 Back
BackRegulation of Gene Expression in Prokaryotes: Lac and Trp Operons
Study Guide - Smart Notes

Gene Expression in Prokaryotes
Overview of Gene Expression Regulation
Gene expression in prokaryotes is tightly regulated to ensure that proteins are synthesized only when needed. This regulation is critical for cellular efficiency and adaptation to environmental changes. The primary mechanisms involve DNA-binding proteins, operon systems, and feedback from metabolic products.
Operon: A cluster of genes under the control of a single promoter and regulatory elements, allowing coordinated expression.
Inducible Operon: Transcription is usually off and needs to be turned on (e.g., lac operon).
Repressible Operon: Transcription is normally on and needs to be turned off (e.g., trp operon).
DNA Binding Proteins: Regulatory proteins (repressors and activators) bind to specific DNA sequences to control transcription.
Key Concepts and Terminology
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
Operator: DNA sequence where regulatory proteins (repressors) bind.
Regulatory Genes: Encode proteins that control operon activity (e.g., lacI and trpR).
Structural Genes: Encode enzymes or proteins involved in metabolic pathways.
Lac Operon: Inducible System in E. coli
Structure and Regulation
The lac operon is a classic example of an inducible operon, regulating lactose metabolism in E. coli. It consists of regulatory and structural genes, and its expression is controlled by the presence or absence of lactose and glucose.
lacI: Encodes the repressor protein.
Operator (O): Binding site for the repressor.
Structural Genes: lacZ (β-galactosidase), lacY (permease), lacA (transacetylase).
Regulation:
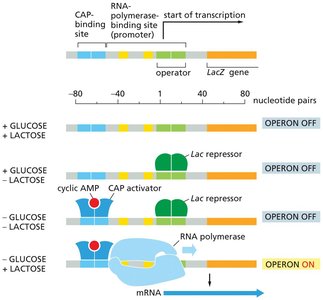
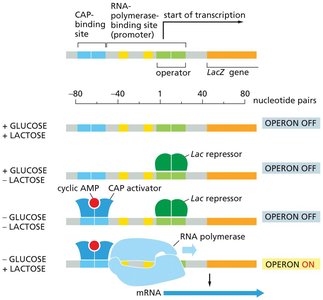
Lactose absent: The lac repressor binds to the operator, blocking transcription.
Lactose present: Lactose (or allolactose) binds to the repressor, causing a conformational change that releases it from the operator, allowing transcription.
Glucose absent: cAMP levels rise, CAP (catabolite activator protein) binds to DNA, enhancing transcription.
Optimal expression: Occurs when lactose is present and glucose is absent.
Mutant Analysis of the Lac Operon
Mutations in regulatory or structural components of the lac operon can alter gene expression patterns.
I- mutant: Repressor protein is absent or altered; cannot bind operator. Structural genes are constitutively expressed.
OC mutant: Operator DNA sequence is altered; repressor cannot bind. Structural genes are constitutively expressed.
IS mutant: Superrepressor; cannot be inactivated by inducer. Structural genes are always repressed.
Summary Table: Lac Operon Mutants
Mutant Type | Effect on Expression | Mechanism |
|---|---|---|
I- | Constitutive | Repressor absent/altered |
OC | Constitutive | Operator altered, repressor cannot bind |
IS | Repressed | Superrepressor, cannot be inactivated |
Trp Operon: Repressible System in E. coli
Structure and Regulation
The trp operon regulates the biosynthesis of tryptophan in E. coli. It is a repressible operon, meaning transcription is normally on and is turned off when tryptophan is abundant.
trpR: Encodes the repressor protein (located elsewhere on the chromosome).
Promoter (trpP): RNA polymerase binding site.
Operator (trpO): Repressor binding site.
Leader (trpL): Contains attenuator sequences.
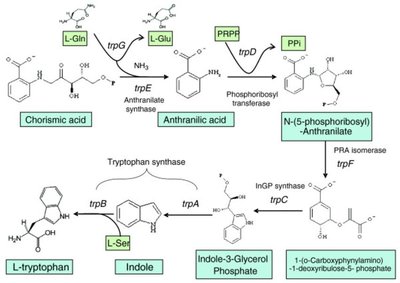
Structural Genes: trpE, trpD, trpC, trpB, trpA (encode enzymes for tryptophan biosynthesis).
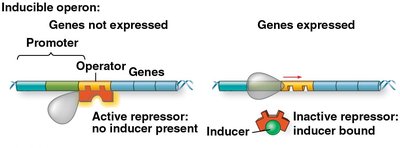
Regulation by Repressor and Corepressor
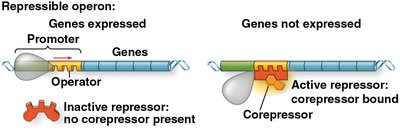
Low tryptophan: Repressor is inactive; operon is transcribed.
High tryptophan: Tryptophan acts as a corepressor, binds to the repressor, activating it. The active repressor binds to the operator, blocking transcription.
Attenuation Mechanism
Attenuation is a secondary regulatory mechanism that fine-tunes trp operon expression based on tryptophan levels. It relies on the coupling of transcription and translation in prokaryotes.
Leader sequence: Contains four regions that can form hairpin structures.
High tryptophan: Ribosome quickly translates leader peptide, allowing region 3 to pair with region 4, forming a termination hairpin that halts transcription.
Low tryptophan: Ribosome stalls at Trp codons, region 2 pairs with region 3, preventing the formation of the termination hairpin, and transcription continues.
Summary Table: Trp Operon Regulation
Condition | Repressor State | Transcription Outcome |
|---|---|---|
Low tryptophan | Inactive | Transcription proceeds |
High tryptophan | Active (corepressor bound) | Transcription blocked |
High tryptophan (attenuation) | Active | Premature termination via hairpin |
Comparison: Inducible vs. Repressible Operons
Inducible Operon (lac): Genes are off unless an inducer (lactose) is present.
Repressible Operon (trp): Genes are on unless a corepressor (tryptophan) is present.
Transcription and Translation Coupling in Prokaryotes
In prokaryotes, transcription and translation occur simultaneously, which is essential for mechanisms like attenuation in the trp operon.
Coupling: Ribosomes begin translating mRNA while it is still being transcribed.
Attenuation: Depends on ribosome position relative to RNA polymerase and leader sequence.
Summary
Gene expression in prokaryotes is regulated by operon systems, with the lac operon as an inducible model and the trp operon as a repressible model.
Mutations in regulatory elements can lead to constitutive or repressed expression.
Attenuation provides an additional layer of regulation for the trp operon.
Transcription and translation coupling is crucial for regulatory mechanisms in bacteria.
