lower eukaryotes.
Dispersed promoters: Initiate transcription from multiple weak start sites over a region; common in higher eukaryotes.
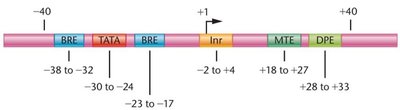
Core-Promoter Elements
Focused promoters contain several conserved elements that facilitate the assembly of the transcription machinery.
Initiator (Inr) element
TATA box
TFIIB recognition element (BRE)
Motif ten element (MTE)
Downstream promoter element (DPE)
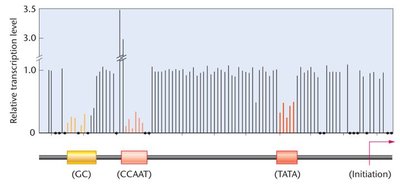
Proximal-Promoter Elements
These elements are located upstream of the core promoter and are important for enhancing transcription levels. Mutations in these regions can significantly affect gene expression.
CAAT box
GC box
Enhancers and Silencers
Enhancers and silencers are cis-acting elements that can be located far from the gene they regulate. They modulate the rate of transcription initiation in a time- and tissue-specific manner.
Enhancers: Increase transcription rates, can function at a distance and in either orientation.
Silencers: Repress transcription, often in a tissue- or temporal-specific manner.
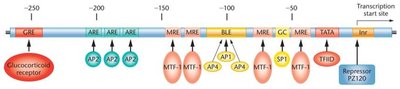
Transcription Regulatory Proteins
Transcription factors bind to promoters, enhancers, and silencers to regulate gene expression. Activators increase transcription, while repressors decrease it. Multiple factors can fine-tune gene expression.
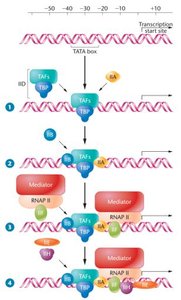
Pre-Initiation Complex (PIC) Formation
The assembly of the pre-initiation complex is a critical step in transcription initiation. It involves RNA polymerase II and general transcription factors (GTFs), beginning with the binding of TFIID to the TATA box.
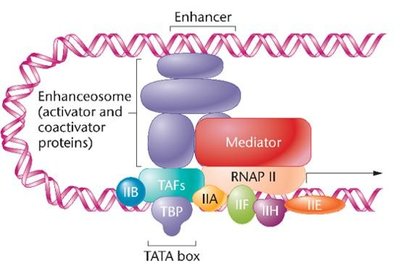
Mechanism of Transcription Activation and Repression
Transcription activators and repressors modulate gene expression by facilitating or hindering the assembly of the transcription machinery. DNA looping allows distant enhancers or silencers to interact with the promoter region.
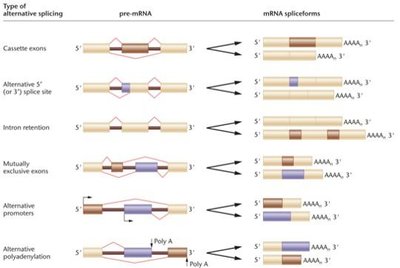
Post-Transcriptional Regulation
Alternative Splicing of mRNA
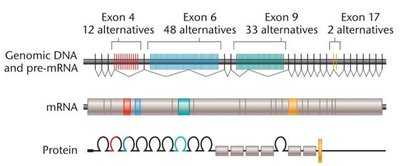
Alternative splicing allows a single gene to produce multiple protein isoforms by including or excluding different exons during mRNA processing. This increases proteomic diversity.
Spliceforms: Different mRNA variants produced from the same pre-mRNA.
Isoforms: Protein variants with potentially different functions.
Alternative Splicing and the Proteome
Alternative splicing greatly expands the number of proteins an organism can produce. For example, the Drosophila Dscam gene can generate over 38,000 different proteins through alternative splicing of its exons.
RNA-Binding Proteins (RBPs) and Spliceopathies
RBPs regulate alternative splicing by binding to specific RNA sequences, influencing splice site selection. Mutations affecting splicing can cause genetic disorders known as spliceopathies (e.g., myotonic dystrophy).
mRNA Stability and Degradation
The steady-state level of mRNA in a cell is determined by the balance between synthesis and degradation. mRNA decay is often initiated by deadenylation (removal of the poly-A tail) and is carried out by exoribonucleases.
Nonsense-Mediated Decay (NMD)
NMD is a surveillance pathway that degrades mRNAs containing premature stop codons, preventing the production of truncated, potentially harmful proteins.
mRNA Localization
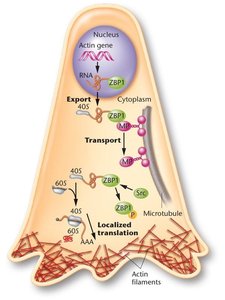
Some mRNAs are transported to specific cellular locations for localized translation. For example, actin mRNA is localized to the leading edge of migrating cells, where it is translated to support cell movement. This process is mediated by zip code sequences and RNA-binding proteins.
Post-Translational Regulation
Posttranslational Modifications
Proteins can be regulated after translation by covalent modifications such as phosphorylation and ubiquitination.
Phosphorylation: Addition of phosphate groups by kinases; removal by phosphatases. Regulates protein activity, localization, and interactions.
Ubiquitination: Attachment of ubiquitin molecules targets proteins for degradation by the proteasome.
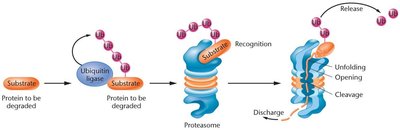
Proteasome-Mediated Protein Degradation
The proteasome is a large protein complex that degrades proteins tagged with ubiquitin. This process regulates protein levels and removes damaged or misfolded proteins, playing a critical role in gene expression regulation.
