 Back
BackRNA Splicing Mechanisms and RNA Editing
Study Guide - Smart Notes

Splicing Mechanisms: Self-Splicing RNAs
Overview of RNA Splicing
RNA splicing is a critical posttranscriptional process in eukaryotic gene expression, involving the removal of non-coding sequences (introns) from precursor RNA transcripts and the joining of coding sequences (exons) to produce mature mRNA. This process ensures that only the necessary coding information is translated into proteins.
Introns: Non-coding sequences removed from the primary RNA transcript.
Exons: Coding sequences that are joined together to form mature mRNA.
Splicing occurs in various RNA types, including those produced in mitochondria and chloroplasts.
Self-Splicing Mechanisms (Group I Introns)
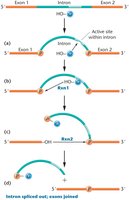
Some introns are capable of self-excision without the need for protein enzymes. These are known as self-splicing introns and are classified as group I or group II introns. The process involves two transesterification reactions that remove the intron and join the exons.
Occurs in certain mRNA and tRNA transcripts, especially in mitochondria and chloroplasts.
Group I introns are found in preliminary transcripts and can catalyze their own removal.
The process involves an endonuclease cutting each end of the intron, followed by ligation of the exons.
Mechanism for Intron Removal in tRNAs
In bacteria, intron removal from tRNA involves endonuclease activity to cut the intron and ligase activity to join the exons. In higher eukaryotes, the mechanism is more complex and often involves additional protein factors.
Endonuclease cuts at each end of the intron.
Ligase joins the terminal ends of adjacent exons.
Splicing Mechanisms: The Spliceosome
The Spliceosome Complex
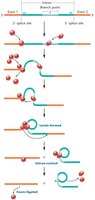
In most eukaryotic genes, intron removal is catalyzed by a large ribonucleoprotein complex called the spliceosome. The spliceosome is composed of small nuclear RNAs (snRNAs) and associated proteins, forming small nuclear ribonucleoproteins (snRNPs).
snRNPs (e.g., U1, U2): Essential components that recognize splice sites and catalyze the splicing reaction.
Spliceosomal splicing is characterized by the formation of a lariat structure during intron excision.
After splicing, the intron is removed and the exons are joined to form mature mRNA.
RNA Editing
Types of RNA Editing
RNA editing refers to posttranscriptional modifications that alter the nucleotide sequence of an RNA molecule after it has been synthesized. This process can diversify the transcriptome and proteome beyond the information encoded in the DNA.
Insertion/Deletion Editing: Nucleotides are added or deleted from the RNA sequence, changing the total number of bases.
Substitution Editing: The identity of individual nucleotide bases is altered, often by deamination or other chemical modifications.
RNA editing is especially prevalent in mitochondrial and chloroplast RNAs in plants.
Summary Table: Mechanisms of Intron Removal
Mechanism | Key Features | Location/Example |
|---|---|---|
Self-Splicing (Group I/II Introns) | Self-catalyzed, two transesterification reactions | Mitochondrial and chloroplast RNA, some nuclear genes |
Spliceosome-Mediated | snRNPs, lariat formation, complex protein machinery | Most nuclear pre-mRNA in eukaryotes |
tRNA Splicing (Bacteria) | Endonuclease and ligase activity | Bacterial tRNA genes |
